* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Comparative mycobacterial genomics Stewart T Cole
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Essential gene wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Protein moonlighting wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Copy-number variation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Oncogenomics wikipedia , lookup
Genetic engineering wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Gene desert wikipedia , lookup
Gene expression programming wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Point mutation wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Ridge (biology) wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Genomic imprinting wikipedia , lookup
Transposable element wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Metagenomics wikipedia , lookup
Gene expression profiling wikipedia , lookup
Microevolution wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
History of genetic engineering wikipedia , lookup
Designer baby wikipedia , lookup
Genomic library wikipedia , lookup
Non-coding DNA wikipedia , lookup
Genome (book) wikipedia , lookup
Human Genome Project wikipedia , lookup
Human genome wikipedia , lookup
Public health genomics wikipedia , lookup
Helitron (biology) wikipedia , lookup
Pathogenomics wikipedia , lookup
Minimal genome wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
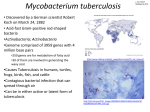
567 Comparative mycobacterial genomics Stewart T Cole Genomics is providing us with a mass of information about the biochemistry, physiology and pathogenesis of Mycobacterium tuberculosis and Mycobacterium leprae. Comparison of the two genome sequences is mutually enriching and indicates that the M. leprae genome appears to have undergone shrinkage and large-scale gene inactivation, which may account for the exceptionally slow growth of this organism. Addresses Unité de Génétique Moléculaire Bactérienne, Institut Pasteur, 28 rue du Docteur Roux, 75724 Paris Cedex 15, France; e-mail: [email protected] Current Opinion in Microbiology 1998, 1:567–571 http://biomednet.com/elecref/1369527400100567 © Current Biology Ltd ISSN 1369-5274 Abbreviations C cytosine G guanine MPTR major polymorphic tandem repeat PGRS polymorphic G + C rich sequence Introduction Intracellular pathogens such as the leprosy and tubercle bacilli are amongst the most genetically intractable microorganisms as a result of their long generation times, fastidious growth requirements and contagiousness. Fresh insight into the biology and pathogenicity of these important agents of human disease has been obtained by harnessing the powerful combination of genomics and bioinformatics. The availability of the complete genome sequence of Mycobacterium tuberculosis and much of the sequence of the chromosome of Mycobacterium leprae has resulted in a quantum leap in our knowledge and understanding. Comparison of the two datasets has identified both common and specific genes that may be of relevance to mycobacterial physiology and the respective diseases. This review presents some of the highlights of mycobacterial genome analysis and illustrates how comparative genomics may help us to understand phenotypic differences between related species. The genome sequence of M. tuberculosis H37Rv The complete genome sequence of the well characterised H37Rv strain of M. tuberculosis comprises 4,411,529 bp and contains around 4000 genes accounting for >91% of the potential coding capacity [1••,2•]. The high guanine (G) and cytosine (C) content of the DNA (65.6%) affects at least two other genomic parameters. First, the genome contains a partial complement of tRNA genes as only 43 of the 61 possible tRNAs are encoded indicating that extensive wobbling must occur during translation of mRNAs. Second, the amino acid content of the proteome is somewhat biased as amino acids encoded by G + C rich codons are statistically more abundant than expected. To identify regions with atypical base composition, that are often indicative of horizontally transferred genes or coding sequences involved in pathogenesis, the G + C composition of the genome was plotted and found to be relatively uniform. Several areas with exceptionally high G + C content (>80%) were detected and shown to correspond to the PGRS (polymorphic G + C rich sequence) gene family described below. Inspection of the regions with higher than average adenine (A) and thymine (T) content revealed genes encoding polyketide synthases or transmembrane proteins (hydrophobic amino acids are encoded by low G + C codons). Since these proteins are probably involved in housekeeping functions this suggests that the acquisition of virulence genes in the form of pathogenicity islands by horizontal transfer, as commonly described in Gram-negative bacteria [3], may not have occurred. The genome of H37Rv contains at least two prophages, phiRv1 and phiRv2 [1••], and one of these was probably lost during the attenuation process of M. bovis that gave rise to bacille Calmette-Guérin (BCG) [4]. PhiRv1 is unlikely to contribute to pathogenesis because it is not present in all clinical isolates of M. tuberculosis, although no comparative studies of the degrees of virulence of isogenic strains with or without the prophage have yet been performed. Genes, their functions and diversity Database comparisons (i.e. similarity of M. tuberculosis genes to other genes of known function) have led to the tentative attribution of functions to roughly 40% of the 3924 protein-coding genes identified and these are predominantly involved in core metabolism. Some functional information or similarity to other gene products was found for a further 44% of the protein-coding genes, although over half of these belong to the class known as conserved hypotheticals, proteins of unknown function but conserved sequence found in a variety of bacteria. The remaining genes are probably characteristic of mycobacteria as they show no similarity to any other microbial sequences. Recent improvements in transposon mutagenesis and gene replacement technology in mycobacteria [5•,6 •], coupled with more molecular methods such as DNA chip technology, serial analysis of gene expression and proteomics, should result in the elucidation of further functions. About 51% of the genes have arisen from gene duplication events. Although this value is close to that seen in Escherichia coli and Bacillus subtilis [7,8] (i.e. eubacteria with similar sized genomes), the degree of sequence conservation is much higher, suggesting that there may be 568 Genomics extensive functional redundancy or that M. tuberculosis is of recent evolutionary descent. Using systematic DNA sequence analysis of 26 loci in a very large number of independent strains, Musser and his colleagues have demonstrated that there is a singular lack of sequence diversity in the M. tuberculosis complex and conclude that this is indicative of recent global dissemination [9••,10]. The basis for this remarkable genetic homogeneity is unknown but most intriguing, and presumably reflects either a very efficient DNA repair system or replication machinery of exceptionally high fidelity. Inspection of the genome for genes associated with these functions provides some insight. First, M. tuberculosis may have an exceptionally clean pool of nucleotide precursors because it has three copies of mutT which encodes the enzyme that removes the oxidised guanines whose incorporation during replication causes base-pair mismatching. Second, the genome does not appear to contain a mismatch repair system because the key genes mutH, mutL, mutS and recJ could not be found. Because mispaired bases are the substrate for the MutHLS system and may occur less frequently in M. tuberculosis than in other bacteria due to the greater cleansing action of MutT, the mismatch repair system may not be required. The predominant gene families More than 20% of the M. tuberculosis chromosome is devoted to genes encoding two different classes of proteins: enzymes involved in fatty acid metabolism and acidic, glycine-rich polypeptides of unknown function, the PE and PPE proteins [1••,11]. The mycobacterial cell envelope contains a dazzling array of lipids, glycolipids, mycolic acids and polyketides [12,13], and it was thus no great surprise to find encoded in the genome examples of every known lipid and polyketide biosynthetic system, including enzymes usually confined to mammals and plants, in addition to the more common bacterial systems. More genes encoding potential lipid biosynthetic activities were uncovered than there are known metabolites thus raising the intriguing possibility that several novel lipid and polyketide species may exist [11]. It is conceivable that these unknown metabolites may only be produced in restricted environments such as the macrophage and that some of them may even