* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download MS Word File
Alternative splicing wikipedia , lookup
Non-coding DNA wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
RNA interference wikipedia , lookup
Molecular evolution wikipedia , lookup
Bottromycin wikipedia , lookup
Point mutation wikipedia , lookup
Metalloprotein wikipedia , lookup
Biochemistry wikipedia , lookup
RNA silencing wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Transcriptional regulation wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Polyadenylation wikipedia , lookup
Gene expression wikipedia , lookup
Messenger RNA wikipedia , lookup
Non-coding RNA wikipedia , lookup
Expanded genetic code wikipedia , lookup
Genetic code wikipedia , lookup
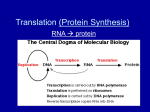
Transcription 6/8/12 Transcription-synthesis of RNA from DNA template Performed by RNA polymerase Holoenzyme made of 5 components in prokaryotes 2’ Three parts to replication: initiation, elongation, termination Initiation-getting RNA Polymerase to bind to DNA at appropriate locations Cis-acting sequences on chromosome direct initation- Promoters Prokaryotes -10 and –35 sequences TATAAT and TTGACA Eukaryotes TATA Box and CAAT box TATA box=AT rich sequence similar to –10; CAAT box=GGCCATTCT within 100 bases of start site is responsible for determining initiation site in prokaryotes alternate subunits promote binding at variable sites example: stress response transcription factors bind to site of initiation and then to RNAP II in eukaryotes Initiation also involves unwinding of DNA, removal of supercoiling, or chromatin restructuring before RNA Polymerase can bind. Elongation-polymerizing complementary ribonucleotide bases into an RNA molecule Similar in prokaryotes and eukaryotes In prokaryotes dissociates after approximately 8 bases have been added Remaining holoenzyme continues elongation Move along DNA strand 3’-5’ Synthesize RNA strand from 5’-3’ First base added is at 5’ end Most recently added base is at 3’ end Incorporates uracil instead of thymine in RNA molecule Termination-polymerase stops synthesizing RNA and falls off of DNA strand Newly formed RNA folds back on itself and makes a hairpin structure Hairpin may be bound by rho protein complex (rho-dependent termination) Hairpin structure helps RNA Polymerase dissociate from DNA strand In Eukaryotes mRNA undergoes maturation and splicing Takes place in nucleus 5’ cap and 3’ poly A tail are added splicing occurs introns removed and exons are spliced together Translation In Eukaryotes mRNA undergoes maturation and splicing Takes place in nucleus 5’ cap and 3’ poly A tail are added splicing occurs introns removed and exons are spliced together Translation-using mRNA template to synthesize protein Same three steps involved-initiation, elongation, and termination Genetic code is read so three nucleotides (codon) encode a single amino acid Genetic code found on page 314 of text book Has wobble-in many cases the first two bases of a codon determine the amino acid and the third is not essential (ACX=threonine) Proteins begin with formyl methionine (fMet) with AUG codon Three termination codons in code UAG, UGA, UAA Ribosomes, mRNA and tRNA are key players Ribosomes made of two subunits 60S and 40S in eukaryotes 50S and 30S in prokaryotes each comprises small proteins and rRNA have 3 sites:A-aminoacyl, P-peptidyl, E-exit tRNA has cloverleaf structure anticodon on loop will bind to complementary codon on mRNA amino acid binding site at 3’ end appropriate amino acid is added by aminoacyl tRNA synthetase separate enzyme for each amino acid (20 total) Initiation-Ribosome binds to mRNA near AUG codon (ribosome binding site) Small ribosomal subunit, fMet tRNA, and mRNA form initiation complex tRNA binds at P site of ribosome and is positioned at AUG codon Large subunit then comes to complete the complex Elongation-addition of new amino acids that are encoded by codons on mRNA Charged tRNA with anticodon sequence that is complementary to codon moves into A site The fMet forms peptide linkage to a.a. at A site The fMet is released by tRNA and is transferred to A site A site now has dipeptide Ribosome moves over one codon Moves uncharged tRNA to E site from which it leaves Moves dipeptide bound to tRNA to P site Opens A site for a new charged tRNA to enter Process repeats as long as codon at A site has complementary tRNA Termination-protein synthesis halts and ribosome dissociates from mRNA Happens when codon for which there is no tRNA enters A site Termination codons have no corresponding tRNA molecules May also happen prematurely when amino acid starvation occurs RNA Polymerase and ribosomes don’t respond to the same signals. Ribosome doesn’t begin it’s job at transcription start site.