* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download H +
Protein moonlighting wikipedia , lookup
Membrane potential wikipedia , lookup
Phosphorylation wikipedia , lookup
SNARE (protein) wikipedia , lookup
Mechanosensitive channels wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
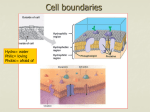
Magnesium transporter wikipedia , lookup
Ethanol-induced non-lamellar phases in phospholipids wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Signal transduction wikipedia , lookup
Theories of general anaesthetic action wikipedia , lookup
Lipid bilayer wikipedia , lookup
Cell nucleus wikipedia , lookup
Cell membrane wikipedia , lookup
Western blot wikipedia , lookup
Model lipid bilayer wikipedia , lookup
1 Cell biology 2014 (revised 21/1 -14), Note Lecture 2 handout. Ester bond Lecture 2: Chapter 10 Chapter 11 Alberts et al 5th edition 617-626 628-636 651-664 Chapter 12 A lot of reading! Focus on principles and topics highlighted in the lecture synopsis 695-699 704-710 Cell Biology interactive media ”video” or ”interactive” Membranes are primary built from phospholipids The major phospholipid: Lipid bilayer 5 -8 nm thick Variable Phosphate Glycerol Fatty acid Glycerides (acylglycerols): esters formed from glycerol and fatty acids Fatty acid Biological membranes are lipid bilayers primary composed of amphipathic phospholipids Hydrophilic head Hydrophobic tails Phosphoglyceride 2 Packing of amphipathic lipids in water - Wedge-shaped lipids form micelles in water H2O is a dipole Red: negative Blue: positive - Cylinder-shaped lipids form bilayers, followed by liposome formation 3 Amphipathic lipids will spontaneously form structures that eliminate the exposure of hydrophobic parts to water Movement of individual lipids within the bilayer Rotational and lateral movement (frequent) Phospholipids can freely and rapidly (mm/s) diffuse within the monolayer Flip-flop (rare) Spontaneous movements between the two monolayers are rare The lipid bilayer is a two-dimensional fluid Similar viscosity as olive oil video 01.2 crawling_amoeba.mov; 13.5 phagocytosis .mov 4 Fatty acid length affects membrane fluidity C=O C=O C=O C=O CH2 CH2 van der Waals CH2 CH2 CH2 CH2 van der Waals CH2 CH2 CH2 CH2 van der Waals Long fatty acid tails CH2 CH2 Short fatty acid tails Weak interactions Strong interactions High fluidity Low fluidity Long aliphatic carbon chains promote van der Waals interactions decreased membrane fluidity 5 Fatty acid saturation affects membrane fluidity An unsaturated fatty acid has a kink Phospholipids containing only saturated fatty acids C=O C=O CH2 CH2 CH2 CH CH2 CH2 CH2 CH Phospholipids containing a unsaturated fatty acid CH2 CH2 Unsaturation's results in steric hindrance decreased van der Waals interactions increased membrane fluidity 6 Effect of lipid composition on membrane fluidity - Membrane thickness - Interactions between fatty acid chains - Membrane fluidity Shorter fatty acid chains and an increased degree of unsaturation make a thinner and more fluid lipid bilayer 7 Anim. 09.1-laser_tweezer; Video 10.1- membrane_fluidity Lipid rafts - clusters of strongly interacting lipids < 100 nm The phospholipid sphingomyelin have long saturated fatty acid tails strong van der Waals interactions Formation of a more static lipid environment Lipid rafts are micro-domains of phospholipids with low fluidity 8 Asymmetry of the plasma membrane Outer monolayer Inner monolayer (facing the cytosol) Phosphatidylcholine Phosphatidylethanolamine Phosphatidylserine Lipid raft former Sphingomyelin 50 40 30 20 10 0 10 20 30 40 50 Percentage of membrane lipids Extracellular space - - molecular_models 10.2-lipids.mov - Phosphatidylinositol, important for cell signaling 9 Different types of membrane proteins Peripheral Integral 3. 1. b-barrel Multi-pass a-helix Single-pass a-helix Mono-topic protein Integral membrane proteins are not tossed into the membrane randomly, but have a specific topology 2. Associated to 1. Lipid 2. Integral protein 3. Glycolipid 10 Dynamics of membrane proteins Rapid movement of proteins within the lipid bilayer Original fluid mosaic model (Singer& Nicolson 1972) Lipid raft Lipid micro-domain (Simons & Ikonen 1997) ~20 % of the 11 plasma membrane Membrane permeability of different molecules • Hydrophobic molecules CO2 O2 Benzene • Small uncharged polar molecules H2O Ethanol • Large uncharged polar molecules H • Charged molecules H + N R C - Glucose O H+ C H H Amino acids O Na+ ClIons 12 Two types of transmembrane transport proteins Carrier Proteins Channel proteins From above Creates a hydrophilic channel Binds a “passenger” at one side of membrane and deliver through the lipid bilayer that is selective for a particular solute it to the other side 13 Ion channels • Most channel proteins are involved in ion transport over