have immunomodulatory activity like the potent polyketide, rapamycin, and account for the pronounced immune suppression observed in mycobacterial diseases. Although there are extensive lipid biosynthetic functions in M. tuberculosis, these are eclipsed by the vast selection of genes and enzymes potentially involved in fatty acid degradation. In addition to the multifunctional FadA/FadB proteins that effect all the reactions of the βoxidation cycle, there are >100 genes encoding enzymes that could catalyse the individual steps. These may well be involved in the degradation of lipids present in host membranes and could make important contributions to energy metabolism [14]. One of the great surprises of the M. tuberculosis genome project was the discovery of two large gene families encoding unusual glycine-rich proteins with basic pIs (isoelectric points) and well conserved amino-terminal domains. These show no significant similarity to proteins of known function and were referred to as the PE and PPE protein families as they contain conserved Pro–Glu and Pro–Pro–Glu motifs at the amino terminus, respectively [1••,11]. There are 99 members of the PE family and 61 of these belong to the PGRS subfamily, containing multiple tandem repetitions of the tripeptide Gly–Gly–Ala (or a variant thereof), whereas the remaining PE proteins have different carboxy-terminal domains. The PPE family comprises 68 members that fall into at least three subfamilies, the most spectacular of which is the MPTR (major polymorphic tandem repeat) class as several of these proteins are predicted to consist of >3000 amino acid residues mainly repetitions of the motif Asn-X-Gly-X-Gly-Asn-X-Gly (where X is any amino acid). The PGRS members of the PE subfamily are also large proteins and can contain up to 1400 amino acid residues. As the names PGRS and MPTR imply, the genes based on these sequences show extensive polymorphisms in the different M. tuberculosis complex strains probably as a result of strand slippage during replication of the many simple sequences comprising the coding sequence [15–17]. Consequently, the corresponding proteins should also display size and sequence variation and it is possible that they may even represent variable protein antigens that are relevant in evasion of the host immune responses. The genome sequence of M. leprae Unlike M. tuberculosis, there were no alternative genetic approaches to tackle the genetics of the leprosy bacillus and systematic mapping and sequencing was thus initiated at an early stage [18]. At the present time close to 90% of the genome has been sequenced and much of this has been analysed and annotated [19,20]. On inspection of the first cosmid sequence [21] it was apparent that the gene density at ~50–60% was unusually low for a bacterium, and this trend has since been observed throughout the genome although there are regional differences [19]. Now that the complete genome sequence of M. tuberculosis H37Rv is available [1••] detailed comparisons can be undertaken and these are proving to be extremely useful in understanding the basis of the biology of these important pathogens. As first indicated by hybridisation mapping [22], the two genomes show synteny but this appears to be local rather than global. In most instances, the same genes or operons occur in the same order but these are often connected by extended regions that are apparently noncoding or contain pseudogenes in M. leprae. An example of these noncoding regions or pseudogenes is shown in Figure 1, where a segment situated between the rif and str operons of both M. tuberculosis and M. leprae is aligned, the genes identified and differences highlighted. It Comparative mycobacterial genomics Cole is immediately clear that in M. tuberculosis this region is both larger and contains more genes than the corresponding stretch from M. leprae. If one makes the reasonable assumption that both mycobacteria are descended from a common ancestor then they should have similar sized genomes. This raises the possibility that either the leprosy bacillus has lost genes or that M. tuberculosis has acquired additional functions as a result of gene duplication or horizontal transfer. Dotplot analysis of the corresponding nucleotide sequences provides support for both interpretations. 