* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Slide 1
Cell nucleus wikipedia , lookup
Cytoplasmic streaming wikipedia , lookup
Theories of general anaesthetic action wikipedia , lookup
Cell encapsulation wikipedia , lookup
Cell culture wikipedia , lookup
Cellular differentiation wikipedia , lookup
Cell growth wikipedia , lookup
Organ-on-a-chip wikipedia , lookup
Cytokinesis wikipedia , lookup
Lipid bilayer wikipedia , lookup
Endomembrane system wikipedia , lookup
Cell membrane wikipedia , lookup
Extracellular matrix wikipedia , lookup
Signal transduction wikipedia , lookup
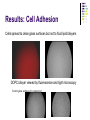
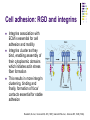
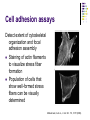
Cell adhesion to supported peptideamphiphile bilayer membranes Badriprasad Ananthanarayanan Advised by Matthew Tirrell PhD Candidacy exam, August 2004 Faculty Committee: Matthew Tirrell Jacob Israelachvili Samir Mitragotri Luc Jaeger Introduction Biomaterials Surface functionalization for increased compatibility and safety Examples Implant materials, e.g. Vascular grafts Seeding with endothelial cells improves graft performance Tissue engineering scaffolds Cells require many signals from matrix to enable proliferation and tissue regrowth Tirrell, M et al., Surface Science, 500, 61 (2000). Biomimetics Engineering biological recognition to create ‘biomimetic’ materials Extra-Cellular Matrix Proteins in the ECM e.g. fibronectin and others provide a structural framework and biochemical signals that control cellular function, e.g. adhesion, growth, differentiation, etc. Creating biomaterials which reproduce these interactions may allow us to direct cell adhesion Tirrell, M et al., Surface Science, 500, 61 (2000). RGD and Integrins • Fibronectin is one of the adhesion-promoting proteins in the ECM • Fibronectin binds to cell-surface receptors known as integrins, transmembrane proteins which regulate a number of cellular processes • The binding site for many integrins in fibronectin is the loop containing the peptide sequence Arg-Gly-Asp (RGD) RGD sites on Fibronectin binding to cell-surface integrins Giancotti, FG, et al., Science, 285, 1028 (1999). Peptide biomaterials: peptide-amphiphiles • Short peptides incorporating the RGD sequence can bind integrins and promote cell adhesion, similar to fibronectin • Using peptides may offer advantages over proteins in terms of convenience, selectivity, and presentation on surfaces Peptide amphiphiles NH2 HN NH HO O O C O C N H O C C O H O N C N C H O H O N C O N C H O H O N C N C H O OH OH C O Hydrophobic ‘tail’ section GRGDSP peptide - headgroup • Peptide headgroups covalently linked to a hydrophobic ‘tail’ segment • Hydrophobic-force driven self-assembly into micelles, vesicles, bilayers, etc. allows us to easily deposit functional molecules on surfaces using selfassembly Self-assembly: Vesicle Fusion • Vesicles are formed from a solution of amphiphiles • When exposed to a hydrophilic surface, vesicles rupture and form bilayer fragments which fuse to form a continuous bilayer on the surface • Clean hydrophobic surfaces are essential for fusion, smaller vesicles are more fusogenic Vesicle Solution on Surface Vesicle Fusion Hydrophilic Substrate Patterned Surfaces Creating Multi-component patterned surfaces Lipid Peptide amphiphile Surfaces: - Glass Barriers: - Proteins, e.g. BSA, deposited by microcontact printing Concentration Gradient: - Microfluidic parallel flow - Fabrication of Microchannels Cell adhesion assays Results: Patterned Bilayers Grid-patterned Stamp Patterned bilayer viewed by Fluorescence Microscopy Results: Cell Adhesion Cells spread to clean glass surfaces but not to fluid lipid bilayers DOPC bilayer viewed by fluorescence and light microscopy Control glass surfaces for comparison: Current work Cell adhesion to bilayers containing peptideamphiphiles Fabrication of microchannels for creating patterned surfaces Effect of Membrane Fluidity on Cell Adhesion SLBs used in our research as a platform for incorporating adhesion-promoting ligands Ease of fabrication by vesicle fusion Inert background: cells show no adhesion to fluid lipid bilayers Retains lateral mobility of membrane components and hence a better mimic of cell membrane Fluidity of SLBs has been used for various purposes Creating micropatterned surfaces Biosensors, etc. Does the fluidity have an effect on cell adhesion? Membrane fluidity in nature Fluid Mosaic model of membranes – proteins and lipids have varying degrees of lateral fluidity Lateral mobility of membrane proteins is an essential step in many signal transduction pathways, e.g. action of soluble hormones, immune recognition, growth, etc. Jacobson, K et al., Science 268, 1441 (1995). Example: Immune Recognition T-cell activation is a critical step in the immune response T-cell activation requires sustained engagement of T-cell receptors by ligands through the ‘immunological synapse’ Formation of this structure involves many receptor-ligand pairs and their transport within the membrane Groves, JT et al., J. Immunol. Meth. 278, 19 (2003). Influence of Ligand Mobility T-cell receptor CD2 and