* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download 33_eukaryote1
Non-coding DNA wikipedia , lookup
Protein adsorption wikipedia , lookup
Molecular evolution wikipedia , lookup
Protein moonlighting wikipedia , lookup
Secreted frizzled-related protein 1 wikipedia , lookup
RNA silencing wikipedia , lookup
RNA interference wikipedia , lookup
Polyadenylation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Transcription factor wikipedia , lookup
Paracrine signalling wikipedia , lookup
Proteolysis wikipedia , lookup
Histone acetylation and deacetylation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
List of types of proteins wikipedia , lookup
Non-coding RNA wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Two-hybrid screening wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Messenger RNA wikipedia , lookup
Gene regulatory network wikipedia , lookup
Epitranscriptome wikipedia , lookup
Gene expression wikipedia , lookup
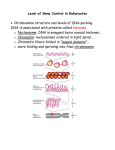
32 Gene regulation in Eukaryotes Lecture Outline 11/28/05 • Gene regulation in eukaryotes – Chromatin remodeling – More kinds of control elements • Promoters, Enhancers, and Silencers • Combinatorial control • Cell-specific transcription – Post transcription gene regulation • mRNA processing • Micro RNAs • Protein degradation – Differentiation and Development • A cascade of transcription regulators • Examples from flowers and fruit flies Gene Regulation in Prokaryotes and Eukarykotes • Prokaryotes – Operons • 27% of E. coli genes • (Housekeeping genes not in operons) – simultaneous transcription and translation • Eukaryotes – No operons, but they still need to coordinate regulation – More kinds of control elements – RNA processing – Chromatin remodeling • Histones must be modified to loosen DNA – Short- and long-term regulation Signal NUCLEUS Chromatin modification: DNA Gene Transcription RNA RNA processing Transport to cytoplasm CYTOPLASM Degradation of mRNA Polypetide Translation Cleavage Chemical modification Transport to cellular destination Active protein Degradation of protein Figure 19.3 Degraded protein DNA Packing 30 nm Nucleosome (b) 30-nm fiber Protein scaffold Loops 300 nm (c) Looped domains (300-nm fiber) 700 nm 1,400 nm Figure 19.2 Scaffold Histone Modification Chromatin changes Transcription RNA processing mRNA Translation degradation Protein processing and degradation DNA double helix Figure 19.4a Histone tails Amino acids available for chemical modification Histone acetylation loosens DNA to allow transcription Unacetylated histones Figure 19.4 b Acetylated histones Densely packed chromatin Activator recruits chromatin remodeling and acetylation proteins RNA Pol Transcription http://cats.med.uvm.edu Review transcription in Eukarkyotes Enhancer (distal control elements) Poly-A signal sequence Proximal control elements Exon Intron Exon Intron Termination region Exon DNA Downstream Upstream Promoter Chromatin changes Transcription Exon Primary RNA 5 transcript (pre-mRNA) Intron mRNA degradation Intron RNA Exon Cleared 3 end of primary transport Coding segment Translation Protein processing and degradation Intron RNA processing: Cap and tail added; introns excised and exons spliced together Transcription RNA processing Exon Poly-A signal mRNA G P P 5 Cap P 5 UTR (untranslated region) Start codon Stop codon Poly-A 3 UTR (untranslated tail region) Many components must be assembled to initiate transcription Those common components are called “General Transcription Factors” There are also many other transcription factors that control transcription of particular genes in particular conditions Control of Galactose metabolism in yeast Two Repressor proteins bind to control region Control of Galactose metabolism in yeast Galactose can bind to repressor complex. Opens activation site to stimulate transcription Enhancers and activators Distal control element Activators Enhancer Promoter Gene TATA box Activator proteins bind to distal control elements. 1 General transcription factors DNA-bending protein Group of Mediator proteins 2 A DNA-bending protein brings the bound activators closer to the promoter. 3 The activators bind to certain general transcription factors and mediator proteins. Fig 19.5 RNA Polymerase II Chromatin changes Transcription RNA processing mRNA degradation RNA Polymerase II Translation Protein processing and degradation Transcription Initiation complex RNA synthesis Transcriptional synergy • Combinations of different enhancers affect the strength of transcription How eukaryotic gene repressors can function: Cell type–specific transcription Enhancer Promoter Albumin gene All cells have the same genes, but only certain genes are expressed in each tissue Control elements Crystallin gene Liver cell nucleus Lens cell nucleus Liver cell Lens cell Albumin gene expressed Fig 19.7 Different set of activator proteins in the two cell types Crystallin gene not expressed Albumin gene not expressed Crystallin gene expressed Long-term control of transcription: methylation • Certain cytosine bases can be methylated, which blocks transcription – Usually CG dinucleotides – Recruits proteins which deacetylate histones, inactivating nearby genes Genomic imprinting: inactivation of maternal or paternal genes Some alleles are tagged by methyl C. Signal NUCLEUS Post-transcription control of gene expression Chromatin modification: DNA Gene Transcription RNA RNA processing Transport to cytoplasm CYTOPLASM Degradation of mRNA Translation Polypetide Active protein Degradation of protein Degraded protein Alternative RNA splicing Chromatin changes Transcription RNA processing mRNA degradation Translation Protein processing and degradation Exons DNA Primary RNA transcript RNA splicing mRNA Fig 19.8 or Micro-RNAs 3 One strand Dicer cuts The micro- dsRNA into of miRNA RNA (miRNA) short segments associates with precursor folds protein. back on itself 1 2 The bound miRNA can basepair with any complementary mRNA 4 5 Prevents gene expresion Chromatin changes Transcription RNA processing mRNA degradation Protein complex Translation Protein processing and degradation Dicer Degradation of mRNA OR miRNA Target mRNA Hydrogen bond Fig 19.9 Blockage of translation Degradation of a protein by a proteasome 1 Chromatin changes Ubiquitin molecules are attached to a protein The ubiquitin-tagged protein is recognized by a proteasome. 2 3 The proteasome cuts the protein into small peptides. Transcription RNA processing Ubiquitin mRNA degradation Translation Proteasome Proteasome and ubiquitin to be recycled Protein processing and degradation Protein to be degraded Fig 19.10 Ubiquinated protein Protein entering a proteasome Protein fragments (peptides) Development Mutant Drosophila with an extra small eye on its antenna Figure 21.1 Determination and differentiation of muscle cells Nucleus myoD Other muscle-specific genes DNA OFF Embryonic precursor cell OFF 1 Determination. Signals from OFF myoD is a “master control” gene: it makes a mRNA transcription factor that can activate other The cell is now protein ireversibly muscle specific genes. MyoD (transcription factor) other cells activate a master regulatory gene, myoD, Myoblast (determined) 2 determined Differentiation. MyoD protein activates other muscle-specific transcription factors, which in turn activate genes for muscle proteins. The embryonic precursor cell is still undifferentiated mRNA MyoD Muscle cell (fully differentiated) Fig 21.10 The cell is now fully differentiated mRNA Another transcription factor mRNA mRNA Myosin, other muscle proteins, and cell-cycle blocking proteins Determination and differentiation of muscle cells Nucleus Master control gene myoD Other muscle-specific genes DNA OFF Embryonic precursor cell 1 Myoblast (determined) 2 Determination. Signals from other cells activate a master regulatory gene, myoD, Differentiation. MyoD protein activates other muscle-specific transcription factors, which in turn activate genes for muscle proteins. OFF OFF mRNA MyoD protein (transcription factor) mRNA MyoD Muscle cell (fully differentiated) Fig 21.10 The cell is now fully differentiated mRNA Another transcription factor The cell is now ireversibly determined to become a muscle cell. mRNA mRNA Myosin, other muscle proteins, and cell-cycle blocking proteins Determination and differentiation of muscle cells Nucleus Master control gene myoD Other muscle-specific genes DNA OFF Embryonic precursor cell OFF 1 Determination. Signals from other cells activate a master regulatory gene, myoD, MyoD protein (transcription factor) Myoblast (determined) 2 OFF mRNA Differentiation. MyoD protein activates other muscle-specific transcription factors, which in turn activate genes for muscle proteins. mRNA MyoD Muscle cell (fully differentiated) Fig 21.10 The cell is now fully differentiated mRNA Another transcription factor The cell is now ireversibly determined mRNA mRNA Myosin, other muscle proteins, and cell-cycle blocking proteins Genetic control of Flower Development Normal Flower “ABC Model” Apetala Class A Pistillata Class B Agamous Class C These genes all code for transcription factors The effect of the bicoid gene, an egg-polarity gene in Drosophila Tail Head T1 T2 T3 A1 A2 A3 A4 A5 A6 A7 A8 Normal larva Tail Tail Figure 21.14 A8 A8 A7 A6 A7 Mutant larva (bicoid) A mutation in bicoid leads to tail structures at both ends (bottom larva). Hierarchy of Gene Activity in Early Drosophila Development Maternal effect genes (egg-polarity genes) Gap genes Pair-rule genes Segment polarity genes Homeotic genes of the embryo Other genes of the embryo Segmentation genes of the embryo Drosophila pattern formation Nurse cells Egg cell 1 Developing egg cell bicoid mRNA 2 Bicoid mRNA in mature unfertilized egg Fertilization Translation of bicoid mRNA 100 µm 3 Bicoid protein in early embryo Anterior end (b) Gradients of bicoid mRNA and bicoid protein in normal egg and early embryo. Homeotic genes Homeotic genes • Regulatory genes that control organ identity • Conserved from flies to mammals