* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Autophagy_Marc
SNARE (protein) wikipedia , lookup
Biochemical switches in the cell cycle wikipedia , lookup
Protein moonlighting wikipedia , lookup
Extracellular matrix wikipedia , lookup
Cell membrane wikipedia , lookup
Cell nucleus wikipedia , lookup
Cytokinesis wikipedia , lookup
Cytoplasmic streaming wikipedia , lookup
Signal transduction wikipedia , lookup
Programmed cell death wikipedia , lookup
Endomembrane system wikipedia , lookup
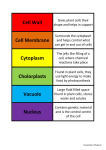
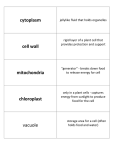
PAINT autophagy annotation Lessons learned Autophagy (or autophagocytosis) (from the Greek auto-, "self" and phagein, "to eat"), is the basic catabolic mechanism that involves cell degradation of unnecessary or dysfunctional cellular components through the actions of lysosomes . The breakdown of cellular components promotes cellular survival during starvation by maintaining cellular energy levels. Autophagy allows the degradation and recycling of cellular components. During this process, targeted cytoplasmic constituents are isolated from the rest of the cell within a double-membraned vesicle known as an autophagosome. The autophagosome then fuses with a lysosome and its cargo is degraded and recycled. There are three different forms of autophagy that are commonly described; macroautophagy, microautophagy and chaperone-mediated autophagy. In the context of disease, autophagy has been seen as an adaptive response to stress which promotes survival, whereas in other cases it appears to promote cell death and morbidity. The 3 types of autophagy The role of autophagy in Parkinson’s disease. Lynch-Day M.A., Mao K., Wang K., Zhao M., Klionsky D.J. Cold Spring Harb Perspect Med. 2012 Apr;2(4):a009357. What’s the origin of the autophagosome / autophagic vacuole ? The autophagosome: origins unknown, biogenesis complex Lamb C.A., Yoshimori T., & Tooze S.A. Nature Reviews Molecular Cell Biology 14, 759–774 (2013) doi:10.1038/nrm3696 Proteins involved in autophagic vacuole assembly are very conserved and well known GO: 0000045 - autophagic vacuole assembly The formation of a double membrane-bounded structure, the autophagosome, that occurs when a specialized membrane sac, called the isolation membrane, starts to enclose a portion of the cytoplasm The emerging role of autophagy in alcoholic liver disease. Ding W.X., Manley S., & Ni H.M. Exp Biol Med (Maywood). 2011 May 1;236(5):546-56. What’s the origin of the autophagosome / autophagic vacuole ? The autophagosome: origins unknown, biogenesis complex Lamb C.A., Yoshimori T., & Tooze S.A. Nature Reviews Molecular Cell Biology 14, 759–774 (2013) doi:10.1038/nrm3696 Pre-autophagosomal structures It is very difficult to discriminate between omegasomes, PAS and autophagosomes. Only recently specific localisation markers such as DFCP1/ZFYVE1 for omegasomes have been identified. New GO terms: PAINT autophagy - summary • 235 primary annotations to autophagy-related terms • 137 primary annotations confirmed for propagation • 98 indirect annotation (IMP, phenotypes) • 48 families annotated • 15 missing families ATGs and autophagic vacuole assembly Protein family PTHR family Assembly Localization Function ATG1 PTHR24348 YES Cytoplasm, PAS Kinase for ATG9 to bind ATG18 ATG2 PTHR13190 YES PAS Localizes ATG18 to omegasome and PAS ATG3 PTHR12866 YES Cytoplasm E2 for ATG8-PI conjugation ATG4 PTHR22624 YES Cytoplasm Cleaves ATG8: reveals C-ter Gly for PI conjugation. Also removes PI ATG5 PTHR13040 YES Cytoplasm, PAS ATG12-ATG5 acts as E3 for ATG8-PI conjugation ATG6/BECN1/VPS30 PTHR12768 YES Endosome, vacuole, PAS VPS34 PI3-kinase complex I, ATG8 conjugation and membrane location ATG7 PTHR10953 YES Cytoplasm, PAS E1 for ATG12-ATG5 and ATG8-PI conjugation ATG8 PTHR10969 YES Omegasome, PAS, vacuole Membrane fusion, autophagosome assembly ATG9 PTHR13038 YES ER, Golgi, PAS (integral) Recruites ATG proteins to the PAS (through ATG18-binding) ATG10 PTHR14957 YES Cytoplasm, PAS E2 for ATG12-ATG5 