569 Figure 1 end rpsL end lpqP fadE8 echA4 Rv0674 Of the 13 genes found in M. tuberculosis in this region (Figure 1), only two have been conserved in M. leprae and these show between 80–90% identity with their counterparts. Lower levels of sequence identity (~70%) can be seen in regions equivalent to six M. tuberculosis coding sequences; in the leprosy bacillus the corresponding regions contain pseudogenes with multiple frameshifts, small deletions and in-frame stop codons which would abolish gene expression. These data suggest that functional genes were once present in M. leprae but that they have been silenced because their activities were no longer required by an obligate intracellular parasite. Three of these genes (fadE8, echA4, echA5) encode putative β-oxidation enzymes that could degrade unknown fatty acids thereby indicating that M. tuberculosis has the potential to metabolise a larger choice of substrates for growth than M. leprae. Functional information is available for a further three genes: lpqP encodes a lipoprotein that is missing from M. leprae, whereas the other two mmpL5 and mmpS5 encode conserved integral membrane proteins that are confined to mycobacteria and may effect specific tasks such as metabolite transport. As the latter genes belong to a family, it is probable that they arose via a duplication event. The presence of large numbers of pseudogenes in M. leprae probably accounts for many of the phenotypic differences between the leprosy and tubercle bacilli and this is exemplified by their respective responses to the key tuberculocidal agent, isoniazid. In M. tuberculosis, katG encodes catalase-peroxidase, a heme-containing enzyme that mediates the toxic effect of the key tuberculocidal agent, isoniazid [23]. Comparison of the corresponding regions of M. leprae and M. tuberculosis revealed the presence of numerous mutations in the M. leprae gene that abolished its activity [24,25]. This undoubtedly explains why the leprosy bacillus produces no catalase-peroxidase and displays high level resistance to isoniazid. Until the genome sequence of M. leprae is completed, the precise number of genes will remain unknown. Preliminary estimates suggest that the proteome may contain as few as 1600 proteins. The genome of M. leprae is roughly 1.4 Mb smaller than that of M. tuberculosis and its G + C content (57%) is significantly lower than those of all other mycobacterial genomes. Although deletion of coding sequences almost certainly led to some genome shrinkage, echA5 mmpL5 mmpS5 Rv0678 Rv0679c Rv0680c Rv0681 rpsL Current Opinion in Microbiology Dotplot alignment of the nucleotide sequences found in the region downstream of the rif operons of M. leprae (horizontal axis) and M. tuberculosis (vertical axis). The M. leprae sequence is 9,186 bp in length and that of M. tuberculosis 12,140. Both sequences start with the initiation codon of the end gene (endonuclease IV) and finish with the termination codon of rpsL (r-protein S12). M. tuberculosis has an additional lipoprotein gene, lpqP, between end and fadE8, and an operon mmpL5mmpS5 that is not found in M. leprae. By contrast, M. leprae appears to have additional DNA in the region between the residual Rv0681 sequences and rpsL. The figure was generated by DOTTER [27] using a window of 25 bases and greyramp settings of 41–101. Where the degree of relatedness of the two sequences is above the average score, a dot is plotted, and its intensity is proportional to its score. When the dots merge to form a line, this indicates that the DNA sequences are highly related and correspond to genes or pseudogenes. See the text for a detailed explanation of similarity values and further biological interpretation. two other factors may also have contributed to the size difference. First, M. leprae contains very few members of the PE and PPE gene families which account for ~450 kb of the M. tuberculosis chromosome [1••]. Second, traces of far fewer insertion sequences (IS) and bacteriophages have been found in M. leprae than in M. tuberculosis H37Rv, where they contribute over 120 kb. Comparisons such as those outlined above with M. tuberculosis will be extremely informative as they will allow the genes and their functions to be classed into three broad groups: those found in many bacteria, those confined to the genus Mycobacterium and those present only in one or other species. It is already clear that a small number of proteins 570 Genomics have no counterparts in M. tuberculosis and these may confer novel biological properties on M. leprae, and be involved in functions such as neurotropism or nerve damage [26••]. Conclusions Genomics will radically change the way mycobacteriologists tackle problems of pathogenesis by enabling more focused experimental approaches to be adopted and providing greater subject diversity. Comparative genomics will lead to the identification of genes restricted to a given mycobacterium that may play unique biological roles, and serve as sources of specific antigens or potential drug targets. Acknowledgements complete genome sequence of Escherichia coli K-12. Science 1997, 277:1453-1462. 