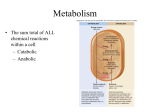
* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download 2005 MCB 3020 Study Objectives, Part 2
Photosynthesis wikipedia , lookup
Biosynthesis wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Epitranscriptome wikipedia , lookup
Metalloprotein wikipedia , lookup
Gene regulatory network wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Expression vector wikipedia , lookup
Deoxyribozyme wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Proteolysis wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Citric acid cycle wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene expression wikipedia , lookup
Biochemistry wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Microbial metabolism wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Electron transport chain wikipedia , lookup
2005 MCB 3020 Study Objectives, Part 2 The Generation of Energy (slides 210-257) • Understand the difference between catabolism and anabolism. • Understand the terms macronutrient, micronutrient, and growth factor. Know that growth factors are organic compounds required in small amounts by some cells; for example, some cells require amino acids, purines, pyrimidines, or vitamins like folate. Recall that sulfanilamide is a growth factor analog that inhibits folate production. Why doesn’t it hurt us? Note that “auxotrophs” (slide 785) are mutants that have a requirement for a growth factor that is not needed by the wildtype. • Understand the classes of microorganisms based on energy source (repeated theme, slide 227). Be prepared to identify examples. Be able to integrate. (eg. nitrification is use of NH3 as an energy source; therefore it is chemolithotrophy.) • No questions on slides 222-224 or 229-239. Know slides 225-228, 240, 242, and 243. (Most enzymes are proteins. A few are ribonucleic acids (These are called “catalytic RNAs” or “ribozymes”). I will not ask about slides 241 and 244-247. • Understand oxidation and reduction. Define electron donor and electron acceptor. Understand the terms primary electron donor (energy source), terminal electron acceptor, intermediate electron carriers. Be prepared to identify examples. Summary: Oxidation is the loss of electrons from a compound. For example, electrons can be lost from glucose to make carbon dioxide (6 CO2). When glucose is oxidized to make CO2 (i.e. when glucose loses electrons), energy is released. Glucose is therefore called the “energy source”. It is also called the “primary electron donor” because it donates electrons to other compounds. The electrons are donated to intermediate electron carriers like NAD+ or FAD, producing NADH or FADH2. The electrons are later donated to other electron acceptors; in this way, the intermediate electron carriers are REOXIDIZED. The LAST compound to receive electrons is called the “terminal electron acceptor”. In aerobic respiration, we use O2 as the terminal electron acceptor. When O2 accepts four electrons, it is reduced to water. In anaerobic respiration, nitrate, sulfate or some compound other than O2 serves as the terminal electron acceptor. Energy Generation and Glycolysis (slides 258-304) • Define oxidant, reductant, and intermediate electron carrier. Know that NAD+ and FAD are INTERMEDIATE electron carriers. Know that NADPH is used mostly in biosynthesis. • Describe how organisms make ATP from glucose in fermentation (slides 277 and 286). Understand that cells derive energy for growth from the OXIDATION of energy sources. This is the ultimate purpose of fermentation and respiration. The energy is saved in the form of highenergy chemical bonds like the phosphoanhydride bond of ATP. • Know that ATP is the energy currency of the cell. You do not need to know the other high energy compounds. • Understand the DETAILS of the differences between fermentation and respiration. • Know the following differences between substrate level and oxidative phosphorylation: Substrate level phosphorylation is ATP (or GTP) synthesis driven by a high energy compound and NOT involving proton motive force. It occurs in the cytoplasm. Oxidative phosphorylation is ATP synthesis using proton motive force and a transmembrane ATP synthase. The proton motive force is generated during electron transfer through intermediate electron carriers in a membrane. Note that a membrane is required for oxidative phosphorylation (also called electron transport phosphorylation). Also note that in respiration ATP production is mainly by oxidative phosphorylation, but that substrate level phosphorylation (in glycolysis and the TCA cycle) also 5 contributes to the number of ATPs produced during aerobic respiration. Prepare for an integrative question comparing substrate level phosphorylation and oxidative phosphorylation. • Understand glycolysis. I will not ask you to memorize the specific steps or intermediates, but I would like you to know the overall reaction (slide 286); that glucose becomes OXIDIZED and that the electrons are transferred to NAD+ making NADH (an intermediate electron carrier); that ATP is produced by substrate level phosphorylation; the net number of ATPs produced (2); that pyruvate is the product of glycolysis; and that glycolysis occurs in the cytoplasm. • Understand fermentation. Why must NAD+ be regenerated? How is this done in fermentation? (Answer: