* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Scenario 2 - people.vcu.edu
Polycomb Group Proteins and Cancer wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Epigenetics wikipedia , lookup
DNA vaccination wikipedia , lookup
Molecular cloning wikipedia , lookup
Gene expression profiling wikipedia , lookup
DNA methylation wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Neocentromere wikipedia , lookup
Gene expression programming wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Point mutation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
DNA supercoil wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Non-coding DNA wikipedia , lookup
Genome editing wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Microevolution wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Nutriepigenomics wikipedia , lookup
History of genetic engineering wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Designer baby wikipedia , lookup
Epigenomics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
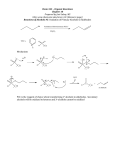
Scenario 2 Anotation: Gene of which something known What follows is a simulation of an orf page in the proposed graphical interface. The interface does not yet exist. As you go through the simulation please consider what capabilities you would want to serve your research and annotation interests. A narrative to help you go through the simulation appears in a red-bordered box, such as the one below. To begin: 1. Click on Slide Show, (on the upper toolbar) 2. Click View Show 3. Click Continue button Continue Main Menu Annotate Options History Anabaena PCC 7120: alr1152 Replicon: Chromosome Coordinates: 1356962 (start-ATG) -> 1357843 (stop) Length = 293 amino acids Human A Function: solitary Type II DNA methyltransferase (GGCC-specific) Classification: Type II alpha (N4) Experiment Human A Activity: Modification: N4-methylcytosine (inner cytosine of GGCC) Protects against: HaeIII In vivo activity: exists Experiment Experiment Experiment A Expression: no data Syny6803:sll1330: (click to expand) Experiment Strand: Direct Gene name(s): dmtB, avaVIIM, dmnB Mutant: Knockout loses GGCC methylation. Otherwise no known difference from wild-type Experiment This is an example of an orf that has substantial human annotation. Cyanobacterial orthologs: Syny6803.C:sll0729 NostPunc.352:003 The source of each assertion is given at the right. Press Continue . Continue A A A A Main Menu Annotate Options History Anabaena PCC 7120: alr1152 Replicon: Chromosome Coordinates: 1356962 (start-ATG) -> 1357843 (stop) Length = 293 amino acids Human A Function: solitary Type II DNA methyltransferase (GGCC-specific) Classification: Type II alpha (N4) Experiment Human A Activity: Modification: N4-methylcytosine (inner cytosine of GGCC) Protects against: HaeIII In vivo activity: exists Experiment Experiment Experiment A Expression: no data Syny6803:sll1330: (click to expand) Experiment Strand: Direct Gene name(s): dmtB, avaVIIM, dmnB Mutant: Knockout loses GGCC methylation. Otherwise no known difference from wild-type Mouse over* to Experiment to the right of the Function Cyanobacterial orthologs: Syny6803.C:sll0729 NostPunc.352:003 specification line to see where the assertion comes from. *In real life, information will pop up when you mouse to an informative position. In this simulation you’ll also need to click. Experiment A A A A Main Menu Annotate Options History Anabaena PCC 7120: alr1152 Replicon: Chromosome Coordinates: 1356962 (start-ATG) -> 1357843 (stop) Length = 293 amino acids Strand: Direct Gene name(s): dmtB, avaVIIM, dmnB Human A IDA: Matveyev et al (2001) Function: solitary Type II DNA methyltransferase (GGCC-specific) Classification: Type II alpha (N4) Experiment Human A Activity: Modification: N4-methylcytosine (inner cytosine of GGCC) Protects against: HaeIII In vivo activity: exists Experiment Experiment Experiment A Expression: no data Syny6803:sll1330: (click to expand) Experiment Mutant: Knockout loses GGCC methylation. Otherwise no known difference from wild-type IDA (Inferred from Direct Assay) is a standard Cyanobacterial orthologs: Syny6803.C:sll0729 NostPunc.352:003 Gene Ontology Consortium descriptor, useful for searching (see Scenario 3 for more). Evidently