8. Kunst F, Ogasawara N, Moszer I, Albertini AM, Alloni G, Azevedo V, Bertero MG, Bessieres P, Bolotin A, Borchert S et al.: The complete genome sequence of the Gram-positive bacterium Bacillus subtilis. Nature 1997, 390:249-256. 9. •• Sreevatsan S, Pan X, Stockbauer KE, Connell ND, Kreiswirth BN, Whittam TS, Musser JM: Restricted structural gene polymorphism in the Mycobacterium tuberculosis complex indicates evolutionarily recent global dissemination. Proc Natl Acad Sci USA 1997, 94:9869-9874. This highly interesting study presents the results of a rigorous attempt to understand the population genetics of the members of the M. tuberculosis complex and demonstrates that there is very little genetic drift in the set of sequences examined. The findings are interpreted in an evolutionary context and suggest that tuberculosis may be a very recent disease of humans. 10. Kapur V, Whittam TS, Musser J: Is Mycobacterium tuberculosis 15,000 years old? J Infect Dis 1994, 170:1348-1349. I gratefully acknowledge support provided by the Wellcome Trust, the Association Française Raoul Follereau, the World Health Organisation and the Institut Pasteur. 11. Cole ST, Barrell BG: Analysis of the genome of Mycobacterium tuberculosis H37Rv. In Genetics and Tuberculosis (Novartis Foundation Symposium 217). Edited by Chadwick DJ, Cardew G. Chichester: John Wiley; 1998:160-172. References and recommended reading 12. Kolattukudy PE, Fernandes ND, Azad AK, Fitzmaurice AM, Sirakova TD: Biochemistry and molecular genetics of cell-wall lipid biosynthesis in mycobacteria. Mol Microbiol 1997, 24:263-270. Papers of particular interest, published within the annual period of review, have been highlighted as: • of special interest •• of outstanding interest 1. •• Cole ST, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, Gordon SV, Eiglmeier K, Gas S, Barry CE III et al.: Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 1998, 393:537-544. The authors describe the complete genome sequence of the most widely used strain of M. tuberculosis H37Rv and interpret the genomic message using various bioinformatic methods. The main themes addressed are slow growth, dormancy, genetic homogeneity, biosynthesis of the cell envelope and the PE and PPE gene families. The possibility that PE and PPE proteins may be sources of antigenic variation or inhibitors of antigen presentation is discussed. 2. • Brosch R, Gordon SV, Billault A, Garnier T, Eiglmeier K, Soravito C, Barrell BG, Cole ST: Use of a Mycobacterium tuberculosis H37Rv Bacterial Artificial Chromosome (BAC) library for genome mapping, sequencing and comparative genomics. Infect Immun 1998, 66:2221-2229. This paper describes the first successful application of BACs in bacterial genomics and shows how complete genome coverage could be obtained in contrast to the situation with cosmids. The complete genome can be represented by 68 BACs. An application in comparative genomics is presented and a 12.7 kb region of difference between the chromosomes of M. tuberculosis, M. bovis and M. bovis BCG characterised. 3. Hacker J, Blum OG, Muhldorfer I, Tschape H: Pathogenicity islands of virulent bacteria: structure, function and impact. Mol Microbiol 1997, 23:1089-1097. 4. Mahairas GG, Sabo PJ, Hickey MJ, Singh DC, Stover CK: Molecular analysis of genetic differences between Mycobacterium bovis BCG and virulent M. bovis. J Bacteriol 1996, 178:1274-1282. 5. • Pelicic V, Jackson M, Reyrat JM, Jacobs WR Jr, Gicquel B, Guilhot C: Efficient allelic exchange and transposon mutagenesis in Mycobacterium tuberculosis. Proc Natl Acad Sci USA 1997, 94:10955-10960. This publication describes the development of a highly efficient vector system that greatly facilitates gene replacements in mycobacteria. This system is certain to find widespread application. Preliminary results of its use as a vehicle for transposon delivery are presented. 