by giving the electrons from NADH to a molecule DERIVED FROM THE SUBSTRATE. For example, pyruvate is made from glucose during glycolysis (i.e. derived from the substrate glucose). You can give the electrons directly to pyruvate to make lactate, slide 301. NAD+ is regenerated as one of the products.) Note that ethanol, lactate/lactic acid, acetate, and other organic acids can be products of fermentation. INTEGRATIVE: Which normal flora use fermentation reactions? How does this affect their environment? (The acids produced lower the pH of the environment.) Aerobic Respiration and the TCA cycle (slides 305-348) • How is ATP generated during the aerobic respiration of glucose? Understand how the following interact to make ATP: glycolysis, pyruvate, acetyl CoA, TCA cycle, electron transport, membrane, proton motive force, ATP synthase, and O2. Know the location of each of these processes or components in prokaryotic cells. See slide 314 for a pictorial summary. What is the primary electron donor? What is the terminal electron acceptor? • Understand the general reactions of the TCA cycle (oxidations, dehydrations, decarboxylations, substrate level phosphorylation, synthesis of carbon skeletons for various biosynthetic processes, production of NADH, FADH2, GTP). I will NOT ask about specific steps of the TCA cycle. • Understand the role of the electron transport system in making ATP. Understand how proton motive force is made. Understand chemiosmosis and the role of the proton gradient (proton motive force) in making ATP. Know the general role of electron carriers in electron transport, but I will NOT ask about specific electron carriers, other than NAD and FAD. • Understand the differences between fermentation, aerobic respiration, and anaerobic respiration. How does oxidative phosphorylation differs from substrate level phosphorylation. Anaerobic Respiration, Chemolithotrophy, and Anabolism (slides 349-386) • Define anaerobic respiration. How do cells derive energy (make ATP) from an energy source during anaerobic respiration? How does this differ from fermentation? • Know that nitrate (NO3-), sulfate (SO4-) can serve as terminal electron acceptors in anaerobic respiration. Note that in anaerobic respiration, ATP is made by proton motive force (i.e. using oxidative phosphorylation or electron transport phosphorylation). • Define anaerobic respiration, chemolithotrophy, autotrophy, heterotrophy, and phototrophy. Know that H2, H2S (hydrogen sulfide), and NH3 (ammonia) are chemolithotrophic substrates. • Know that ATP & reductant (NADPH) are required for many biosynthetic/anabolic processes. • Understand gluconeogenesis and what it is used for. • Know that pentoses for DNA and RNA synthesis are made from hexoses in the pentose phosphate pathway. NADPH is also made in the pentose phosphate pathway. • Know that purine biosynthesis requires folic acid and NADPH (among other things). Understand how sulfanilamide affects purine biosynthesis and cell growth. Microbial Growth (slides 387-434) • Understand the four phases of microbial growth in a batch culture and what is going on in each 6 phase. Know during which growth phase antibiotics (chemicals produced by microbes to inhibit the growth of competing microbes) are made. Integrative: Recognize that a cell that makes antibiotics probably has resistance to the antibiotic. We studied resistance mechanisms later.) NO questions on measuring cell growth. • Be able to define the terms mesophile, thermophile, hyperthermophile, acidophile, halophile, aerobe, facultative anaerobe, strict (obligate) anaerobe. Know where each of these organisms might live. Be prepared for an integrative question. • Know the toxic forms of oxygen, but NOT the detoxifying enzymes. Recall that some toxic forms of oxygen occur in the lysosome of macrophages. Recall from a later lecture that some pathogens can defend themselves against toxic reactive forms of oxygen in the lysosome. What two defense mechanisms do they use? (For answer, see slide 1155.) Replication (slides 435-481) • Understand the terms genetics, heredity, replication, transcription, translation, gene, genome, and the central dogma of molecular biology. • Know that the typical bacterial genome includes one circular DOUBLE-STRANDED DNA chromosome and sometimes plasmids. Understand the structure of a typical virus genome. Know that a typical eukaryote has linear chromosomes. There will be NO questions on the numbers of genes in genomes. • Understand the DETAILS of DNA structure. (See study guide on slide 482.) Be able to contrast this with RNA structure. Note that the “origin” is the part of the DNA where replication starts (slide 456). You do NOT need to recognize the chemical structures. • Explain how DNA replicated (copied) in prokaryotes. Know the names and functions of the six proteins involved in DNA replication. Note that complementary base pairing of G-to-C and Ato-T results from (noncovalent) hydrogen bonding between the complementary bases. • Understand that telomeres are made by telomerase and that they solve the problem of RNA primers at ends of linear eukaryotic chromosomes. NO other telomerase details. Transcription (slides 483-532) • Know the functions of the 3 major RNAs (slide 486). • Understand the DETAILS of RNA structure. (slides 487-491) (Integrative) Know what RNA