the membrane and are therefore called ion channels • Ion channels are regulated and ion specific Closed Ion Open Ion Ion Ion A Ion B Ion A 14 Mechanisms behind membrane transport Concentration gradient Simple diffusion Facilitated specific diffusion Energy independent (down-hill) Active transport Energy dependent (up-hill) 15 Different types of active membrane transport Transport of molecules against a concentration gradient requires energy. Cells uses two distinct strategies. ATP-driven pumps ATP Coupled transporters (symporters) ADP + P “Up-hill” transport coupled directly to hydrolysis of ATP “Up-hill” transport of molecule coupled to “down-hill” transport of molecule . The “down-hill” gradient depends on a ATP-driven pump 16 Example of active transport - Na+/K+ pump Na+ 145 mM K+ 5 mM 1. 2. P ++ Na Na Na+ P ++ Na Na Na+ Na+ 10 mM ATP ADP K+ 140 mM 3. KK++ 4. 1 cycle 10 milliseconds P KK++ Anim. 11.2-carrier_proteins , Anim. 11.1-Na_K_pump 17 Using concentration gradients of Na+ and K+ 18 1. Active transport of Na+ and K+ creates concentration gradients 2. The Na+ gradient provides the energy for “up-hill transport” 3. Coupled transport of sucrose into the cytosol K+ Na+ Na+ Na+ Na+ Na+ Na+ 1. K+ K+ K+ Na+ 2. Glucose 3. Glucose Glucose Glucose The ATP driving the Na+/K+ pump is the energy source for concentrating sugars and amino acids within cells Example of trans-cellular transport by a symporter Na+ Na+ Intestinal lumen 2. Na+ Na+ Glucose Blood vessels Glucose K+ Glucose Glucose Glucose 1. Na+/K+ pump establish Na+ gradient 2. Active transport: Na+ driven 3. glucose symport (“cotransporter”) 3. Passive transport: facilitated Glucose “specific” diffusion of glucose to blood Anim. 11.3-glucose_uptake Glucose K+ Na+ K+ K+ ATP 1. K+ Na+ Na+ Na+ Na+ 19 Compartments/organelles of eukaryotic cells Compartment Main function Cytosol Protein synthesis, metabolism Nucleus DNA & RNA synthesis Mitochondrion ATP production Endoplasmic reticulum (ER) Lipid synthesis, synthesis of proteins that enters the secretory pathway Golgi Sorting and packaging for delivery to cell surface or lysosome Lysosome Protein degradation 20 The nucleus – the instruction book of the cell Nuclear processes: 1. DNA replication 2. Transcription mRNA, rRNA and tRNA 3. Ribosome subunit assembly 1. 2. rRNA + 3. proteins 3-10 mm Nuclear pore 21 One reason for a nucleus in eukaryotes Prokaryote Eukaryote Transcription Transcription mRNA processing Translation Translation In eukaryotes mRNA has to be processed prior to initiation of translation, which requires spatial separation of transcription 22 and translation (Note cloning of an ORF cDNA synthesis) Transport in and out of the nucleus 1. Transcription mRNA 1. tRNA Nuclear Nuclear rRNA pore pore 2. 2. DNA replication Protein synthesis in the cytosol 23 The nuclear pore complex (NPC) A typical cell contains 3000-4000 nuclear pore complexes 120 nm Outer nuclear membrane Inner nuclear membrane Annular subunit; the gatekeeper 24 Proteins less than 60 kDa can diffuse ”freely” between cytosol and nucleus Nuclear import of proteins (>60kD) Nuclear Localization Sequence (NLS) = sequence in a protein that mediates nuclear uptake N NLS N C NLS C N N N L N N L S C Could be localized anywhere in the protein NLS C S C Even distant apart in the primary structure of the protein Which becomes adjacent in the folded protein 25 The process of facilitated nuclear protein import 1. NLS Nuclear import receptor (importin) NLS 2. 3. NLS 1. Association of target protein and nuclear import receptor in the cytosol 2. Binding to the nuclear pore complex mediated by the nuclear import receptor 3. ”Walking” through the gate-keepers of the pore NLS 4. 4. Dissociation of target protein and nuclear import receptor inside the nucleus 26 The nuclear import cycle Cytosol NLS Nucleus 1. Importin NLS Importin Importin NLS GDP Ran <60 kDa 4. GTP Ran 2. NLS GDP Importin Ran GTP Importin Ran +Pi GTP Importin Ran 3. 27 The driving forces behind nuclear import NLS NLS Cytosol Nucleus Importin NLS Importin NLS GDP Ran Importin GDP Ran <60 kDa GDP Energy cost! GTP Importin Ran Video 02.3-brownian_motion.mov GTP GTP Ran GTP Importin Ran 28 Directionality in nuclear import – the Ran cycle Guanine-nucleotide Exchange Factor (GEF) GDP << GTP GDP G protein GTP G protein GTPase Activating Protein (GAP) GDP Ran GDP Ran Pi Ran-GEF Ran-GAP GTP Ran GTP Ran 29 Cytosol Nucleus Nuclear export Nuclear export of proteins is mediated by an intrinsic Nuclear Export Signal (NES). Proteins with NES include: NES NLS NES Small protein that should not be nuclear Protein that shuttle between cytosol and nucleus Video 12.2-nuclear_import.mov Export of mRNA is dependent on successful splicing N E S Proteins responsible for splicing Splicing; removal of introns from mRNA N ES Spliced mRNA ready for nuclear export 30