6. • Bardarov S, Kriakov J, Carriere C, Yu S, Vaamonde C, McAdam R, Bloom BR, Hatfull GR, Jacobs JWR: Conditionally replicating mycobacteriophages: a system for transposon delivery to Mycobacterium tuberculosis. Proc Natl Acad Sci USA 1997, 94:10961-10966. The authors describe the construction of a synthetic transposon and its application as an insertional mutagen in M. tuberculosis. The results are interpreted in the context of the genome sequence and suggest that this approach could be very useful for identifying and inactivating nonessential genes. Some bias in the sites of insertion was seen that reflects the locations of the endogenous IS (insertion sequence) elements in the genome. 7. Blattner FR, Plunkett G III, Bloch CA, Perna NT, Burland V, Riley M, Collado-Vides J, Glasner JD, Rode CK, Mayhew GF et al.: The 13. Besra GS, Chaterjee D: Lipids and carbohydrates of Mycobacterium tuberculosis. In Tuberculosis: Pathogenesis, Protection, and Control. Edited by Bloom BR. Washington DC: American Society for Microbiology; 1994:285-306. 14. Wheeler PR, Ratledge C: Metabolism of Mycobacterium tuberculosis. In Tuberculosis: Pathogenesis, Protection, and Control. Edited by Bloon BR. Washington DC: American Society for Microbiology; 1994:353-385. 15. Hermans PWM, van Soolingen D, van Embden JDA: Characterization of a major polymorphic tandem repeat in Mycobacterium tuberculosis and its potential use in the epidemiology of Mycobacterium kansasii and Mycobacterium gordonae. J Bacteriol 1992, 174:4157-4165. 16. Poulet S, Cole ST: Characterisation of the polymorphic GC-rich repetitive sequence (PGRS) present in Mycobacterium tuberculosis. Arch Microbiol 1995, 163:87-95. 17. Poulet S, Cole ST: Repeated DNA sequences in mycobacteria. Arch Microbiol 1995, 163:79-86. 18. Eiglmeier K, Honoré N, Woods SA, Caudron B, Cole ST: Use of an ordered cosmid library to deduce the genomic organisation of Mycobacterium leprae. Mol Microbiol 1993, 7:197-206. 19. Smith DR, Richterich P, Rubenfield M, Rice PW, Butler C, Lee H-M, Kirst S, Gundersen K, Abendschan K, Xu Q et al.: Multiplex sequencing of 1.5 Mb of the Mycobacterium leprae genome. Genome Res 1997, 7:802-819. 20. Fsihi H, De Rossi E, Salazar L, Cantoni R, Labò M, Riccardi G, Takiff HE, Eiglmeier K, Bergh S, Cole ST: Gene arrangement and organisation in a ~76 kilobase fragment encompassing the oriC region of the chromosome of Mycobacterium leprae. Microbiology 1996, 142:3147-3161. 21. Honoré N, Bergh S, Chanteau S, Doucet-Populaire F, Eiglmeier K, Garnier T, Georges C, Launois P, Limpaiboon P, Newton S et al.: Nucleotide sequence of the first cosmid from the Mycobacterium leprae genome project: structure and function of the Rif-Str regions. Mol Microbiol 1993, 7:207-214. 22. Philipp WJ, Poulet S, Eiglmeier K, Pascopella L, Subramanian B, Heym B, Bergh S, Bloom BR, Jacobs WR Jr, Cole ST: An integrated map of the genome of the tubercle bacillus, Mycobacterium tuberculosis H37Rv, and comparison with Mycobacterium leprae. Proc Natl Acad Sci USA 1996, 93:3132-3137. 23. Zhang Y, Heym B, Allen B, Young D, Cole S: The catalaseperoxidase gene and isoniazid resistance of Mycobacterium tuberculosis. Nature 1992, 358:591-593. 24. Eiglmeier K, Fsihi H, Heym B, Cole ST: On the catalase-peroxidase gene, katG, of Mycobacterium leprae and the implications for treatment of leprosy with isoniazid. FEMS Microbiol Lett 1997, 149:273-278. Comparative mycobacterial genomics Cole 25. Nakata N, Matsuoaka M, Kashiwabara Y, Okada N, Sasakawa C: Nucleotide sequence of the Mycobacterium leprae katG region. J Bacteriol 1997, 179:3053-3057. 26. Rambukkana A, Salzer JL, Yurchenco PD, Tuomanen EI: Neural •• targeting of Mycobacterium leprae mediated by the G domain of the laminin-a2 chain. Cell 1997, 88:811-821. This is the most significant publication in the past decade addressing the tropism and mechanism of cell invasion of M. leprae. The authors demonstrate that the bacillus interacts with the G domain of the laminin 571 alpha 2 chain and that this is necessary and sufficient for adherence to Schwann cells. This protein probably acts as a tissue-restricted bridging molecule and would explain why the leprosy bacillus is found in peripheral nerves and muscle cells. It should now be possible to identify the ligand on the bacterial surface that interacts with the G domain. 27. Sonnhammer ELL, Durbin R: A dot-matrix program with dynamic threshhold control suitable for genomic DNA and protein sequence analysis. Gene 1995, 167:GC1-10.