there’s some hard evidence for the assertion. To learn more, click on Experiment. Experiment A A A A Back to Annotation page Nucleic Acids Research (2001) 151:1491-1506 Full text article at nar.oupjournals.orf PubMed Central access FREE full text articles DNA methyltransferases of the cyanobacterium Anabaena PCC 7120 Andrey Matveyev, Kathryn T Young, Andrew Meng, and Jeff Elhai Dept. of Biology, University of Richmond, Richmond VA USA From the characterization of enzyme activities and the analysis of genomic sequences, the complement of DNA methyltransferases (MTases) possessed by the cyanobacterium Anabaena PCC 7120 has been deduced. Anabaena has nine DNA MTases. Four are associated with Type II restriction enzymes (AvaI, AvaII, AvaIII and the newly recognized inactive AvaIV), and five are not. Of the latter, four may be classified as solitary MTases, those whose function lies outside of a restriction/modification system. The group is defined here based on biochemical and genetic characteristics. The four solitary MTases, DmtA/M.AvaVI, DmtB/M.AvaVII, DmtC/M.AvaVIII and DmtD/M.AvaIX, methylate at GATC, GGCC, CGATCG and rCCGGy, respectively. DmtB methylates cytosines at the N4 position, but its sequence is more similar to N6-adenine MTases than to cytosine-specific enzymes, indicating that it may have evolved from the former. The solitary MTases,are appear to betoofreferences ancient origin Annotations linked at within cyanobacteria, while the restriction MTases appear to have arrived by recent horizontal transfer as did five now inactive Type I PubMed and from there (sometimes) to restriction systems. One MTase, M.AvaV, cannot reliably be classified as either a solitary or full length articles. Click on button restriction MTase. It is structurally unusualtoand along with a few proteins of prokaryotic and return annotation page.class of MTases distinct from all previously described. eukaryotic origintodefines a structural Main Menu Annotate Options History Anabaena PCC 7120: alr1152 Replicon: Chromosome Coordinates: 1356962 (start-ATG) -> 1357843 (stop) Length = 293 amino acids Human A Experiment Human A Experiment Experiment Experiment A Strand: Direct Gene name(s): dmtB, avaVIIM, dmnB Function: solitary Type II DNA methyltransferase (GGCC-specific) Classification: Type II alpha (N4) Activity: Modification: (inner cytosine of GGCC) The human N4-methylcytosine annotations are propagated Protects against: HaeIII throughout all cyanobacterial orthologs. Click In vivo activity: exists on Syny6803.C:sll0729 to see how this works. Expression: no data Syny6803:sll1330: (click to expand) Mutant: Knockout loses GGCC methylation. Otherwise no known difference from wild-type Cyanobacterial orthologs: Syny6803.C:sll0729 NostPunc.352:003 A A A Experiment Experiment A Main Menu Annotate Options History Synechocystis PCC 6803: sll0729 Replicon: Chromosome Coordinates: 3424667 (stop) <- 3425524 (start) Length = 285 amino acids System Strand: Complement Function: probable DNA methyltransferase System Anab7120.C:alr1152: solitary Type II DNA methyltransferase (GGCC-specific) Experiment Motifs: pfm:MethyltransfD12: D12 class N6 adenine-specific DNA methyltransferase ps:N6_MTASE: N-6 Adenine-specific DNA methylases signature. System System Expression: (click to expand) Mutant: none Anab7120.C:alr1152: Knockout loses GGCC methylation. Otherwise no known difference from wild-type Experiment Experiment Cyanobacterial orthologs: Anab7120.C:alr1152 NostPunc.352:003 The annotation for the Anabaena gene appears automically on the annotation page of its orthologs. This gives a hint to visitors to the Synechocystis page that the system’s suggestion that Sll0729 is an adenine methyltransferase may be [and is] wrong. A End A Scenario 2 Anotation: Gene of which something known Summary • Each assertion is linked to a justification. • Justifying references are made available on-line. • Annotations of an orf in one organism are propagated through orthologs to other cyanobacteria. The integration of annotation across all cyanobacterial genomes makes possible the ideal of consideration of each gene by a person expert in the field. This is achieved by dividing genes vertically, by function, rather than horizontally, by species.