* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Activation sites and enhancer proteins
Bottromycin wikipedia , lookup
Protein moonlighting wikipedia , lookup
Biochemistry wikipedia , lookup
Gene expression profiling wikipedia , lookup
Non-coding DNA wikipedia , lookup
Histone acetylation and deacetylation wikipedia , lookup
Expanded genetic code wikipedia , lookup
RNA interference wikipedia , lookup
Transcription factor wikipedia , lookup
List of types of proteins wikipedia , lookup
Molecular evolution wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Two-hybrid screening wikipedia , lookup
RNA silencing wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genetic code wikipedia , lookup
Point mutation wikipedia , lookup
Gene regulatory network wikipedia , lookup
Biosynthesis wikipedia , lookup
Polyadenylation wikipedia , lookup
Messenger RNA wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Non-coding RNA wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Gene expression wikipedia , lookup
Epitranscriptome wikipedia , lookup
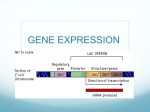
The “Central Dogma” Overview Of DNA DNA Transcription mRNA Translation proteins Enzymes Replication Structure Movement Hormones Gas exchange Amino Acid Storage Protein Synthesis How does DNA control the structure and function of the cell? it makes proteins! – Structure: collagen, elastin, keratin – Enzymes: catalase, amylase, sucrase, etc – Hormones: insulin, glucagon, etc – Amino acid storage: albumin, ovalbumin, etc What is similar about protein synthesis in prokaryotes and eukaryotes? What is different? Protein Synthesis! Transcription http://www.johnkyrk.com/DNAtranscri ption.html Translation http://www.johnkyrk.com/DNAtranslati on.html Notas – From Gene to Protein Metabolism teaches us about genes Metabolic defects caused by non-functional enzyme Studying metabolic diseases suggested that genes specified proteins – PKY – Alkaptonuria (black urine) Genes dictate the phenotype 1 gene – 1 enzyme hypothesis Beadle and Tatum – 1941 – Compared different nutritional mutants of bread mold, Neurospora – Created mutations by X-ray treatments Xrays break DNA) – Wild type grows on “minimal” media (sugar) – Mutants require different amino acids because each mutant lacks a certain enzyme needed to produce a certain amino acid – Conclusion: Broken gene = non-functional enzyme 1 gene – 1 enzyme hypothesis Beadle and Tatum – 1941 Problems with: –One gene – one enzyme not all proteins are enzymes, and they’re coded by genes too –One gene – one protein many proteins consist of several polypeptide, and each polypeptide has it’s own gene One gene – one polypeptide? Defining a gene… “Defining a gene is problematic because small genes can be difficult to detect, one gene can code for several protein products, some genes code only for RNA, two genes can overlap, and there are many other complications.” – Elizabeth Pennisi, Science 2003 How would YOU define a gene in your own words? From nucleus to cytoplasm… Where are the genes? in DNA on chromosomes in the nucleus Where are proteins synthesized? on ribosomes (free or on the ER) in the cytoplasm How does the information get from the nucleus to the cytoplasm? mRNA is made in the nucleus and can travel into the cytoplasm to the ribosomes deoxyribose A-T, C-G Double ribose T-A, A-U, C-G Single Transcription! http://highered.mcgrawhill.com/sites/0072507470/student_vie w0/chapter3/animation__mrna_synthe sis__transcription___quiz_2_.html http://www.johnkyrk.com/DNAtranscri ption.html Transcription Basics Initiation 1. RNA polymerase binds to promoter sequence on DNA 1 2 3 2 Elongation 3 where to start reading = Promoter (initiation site) which strand to read = template strand direction on DNA = reads 3’5 builds 5’ 3’ RNA polymerase unwinds DNA ~20 bp at a time Reads DNA 3’ 5’ Builds RNA 5’ 3’ No proofreading, about 1 error/105 bases Many copies, short life, no problem Termination RNA polymerase stops at termination sequence mRNA leaves nucleus through pores promoter Terminator Transcription RNA polymerase Template Strand Initiation Elongation mRNA Termination Completed mRNA transcript RNA Processing or Editing 5’ cap – protection – targets mRNA for ribosome Poly-A tail – protection – leads mRNA out of nucleus Spliceosome – composed of snRNPs (small nuclear ribonucleoproteins) – introns – intervening, interrupting = removed by spliceosome – exons – expressed Sliceosome snRNPs = small nuclear ribonucleoproteins snRNPs Other proteins Spliceosome Intron Exons Animations http://www.johnkyrk.com/DNAtranscri ption.html Putting it Together – Transcription to Translation How does mRNA code for proteins? How can you code for 20 aa with only 4 nucleotide bases (A, U, G, C)? How can an alphabet of 4 letters (nucleotides) translate into an alphabet of 20 letters (aa)?! Breaking