* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Transkriptom a proteom - Univerzita Karlova v Praze
List of types of proteins wikipedia , lookup
Exome sequencing wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Non-coding RNA wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
Gene desert wikipedia , lookup
Non-coding DNA wikipedia , lookup
Molecular evolution wikipedia , lookup
SNP genotyping wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Messenger RNA wikipedia , lookup
Genome evolution wikipedia , lookup
Gene expression profiling wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Epitranscriptome wikipedia , lookup
Gene regulatory network wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Gene expression wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Silencer (genetics) wikipedia , lookup
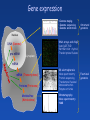
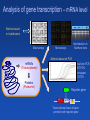
Transcriptome and analysis of gene transcription Gene expression • Genome maping • Genome sequencing • Genome annotations Structural genomics Nucleus DNA (Genome) pre-mRNA Cytoplasm • DNA arrays and chips • (semi) qRT-PCR • Northern blot + hybrid. • Transkriptional fusions mRNA mRNA (Transcriptome) Proteins (Proteome) Metabolites (Metabolome) • 2D electrophoresis Mass spectrometry Protein sequencing • Translational fusional • Immunodetection • Enzyme activities • Chromatography • Mass spectrometry • NMR Functional genomics Transcriptome - set of all mRNAs present in certain cell, tissue, organ, … - mRNA level results from intensity of transcription and mRNA stability Transcriptomics – expression analysis of populations of genes - analysis of differences in expression of gene populations (under different conditions, treatments, developmental stages) Analysis of gene transcription – mRNA level Methods based on hybridization Macroarrays Hybridization on Northern blots Microarrays Methods based on PCR mRNAs (Transcriptome) Real time PCR qRT-PCR; Semiquant. RT-PCR DNA (Genome) Proteins (Proteome) Reporter gene P gene T Transcriptional fusion of gene promotor with reporter gene 1. Transcriptional fusion of promoter with reporter gene encoding glukuronidase or GFP Reporter gene P gen T - easy analysis of the sites of certain gene expression in planta Arabidopsis thaliana Compare with promoter-trap mutagenesis ! qRT-PCR a Semiquantitative RT-PCR original level of template measured as: - PCR product level after certain number of cycles - number of PCR cycles necessary to reach certain product level mRNA isolation Semiquantitative RT-PCR Reverze transcription (oligo T-primer, specific reverze primer) cDNA qRT(real time)-PCR Proper number of cycles has to be determined for semiq. RT-PCR Electrophoretic detection limit qReal Time - PCR Detection of product level – fluorescent probes improve specificity Fluorescent labels: R …reporter Q …quencher D … donor A … acceptor Principle of detection of nucleic acids by hybridization Probe - strand of NA with known sequence used for detection of complementary strand in a mixture of NAs (e.g. transcripts, cDNAs, genomic fragments) Two phases system (): hybridization of complementary single-stranded NA: immobilized (bound on membrane, glass) mobile phase (NA in solution) immobilized probes Arrangement I: (on known positions) mobile, labeled mixture of NA immobilized mixtures Arrangement II: mobile labeled probe Labeled probes for hybridization - labelling by usually by incorporation of labelled nucleotide during NA synthesis Types of labeling – radioactive (most frequently 32P) - fluorescent - digoxygenin, biotin etc. + (followed by detection with a specific antibody) Hybridization on Northern blots RNA isolation Electrophoretic separation Macroarrays Microarrays Blotting = transfer of mRNA from gel onto a membrane Hybridization with labelled probe, detection Hybridization on Northern blots Immobilized phase – analyzed mixture of mRNAs Mobile phase = labeled probe of certain gene (signal = presence of certain transcript + info about the transcript size) x Hybridization on DNA arrays or chips Immobilized phase – multiple probes with known sequences bound on certain places of the solid support Mobile phase = labeled mixture of analyzed NAs (simultaneous detection of presence and quantity of many sequences) DNA arrays and DNA chips - principle Hybridization Fluorescent (RI) signal Fluorescently (RI) labelled analyzed NAs (mobile phase) Pozition Array, chip (imobilized probe) Identity Intensity Amount Terminology: arrays, chips Preparation Support Macroarray (High Density Array) Printing of oligonucleotids or PCR fragments Membrane e.g. glass Microarray Printing of oligonucleotids or PCR fragments Direct synthesis on the support e.g. glass Chip Density [probes/cm2] max. 64 up to 10 4 up to 2.5 *10 5 Arrays Probes Mobile phase (usually labelled cDNA) Imobilized phase (array) cDNA (ESTs) Imobilized probes Genome sequences Oligonucleotides, … Automated preparation of macroarrays contact printing 4.5 mm Comparison of gene expression using differential labelling on arrays Situation I RNA isolation Hybridization Labelling Situation II Alternative approach: independent hybridization and comparison of the results Identification of differentially expressed genes Troubles with hybridization on arrays 1. Non-specific (cross-) hybridizations, background 2. Signal intensity depends also on sequence (differences in efficiency of hybridization) 3. Reproducibility Solutions: • every probe on different positions on the array • several different probes for every gene • Affimetrix chips Oligonucleotide chips from Affymetrix - mutiple probes for every gene (20 pairs), direct synthesis on the chip - probes from 3' end of mRNA (for Eucaryots) - every oligonucleotide in perfectly matching version and with one missmatch mRNA sequence 3‘ 5‘ Pairs of oligonuceotide probes Gene sequence Perfect match Single NT missmatch Perfect match Fluorescens intensity Single NT missmatch Differences in fluorescence intensity between perfect and missmatched oligonucleotide are averaged for all probe pairs Sample labeling for hybridization on Affimetrix chips reverze transcription mRNA AAAAAAAAA cDNA with T7 promoter AAAAAAAAA TTTTTTTTTT- T7 promoter AAAAAAAAA AAAAAAAAA AAAAAAAAA Ligation with promoter in vitro transcription with biotinilated NTP B B B B B B B B B B B B B B fragmentation B B B Sample ready for hybridization with the chip B UUUUUUUUU B B UUUUUUUUU B B B UUUUUUUUU B B B B B B B B B B Biotin-labelled cRNA B B UUUUUUUUU Affymetrix chips - hybridization and result analysis BB B B B B B B B B streptavidin- phycoerythrin binds to biotin BB B B B B B B B B Image analysis detector Emission at 570 nm BB B B B B B B B excitation at 488 nm B Genevestigator https://www.genevestigator.com partially free approach to chip results Selection of: - species - genes - chips (experiments) Affymetrix chip preparation Photolithography