stem loops are and where they can be found. Complementary base pairing in RNA stem loops is due to what kind of bond? • What are operons and polycistronic messages? Do operons occur in eukaryotes? Understand the DETAILS of the differences between eukaryote and prokaryote transcription. Know that introns, capping, splicing, and tailing are part of eukaryotic transcription, but NOT prokaryotic transcription. • Understand the RNA polymerase (core versus holoenzyme), the promoter, the role of the sigma factor, the structure of a typical sigma-70 promoter (-35 and –10 consensus sequences), the prokaryotic intrinsic terminators (but not rho-dependent terminators). Know that the Pribnow box is another name for the prokaryotic –10 consensus sequence. • Be sure you can distinguish between components/events that occur in transcription versus translation. For ex., RNA polymerase binds to the promoter in transcription; the 3’ end of 16S rRNA binds to the Shine-Dalgarno sequence (prokaryotic ribosome-binding site) in translation. Translation (Protein synthesis) (slides 533-573) • Define translation. What covalent bond is formed between amino acids during translation? On a blank sheet of paper, you should be able to describe or draw the events involved in protein synthesis in prokaryotes. What are the roles of the three RNA molecules, ribosomes, amino 7 acids, aminoacyltRNA synthetases, etc.? Define Shine-Dalgarno sequence, codons, start codons, and stop (nonsense) codons, but you do NOT need to memorize any specific sequences or codons. What bonds are involved in base pairing mRNA codons to tRNA anticodons? • Compare & contrast prokaryotic and eukaryotic translation (Slide 556). Note that eukaryotes do NOT have a Shine-Dalgarno sequence. How do eukaryotic ribosomes bind? Know that the prokaryotic ribosome is composed of 30S and 50S subunits that together make a 70S ribosome. Be prepared for an integrative question. Ex., how do we take advantage of the differences between prok. and euk. gene expression? (Think of ribosomal subunit sizes and antibiotics.) • NO questions on degeneracy of the genetic code, universality, codons pairs and families, codon usage bias, or wobble base pairing. Regulation of Gene Expression (slides 574-654) • Define constitutive gene expression, regulated gene expression, gene induction, repression. • Understand the roles of the following in regulation: effectors, co-repressors, inducers, repressor proteins, activator proteins, operator site, activator binding site. Note that biosynthetic products (like tryptophan) are often co-repressors that repress the synthesis of biosynthetic proteins. (If you have a lot of tryptophan, you don’t need to make biosynthetic enzymes to make more.) Note that catabolic substrates like lactose are typically inducers. (You want to make the enzymes to degrade lactose as an energy source ONLY if lactose is available.) • Know that binding of effectors to regulatory proteins (repressor or activator proteins) changes the conformation (shape) of the regulatory protein, which affects its ability to bind DNA. • Understand how repressor proteins can mediate either gene repression (eg. arginine binds to the repressor protein and causes the repressor to bind to the operator site on DNA. This prevents RNA polymerase from binding to the promoter) or gene induction (When lactose is absent, the lac repressor binds to DNA and blocks transcription. When lactose is present, lactose binds to the repressor. This causes a conformational change in the protein, and the repressor does not bind DNA.) • Understand how activator proteins work in the lac operon. • What is global regulation? Catabolite repression is a global regulatory system. What is the purpose of catabolite repression? Describe the roles of cyclic AMP, CAP (catabolite activator protein), and glucose in catabolite repression. Know that for maximal expression of the lac operon, lactose must be present, glucose must be absent, and cAMP/CAP must bind to the activator site. • I will NOT ask about the trp attenuation system or two-component regulatory systems. Viruses (slies 655 to 722) • Understand the general properties of viruses (slide 658). Know the genome types (slide 664). Understand the structure and function of capsids (protein coats), viral envelopes / envelope proteins, and packaged proteins. Be able to give specific examples from HIV (not polio and flu). Know that the lipid part of the envelope is derived from the host. Be sure you know the difference between the capsid and envelope proteins. • Know the steps involved in phage (bacterial virus) reproduction. Understand the basics of viral reproductive and be able to compare and contrast general aspects of animal and bacterial virus reproduction (attachment, penetration, gene expression and replication). • Know that the specificity of a virus for the host depends on binding of a specific viral protein to a specific host receptor. Understand the process for HIV. • Understand lysogeny. Define lysogen, prophage, and prophage induction (excision). • Understand the reproductive cycle of HIV and how it relates to treatment. Know the function of the envelope protein, reverse transcriptase, integrase, and protease. What is a retrovirus? Note that the HIV protease cleaves the HIV polyprotein into smaller proteins that are active. 8