the code Nirenberg and Matthaei Determined 1st codon – amino acid match – UUU coded for phenylalanine Created artificial poly(U) mRNA Added mRNA to test tube of ribosomes and nucleotides – mRNA synthesized a single amino acid polypeptide chain: phe-phe-phe-phe-phe-phe DNA Gene 1 DNA 3’ 5’ Transcription mRNA 5’ Translation 3’ codon Protein Amino acids The CODE!! For ALL life!! (yes, even prokaryotes…) – Strongest support for common origin for all life Code is redundant – Several codons for each amino acid Start codon: AUG = methionine Stop codons: UGA, UAA, UAG Translation Ribosome reads mRNA in codons tRNA brings in correct amino acid tRNA matches codon of mRNA = anticodon Amino acids assembled into polypeptide chain tRNA Structure “clover leaf” structure – anticodon on “clover leaf” end – amino acid on 3’ end – anticodon written 3’ 5’ to match codons which are 5’ 3’ Aminoacyl tRNA synthetase enzyme which bonds amino acid to tRNA – endergonic reaction (does it require energy?) – ATP AMP (how many phosphates do we use?) – Energy stored in tRNA-aa bond Unstable Ribosomes Facilitate coupling of tRNA anticodon to mRNA codon – Organelle or enzyme? Structure – Ribosomal RNA and proteins – 2 subunits: large and small – A site (aminoacyl-tRNA site) Holds tRNA carrying next amino acid to be added to chain – P site (peptidyl-tRNA site) Holds tRNA carrying growing polypeptide chain – E site (exit site) Discharged tRNA leaves ribosome from exit site Building a Polypeptide 1. Initiation 2 3 Brings together mRNA, ribosome subunits, proteins and initiator tRNA Elongation Termination Release polypeptide “release protein” bonds to A site Bonds water molecule to polypeptide chain Polyribosomes Many ribosomes read single mRNA simultaneously making many copies of a protein! Protein Targeting Signal polypeptide – ~20 aa at the beginning of the polypeptide – Recognized by SRPs (signal recognition particles) – SRP brings polypeptide and ribosome to ER so that polypeptide is secreted into the ER as it’s built. Destinations – other signal polypeptides used to target – – – – – – Secretion Nucleus Mitochondria Chloroplasts Cell membrane Cytoplasm Protein Targeting Comparing! Mutations Are all mutations bad? Do all mutations lead to changes in amino acids? Sickle Cell Anemia - Mutation K-ras Oncogene – Point Mutation Mutations Point Mutations – 1 base pair change – Base-pair substitution Silent mutation: no amino acid change because of redundancy in code Missense: change amino acid Nonsense: change to stop Mutations Insertions – adding base(s) Deletions – losing base(s) BOTH cause frameshift Question Which mutations, point mutations or frameshift mutations, do you think are more harmful and why? This weekend… Work on Quest! You should be able to answer everything except the questions about gene regulation which we will do on Monday in class Quest will be due Monday at midnight Gene Regulation In a nutshell… Prokaryotes regulate transcription through operons. Area where RNA Polymerase binds Area where repressor can bind to stop binding of RNA polymerase Segment of DNA containing several Regulatory Gene genes, a promoter, and an operator Codes for repressor; gene is upstream or downstream from operator 2 Kinds of Feedback Prokaryotes regulate transcription through operons. Area where RNA Polymerase binds Area where repressor can bind to stop binding of RNA polymerase Segment of DNA containing several Regulatory genes, promoter, and operator gene = codes for repressor; gene is upstream/do Too much product, stop; wnstream not enough, keep going!! from operator Keep going if you can! 2 Examples of Negative Feedback can bind to stop Prokaryotes regulate transcription through operons. binding of RNA polymerase Segment of DNA containing several Regulatory genes, promoter, and operator gene = codes for repressor; gene is upstream/do Too much product, stop; not wnstream from operator enough, keep going!! •Default “on” because repressor not bound •Product binds to repressor to “activate” and turn “off” transcription when enough product has been made •Usually anabolic pathways (ex: trp) Keep going if you can! can bind to stop Prokaryotes regulate transcription through operons. binding of RNA polymerase Segment of DNA containing several Regulatory genes, promoter, and operator gene = codes for repressor; gene is upstream/do wnstream Too much product, stop; not from operator enough, keep going!! •Default “on” because repressor not bound Keep going if you can! •Default “off” because repressor bound •Product binds to repressor to “activate” and turn “off” transcription •Inducer binds to repressor to “inactivate” and release the repressor and turn “on” transcription only when there is substrate to be broken down •Usually anabolic pathways (ex: trp) •Usually catabolic pathways (ex: lac) can bind to stop Prokaryotes