conjugation ATG101 PTHR13292 YES Cytoplasm, PAS Stabilizes ATG13, protecting it from proteasomal degradation ATG11 PTHR23160 YES PAS Scaffold protein that recruits ATG proteins to the PAS ATG12 PTHR13385 YES Cytoplasm, PAS Ubiquitin-like. ATG12-ATG5 acts as E3 for ATG8-PI conjugation ATG13 PTHR31546 YES Cytoplasm, PAS ATG1 kinase activator ATG14/BAKOR PTHR13664/15157 YES Cytoplasm, PAS VPS34 PI3-kinase complex I. Mediates binding of the complex to the PAS ATG15 PTHR21493 - PAS, vacuole Lipase involved in the lysis of intravacuolar multivesicular body ATG16 PTHR19878 YES Cytoplasm, PAS Stabilizes the ATG5-ATG12 conjugate ATG18/WIPI1/WIPI2 PTHR11227 YES Omegasome/PAS PI(3,5)P2 regulatory complex. ATG9 trafficking to the PAS ATG20 PTHR10555 YES Endosome, PAS Required for proper sorting of the v-SNARE protein SNC1 ATG22 PTHR11360 - Cytoplasm, PAS Vacuolar effluxer, mediates the efflux of amino acids resulting from autophagic degradation Proteins without a PTHR family - 17 distinct autophagy-related proteins do not belong to any PTHR family . - A small part is excluded from families due to highly divergent primary sequences. - The majority were too specific to one species or a small clad of species, so no family could be created. Example: Pichia pastoris ATG30 An example: the ATG1/ULK1 family PTHR24348 Autophagy Response to starvation Phosphorylation Regulation of cell size (?) Regulation of cell growth (?) DROME_ATG1 CEAEL_Unc-51 BPs: High majority of IMPs and IGIs ! Relevance ? Issues with annotations: - Wrong annotations: Disputes: no real stats; but we could dispute at least one annotation per family . Too bad stats are not maintained - it would help identify areas where more annotation guidelines are needed - BP versus regulation of BP: When should we use regulation and when not ? - Over-annotation: IMP/IGI annotations lead to a lot of phenotypic annotations (cell proliferation, cell growth, apoptosis, …). Sometimes it is totally indirect. - Missing annotation: It would be helpful to annotate directly in PAINT - or have a faster turnover between Protein2GO and the GO db. - HTP annotations: Exclusion list for now; is this the best solution ? Even if there are many false positives, the majority should be true positives and are sometimes the only information we have. Issues with propagation: - Issue of species-specific terms: Some terms such as “fungal-type vacuole” are very specific. What's the advantage of having this as a separate term from vacuole ? In animals, the corresponding organelle is the “lysosome”. - Issue of species-specific functions: Some proteins, functions, or processes are very specific to certain species and cannot be propagated. As an example, mitochondrial data should not be propagated in plants, because could be chloroplastic. The case of ATG26 "Atg26 is not involved in autophagy-related pathways in Saccharomyces cerevisiae." Cao Y., Klionsky D.J. Autophagy 3:17-20(2007) … and - PAINT annotation requires to go back to papers most of the time. This is also why it is better to annote related families (i.e. same process). This allows to to: - Save some time. - Make a coherent annotation for the processes. - Annotation of big families is easier when done in teams or based on subject. Part can be left to curators with certain expertise. Example: PTHR24073 (Rab proteins family): over 2700 proteins Function in vesicular transport. Autophagy / Apoptosis The same work has been done by Pascale about Apoptosis which allows some comparisons of annotation between those 2 processes. In general, we found the same issues. However, there are by far less false positive annotations. This is probably due to the fact that there is no «easy» test such as DNA-fragmentation which could be applied to every single mutant. Some suggestions - Work on specific topics, expertise - Grouping for big families. - Direct annotation in PAINT - Faster turnover between Protein2GO and the GO db. - Use a new qualifiers ? (upstream_of and participant_in) - Clean ontology (synonyms, missing relationships,…)