* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download from Chapter 11: Gene Regulation
Epigenetics of neurodegenerative diseases wikipedia , lookup
Essential gene wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
History of genetic engineering wikipedia , lookup
Gene expression programming wikipedia , lookup
Genetic code wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genomic imprinting wikipedia , lookup
Microevolution wikipedia , lookup
Designer baby wikipedia , lookup
Epitranscriptome wikipedia , lookup
Protein moonlighting wikipedia , lookup
Genome evolution wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Point mutation wikipedia , lookup
Primary transcript wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Non-coding RNA wikipedia , lookup
Ridge (biology) wikipedia , lookup
Genome (book) wikipedia , lookup
Minimal genome wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene expression profiling wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
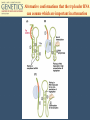
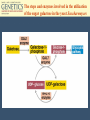
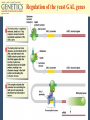
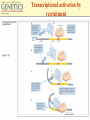
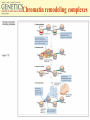
Chapter 11 Molecular Mechanisms of Gene regulation Jones and Bartlett Publishers © 2005 The 4 critical sequences in the lac operon bound by CRP, RNA polymerase, repressor and the ribosome See Figure 11.8. lac operon repression loop. Binding of tryptophan (the co-repressor) activates an inactive repressor into an active form capable of binding to the trp operator site The terminal region of the trp attenuator sequence. Structure of the leader polypeptide in the trp operon The two tandem tryptophans in the leader peptide act as “stalling sequences” in the absence of tryptophan in the cell Alternative conformations that the trp leader RNA can assume which are important in attenuation Other operons with repeated amino acid sequences that act as “stalling sequence” during attenuation Other amino acid operons using attenuation: leucine and isoleucine, TRAP • B. subtilis utilizes a different way of controlling the expression of trp operon. • trp operon is not repressible in B. subtilis. • No aporepressor, no short leader polypeptide sequence. • TRAP: trp RNA-binding attenuation protein. • Each monomer binds a tryptophan molecule. • 11-mer radially-symmetric TRAP is active form. • Binds to regions 1+2 in figure 11.13. Riboswitches • A 5’ untranslated leader mRNA can have a small molecule bind to it to change its conformation between two states: • Antiterminator • Terminator • yitL gene in B. subtilis methionine biosynthesis Riboswitch regulation of transcription termination The genetic map of bacteriophage l Control of transcription in bacteriophage l life cycle by the anti-terminators N and Q proteins, the activator CII protein and the repressor/activator CI protein • cI and cro proteins function as a genetic switch: • cI turns on lysogeny. • cro turns on the lytic cycle. Transcriptional regulation in eukaryotes • Many housekeeping genes are expressed constitutively. • Levels of expression can be 2-10 fold different in induced vs. uninduced genes. • Galactose metabolism in yeast is a good case study. GAL pathway in yeast • Extensively studied system. • Eight genes of this pathway are involved in galactose metabolism and regulation. • Five genes (GAL2, MEL1, GAL4, GAL80, and GAL5) are on different chromosomes. • Three genes are clustered on chromosome II (GAL7, GAL10, GAL1). The steps and enzymes involved in the utilization of the sugar galactose in the yeast Saccharomyces Function of the yeast GAL genes GAL3, GAL4, and GAL80 are regulatory genes. The transcriptional orientation of the 3 genes coding for enzymes important in galactose utilization in Saccharomyces Their synthesis is regulated by the transcription activator Gal4 protein. Regulation of the yeast GAL genes Transcriptional Activator Proteins Two broad types of transcriptional activator proteins: helix-turn-helix motif, so it fits into grooves of DNA zinc finger motif, because zinc ions are involved. Transcriptional Activator Proteins Transcriptional Enhancers (Silencers – opposite of enhancers) Deletion scanning Transcriptional Enhancers (Silencers – opposite of enhancers) Deletion scanning Transcriptional activation by recruitment Chromatin remodeling complexes Alternative promoters