regulate transcription through operons. binding of RNA polymerase Segment of DNA containing several Regulatory genes, promoter, and operator gene = codes for repressor; gene is upstream/do wnstream Too much product, stop; not from operator enough, keep going!! Keep going if you can! •Presence of activator turns “on” •Default “on” because repressor not bound •Ex: lac with lactose and no glucose •Default “off” because repressor bound •Product binds to repressor to “activate” and turn “off” transcription •Product binds to repressor to “inactivate” and release the repressor and turn “on” transcription •Usually anabolic pathways (ex: trp) •Usually catabolic pathways (ex: lac) Summary of Prokaryotic Gene Regulation: Operons • Negative Feedback – Repressible – Inducible • Positive Feedback Gene Regulation In a nutshell… Eukaryotes regulate gene expression pre-transcription, during transcription, and post-transcription. Before transcription; which genes are “on” and “off” •Controls access of RNA polymerase to promoter •Histone acetylation •Acetylated = less bound = easier access •DNA methylation •methylated = more bound = less access Eukaryotes regulate gene expression pre-transcription, during transcription, and post-transcription. •Histone acetylation •Acetylated = less bound = easier access •Controls access of RNA polymerase to promoter •Aids in RNA polymerase binding •DNA methylation •methylated = more bound = less access •Transcription Factors = proteins that help RNA poly binding at promoter •Activation sites and enhancer proteins = also aid in RNA poly binding; 1000s of bp away polymerase to promoter •methylated = more expressionbound pre-transcription, = less access Eukaryotes regulate gene during transcription, and post-transcription. •Aids in RNA polymerase binding After mRNA has been made Translational Regulation RNA Processing mRNA Degradation •Transcription Factors = proteins that help RNA poly binding at promoter •Activation sites and enhancer proteins = also aid in RNA poly binding; 1000s of bp away Protein Processing/Regulation polymerase to promoter •methylated = more expressionbound pre-transcription, = less access Eukaryotes regulate gene during transcription, and post-transcription. •Aids in RNA polymerase binding After mRNA has been made Translational Regulation RNA Processing •Alternate splicing = different combos of exons are expressed (some can be removed) •Essential in making antibodies mRNA Degradation •Transcription Factors = proteins that help RNA poly binding at promoter •Activation sites and enhancer proteins = also aid in RNA poly binding; 1000s of bp away Protein Processing/Regulation polymerase to promoter •methylated = more expressionbound pre-transcription, = less access Eukaryotes regulate gene during transcription, and post-transcription. •Aids in RNA polymerase binding After mRNA has been made Translational Regulation RNA Processing mRNA Degradation •Transcription Factors = proteins that help RNA poly binding at promoter •Activation sites and enhancer proteins = also aid in RNA poly binding; 1000s of bp away Protein Processing/Regulation •Poly-A tail/5’ cap •3’ and 5’ UTR (untranslated region) = nucleotides before the start codon (AUG) and/or after the stop codon •RNAi = RNA interference small interfering RNAs (siRNAs) and microRNAs (miRNAs) = bind to mRNAs and prevent them from being translated or trigger their degradation polymerase to promoter •methylated = more expressionbound pre-transcription, = less access Eukaryotes regulate gene during transcription, and post-transcription. •Aids in RNA polymerase binding After mRNA has been made Translational Regulation RNA Processing •Poly-A tail/5’ cap = mRNAaffect assembly/binding of Degradation ribosome •3’ and 5’ UTR (untranslated region) = nucleotides before the start codon (AUG) and/or after the stop codon •Transcription Factors = proteins that help RNA poly binding at promoter •Activation sites and enhancer proteins = also aid in RNA poly binding; 1000s of bp away Protein Processing/Regulation polymerase to promoter •methylated = more expressionbound pre-transcription, = less access Eukaryotes regulate gene during transcription, and post-transcription. •Aids in RNA polymerase binding After mRNA has been made Translational Regulation RNA Processing mRNA Degradation •Transcription Factors = proteins that help RNA poly binding at promoter •Activation sites and enhancer proteins = also aid in RNA poly binding; 1000s of bp away Protein Processing/Regulation •Processing= some proteins need to get folded, spliced (parts cut off), or have groups added •Degradation= all proteins need to be “marked” for degradation/to get broken down; most “marked” by ubiquitin and broken down by proteosomes Summary of Eukaryotic Gene Regulation • Pre-transcriptional • During transcription • Post-transcriptional