its counter-receptor CD58 (LFA-3) – one of the receptor-ligand pairs involved in T-cell signalling CD58 found in two forms: lipid-anchored (GPI) and transmembrane (TM) lipid-anchored form was mobile, TM form immobile Adhesion of T-cells to GPI-anchored form at lower densities, and adhesion strength also higher Chan, P-Y et al., J. Cell. Bio. 115, 245 (1991). Cell adhesion: RGD and integrins Integrins association with ECM is essential for cell adhesion and motility Integrins cluster as they bind, enabling assembly of their cytoplasmic domains which initiates actin stress fiber formation This results in more integrin clustering, binding and finally, formation of focal contacts essential for stable adhesion Ruoslahti, E et al., Science 238, 491 (1987); Giancotti FG et al., Science 285, 1028 (1999). Effect of RGD clustering The effect of RGD surface density is well known Average ligand spacing of 440 nm for spreading, 140 nm for focal contacts Some evidence that clustering of ligands facilitates cell adhesion (RGD)n-BSA conjugates show equivalent adhesion at much lower RGD densities for higher values of n Synthetic polymer-linked RGD clusters show more efficient adhesion and well-formed stress fibers for nine-member clusters Danilov YN et al., Exp. Cell Res. 182, 186 (1989). Effect of RGD clustering • There is a definite effect of nanoscale clustering of ligands on cell adhesion Maheshwari G et al., J. Cell Sci. 113, 1677 (2000). Simulation of RGD clustering Single-state model – clustering of ligands does not change binding affinity KD No effect observed on ligand clustering other than receptor clustering Two-state model – ligand clustering causes increase in KD – represents activation of receptor in vivo Significantly higher number of receptors bound, especially at low average ligand density This translates into stronger adhesion and better assembly of focal contacts Irvine, DJ et al., Biophys. J. 82, 120 (2002). Effect of bilayer fluidity Spatial organization of ligand has a great effect on cell adhesion, hence fluidity of SLB may have an effect Experimental plan Controlling fluidity in SLBs Characterizing fluidity – FRAP Cell adhesion assays SLB microstructure – formation of domains SLB – controlling fluidity Polymerizable Lipid tails Diacetylenic moieties in lipid tails – can be polymerized by UV irradiation Polymerizable tails can be conjugated to RGD, or lipids with polymerizable tails can be used as a background Control fluidity by varying the degree of polymerization as well as the concentration of polymerizable molecules Tu, RS, PhD thesis, UCSB (2004). SLB – controlling fluidity Quenching mixed-lipid bilayers below the melting temperature e.g. mixed DLPC/DSPC vesicles quenched from 700C to room temperature Results in formation of small lipid domains These domains act as obstacles to lateral diffusion in the bilayer When solid-phase area fraction is very high, diffusion of fluid-phase molecules goes to zero Ratto TV et al., Biophys J. 83, 3380 (2002). Characterizing Fluidity – FRAP Fluorescence Recovery After Photobleaching Fluorescent molecules bleached by high-intensity light source or laser pulse The same light source, highly attenuated, is used to monitor recovery of fluorescence due to diffusion of fluorescent molecules into the bleached area Spot bleaching or Pattern Bleaching Curve fitting gives diffusion constant and mobile fraction Groves, JT et al., Langmuir 17, 5129 (2001). FRAP – analysis Diffusion equation for one species Solution: Gaussian beam intensity profile, circular spot C (r , t ) D 2 C (r , t ) t Curve fitting gives diffusion constant Axelrod, D et al., Biophys J. 16, 1055 (1976); Ratto TV et al., Biophys J. 83, 3380 (2002). FRAP – instrument setup • Light source: High-power lamp or laser • Electromechanical shutter system used to switch between high-intensity beam and fluorescence observation light • PMT vs. Camera – camera allows spatial resolution of intensity, and hence we can monitor background fluorescence recovery, other transport processes • Data analysis by image-analysis software Meyvis, TLK, et al., Pharm. Res. 16, 1153 (1999). Cell adhesion assays Determining adhesion strength Centrifugal detachment assay Sample plate spun in centrifuge, adherent cells counted before and after Low detachment forces applied Hydrodynamic flow Shear stress applied due to flow Many configurations possible Detachment force may depend on cell morphology Garcia, AJ et al., Cell Biochem. Biophys. 39, 61 (2003). Cell adhesion assays Detect extent of cytoskeletal organization and focal adhesion assembly Staining of actin filaments to visualize stress fiber formation Population of cells that show well-formed stress fibers can be visually determined Maheshwari, G et al., J. Cell. Sci. 113, 1677 (2000). Conclusions Constructing supported bilayer membranes incorporating peptide-amphiphiles for cell adhesion Creating micropatterned surfaces for displaying spatially varied ligand concentrations Effect of bilayer fluidity on cell adhesion strength and focal adhesion assembly Design of efficient biomimetic surfaces for analytical or biomedical applications