* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download GMO and Biotechnology - Western Washington University
Genomic library wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Essential gene wikipedia , lookup
Oncogenomics wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Non-coding DNA wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Point mutation wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Public health genomics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Genetically modified organism containment and escape wikipedia , lookup
Pathogenomics wikipedia , lookup
Gene expression programming wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genetically modified food wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genomic imprinting wikipedia , lookup
Helitron (biology) wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Ridge (biology) wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Genome (book) wikipedia , lookup
Minimal genome wikipedia , lookup
Genetically modified crops wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genome evolution wikipedia , lookup
Gene expression profiling wikipedia , lookup
Genetic engineering wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
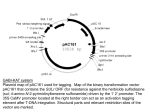
Genetically Modified Plants Biotechnology: underlying science Potential Risks vs.(Potential) Benefits Assigned Reading: Chapter 10.5 Genetically Modified Organisms Types of GMOs? - artificial selection and traditional breeding, - transgenic organisms, - other approaches, - targeted mutagenesis, - gene introgression, - ? Old Science Humans (~30,000 years) Humans (~30 years) Bacteria (eons) Humans (~15 years) Bacteria (eons) Desirable Agronomic Traits (traditional or modern) • Increased yields, more nutritious, quality, etc., • More resistant to pestilence, weeds, water and nutrient deprivations, • Ability to withstand marginal growth conditions, – and thrive in new environmental ranges, • Profit. Traditional Breeding ~25,000 genes ~45,000 genes • technology is not essential, • limited by species boundaries, • all genes/traits are mixed. Introgression …incorporation of genes of one genome into the genome of another cultivar, – standard breeding techniques are laborious (if possible at all), – genomics and related sciences greatly accelerates standard breeding techniques. Genome Era Traditional Breeding Wild tomato Cultivar w/ 1 wild gene replacement Transgenic Plants • based on DNA technology, • single genes/traits can be transferred, • species boundaries are not limiting. How are GMOs generated? ...uses tools of molecular genetics, - i.e. applied bacteria and virus genetics. insert into plant …via biolistics - or - Agrobacterium tumefaceins Biolistics Kalanchoe Stem w/ infection. Agrobacterium tumefaciens Natural soil bacterium that infects plants, hosts: 160 Genera, families: > 60, effect; poor growth, low yield. Nature Ti-Plasmid Transfer-DNA Plant Cells Ti: tumor inducing Plasmid: extrachromosomal DNA evolved for genetic transfer. Agrobacterium Hormone Opines genes genes Lab T-DNA Out: Ti genes, opine genes, In: DNA of choice. Any Gene Selectable Markers, etc T-DNA (Transfer DNA) …with gene of interest, Construct T-DNA bcarotene, - herbicide resistance, etc.. transform, select for agro with T-DNA Agrobacterium infect plant, select for plants with T-DNA Plant chromosome with T-DNA insert. T-DNA (Transfer DNA) selection genes Construct T-DNA virulence genes …gene of interest, bcarotene, - herbicide resistance, etc.. Virulence genes: facilitate Agro infection, T-DNA transfer, • not usually transferred in commercial applications, Selection genes: used to identify transgenics, • usually antibiotic or herbicide resistance, etc. (i.e. only the organisms with the T-DNA live in a selection experiment), Gene of interest: protein coding region, plus a “promoter”. Promoters Control Expression Foreign DNA is common (via nature) in most genomes, Transgenes must be expressed in order to function, Promoters control where, when and how much protein is produced. Gene Structure chromosome (megabases) gene (kilobases) ...ata cgt act atc... ...ttaggttctatc... |||||||||||| ...aatccaagatag... ||| ||| ||| ||| ...tat gca tga tag... promoter specific sequences. protein coding Promoter Specifies Expression General Promoter: all tissues, all the time. Vegetative Promoter: no flower, no fruit expression. Root Promoter: only root expression. Expression = Protein Production Protein and protein functions only present in tissue with active promoter. Tissue Specific Expression Time Specific Expression “Suicide” Promoters, etc. Brief History of Transgenic Organisms • Transgenic E. coli, – not demonstratively dangerous, – demonstratively beneficial (probably). • Transgenic virus, – not demonstratively dangerous, – demonstratively beneficial (probably). • Transgenic plants, – demonstratively dangerous? (not yet), – demonstratively beneficial (?). Potential Risks • Risk of invasion. • Direct nontarget Effects • Indirect nontarget Effects. • New Viral Diseases. • Variability and Unexpected Results. Potential Risks (risk of invasion) • 50,000 invaders in USA the old fashioned ways, – self-sustaining cultivars, • low anticipated risk, – hybridization with (native) neighbors, • transgene introgression, • introgression of domestic cultivar genes with natives has occurred, resulting in negative impacts on native species, – time lags. Direct (nontarget) • Risk to non-target species, – pollinators, – passers-by, • soil ecosystems, – decomposition rates, – carbon cycle, – nitrogen cycle. Indirect (nontarget) • kill weeds = kill species that live “on” or eat the weeds, • bioaccumulation, – nontarget species eat plants, store toxins, – those species are eaten, amassing the toxin, – on up the food chain. Bee on Red Clover. New Viral Diseases • virus resistant plants promote virulent strains, – mutations, – recombination, • heteroencapsulation, – virus move genes from one organism to another, – not presently a risk, but a potential risk. Variability and Unexpected Results • time scale, • numbers, • environmental and cultivar differences, • application, culture and consistency. Other Issues • Economic hegemony of GMP seed producing countries, companies, • Cultural shifts in farming due to the introduction of GMOs, • Potential allergies to genetically modified crops, • The preservation of natural genetic crop-lines, • The lack of an adequate risk assessment methodology to quantify unintended ecological consequences. The Precautionary Principle Biotechnology in General Scenario 1 Scenario 2 Works great Bad Environmental Consequences Increase Carrying Capacity for Humans Human Population Growth Negative impacts on, • select species, • crops, • ecosystems, • etc. Negative impacts on, • select species, • crops, • ecosystems, • etc. Schedule - option - • Monday: Sugar and Genetics – PDF of paper available on WEB page, • print pp. 7 - 15. ….or? 12. (16 pts) In bacterial matings , prophage can be transferred from Hfr to F-. The prophage is auto ma tic all y induced when it enters F- cell s when ther e is no ph age repressor, and the cell is then lysed . Seve ral new Hfr strains of E. coli were independ ently isolated. All were wild type , exc ept for Hfr 1 which was lysogen ic for phage la mbd a. All Hfrs were then mated to a F- strain carrying mutations in the foll owing genes : ara, gal, lys, pro, pyr, rha . The times of f ir st appearanc e of individua l Hfr genes (w il d-type alleles) among the recombinan ts were as foll ows (in minutes): Hfr Marker ara gal his lys pro pyr rha Hfr 1 Hfr2 Hfr3 8 24 No recombinants No recombinants 20 No recombinants No recombinants 60 44 21 73 89 12 4 29 48 40 85 93 84 49 Draw a complete (circular) ma p of the E. coli chro mosome, show ing the distance (in terms of tim e) between each of the markers and the approximate location of the lambda prophage . Show the orientation and location of the F factor (i. e., the arrow). Assume a 100-minute map.