* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download transcription
Site-specific recombinase technology wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Genetic code wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
History of genetic engineering wikipedia , lookup
Transfer RNA wikipedia , lookup
X-inactivation wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Gene expression profiling wikipedia , lookup
Designer baby wikipedia , lookup
Microevolution wikipedia , lookup
Non-coding DNA wikipedia , lookup
Point mutation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Nutriepigenomics wikipedia , lookup
RNA interference wikipedia , lookup
Transcription factor wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Polyadenylation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
RNA silencing wikipedia , lookup
Epigenetics of human development wikipedia , lookup
History of RNA biology wikipedia , lookup
Messenger RNA wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Non-coding RNA wikipedia , lookup
Transcription in Prokaryotes
by Jean-Pierre Herveg, Etienne De Plaen and a lot of friends at
the Brussels Branch of the Ludwig Institute for Cancer research (Licr) and the Christian de Duve*
Institute for cellular Patholgy (ICP).
April 2006
Université Catholique de Louvain
Avenue E. Mounier, 1200 Brussels (Belgium)
Questions
what is the meaning of the sentence :” The sequence of a prokaryotic protein
is colinear with the translated mRNA
2. which sugar replaces desoxyribose in ARN ?
3. which base anneals to uracil ?
4. what’s a polysome ?
5. what’s an operon, an operator ? Describe !
6. what is X-gal.
7. what is lacZ?
8. what is IPTG?
9. describe a promoter.
10. what is the role of sigma in prokaryotes
11. why are there a pUR278, pUR289 and a PUR288 to make fusion proteins
12. explain the control of protein production by pET-3a
1
Transcription in prokaryotes
The sequence of a prokaryotic protein is colinear with the translated mRNA;
that is, the transcript of the gene is the molecule that is translated into the polypeptide.
In prokaryotes, ther is no nucleus and then replication, transcription and translation take place in
The same compartment.
Reverse transcription is the opposite
The enzymes involved are RNA polymerase for transcription and reverse transcriptase for the opposite.
-------------------------question
what is the meaning of the sentence :” The sequence of a prokaryotic protein is colinear with
the translated mRNA
Ribose:
Is a sugar containing 5 carbon atoms and a hydroxyle in the position 2’
Uracile
Is a RNA base which is represented by a U. U anneals with A.
-------------------------------------La ribosa
Es un azúcar de 5 carbonos. Es un componente estructural de la estructura del ARN, como el ATP, GTP, CTP y TTP.
Uracilo
Es una de las 4 bases del ARN. Se representa con la letra U. En el ARN, El uracilo reemplaza a la timina.
El uracilo se aparea con la adenina.
--------------------------------------Cuestiones para el examen
1. ¿ Cuál es el azúcar y la basa específicos del ARN
2. ¿ Con cuál base se aparea el uralcilo?
There are 5 types of RNA, each encoded by its own type of gene:
mRNA -(80 % in eucaryoyes) Messenger RNA: Encodes the amino acid sequence of a polypeptide.
tRNA - (15 % in eucaryoyes) Transfer RNA: Brings amino acids to ribosomes during translation.
mRNA - (5 % in eucaryoyes) Ribosomal RNA: with ribosomal proteins, makes up the ribosomes.
rRNA possess an enzymic activity. The ribosome is the organelles that translate the mRNA into polypeptide.
snRNA - Small nuclear RNA: With proteins, forms complexes that are used in RNA processing
siRNA and miRNA: are inhibitor od gene expression
Transcription and translation take place in the same compartment.
In prokaryotes, there is simultaneous transcription and translation.
Here, in E. Coli you can see a long fiber running from top to bottom (green arrow ). This fiber is a
Segment of the E. coli chromosome (dsDNA).
This picture comes from:
http://users.rcn.com/jkimball.ma.ultranet/BiologyPages/M/Miller_Hamkalo.html
Extending from the DNA are polysomes (red arrow),
A polysome is a backbone of messenger RNA (mRNA) to which ribosomes are
attached.
On the left picture, you can see that the size of polysomes increases from top
to bottom.
Each polysome is attached to the DNA fiber by a complex of proteins
that includes a molecule of RNA polymerase.
Thus
the DNA is transcribed by RNA polymerase molecules moving
from top to bottom, and
the growing mRNA molecules are translated by ribosomes
moving in a proximal -> distal direction.
In E. coli, then, and probably in all prokaryotes,
the transcription of DNA into mRNA and the translation of mRNA into
polypeptides (not visible here) are closely coordinated
in both time and space.
(In eukaryotes, in contrast, while all transcription takes place in the
nucleus, most (but not all) translation of mRNA occurs later in the cytosol).
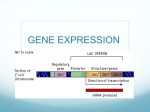
Defintion:
An operon is a group of genes including a common promoter and an operator, which control the
transcription of these genes to produce a common messenger RNA (mRNA).
A promoter is a DNA sequence that enables a gene to be transcribed.
The promoter is recognized by RNA polymerase, which then initiates transcription.
An operator is a segment of DNA that recognized by a regulatory protein (inducer or or repressor).
In negative inducible operons (the lac operon), a regulatory repressor protein is normally
bound to the operator and it prevents the transcription of the genes on the operon
If an inducer molecule is present, it binds to repressor and changes its conformation
so that it is unable to bind to the operator.
This allows for the transcription of the genes controlled by the operator.
In negative repressible operons, (the trp operon) transcription of the genes on the operon
normally takes place. Repressor proteins are produced by a regulator gene but they are
unable to bind to the operator in their normal conformation. However certain molecules
called corepressors can bind to the repressor protein and change its conformation so that
it can bind to the operator. The activated repressor protein binds to the operator and
prevents transcription.
Operons can also be positively controlled. With positive control, an activator protein
stimulates transcription by binding to DNA (usually at a site other than the operator).
a negative inducible operons:
the lac operon consists of a regulatory gene and 3 structural genes (z, y, and a).
The i gene codes for a repressor. The z gene codes for ß-galactosidase, which hydrolyses lactose into
galactose and glucose. The y gene codes for permease, which increases permeability to ß-galactosides.
The a gene encodes a transacetylase.
In the presence of an inducer of the lac operon, the repressor protein binds the inducer. RNA polymerase is
thus able to bind at the promoter region, and transcription of the operon ensues.
Lactose analogues
1. IPTG is an inducer (acts like lactose, but is not metabolized by
ß-galactosidase).
A number of lactose derivatives or analogs have been described that are useful for work with the lac operon.
These compounds are mainly substituted galactosides, where the glucose moiety of lactose is replaced
by another chemical group.
Isopropyl-β-D-thio-galactoside (IPTG) is frequently used as an inducer of the lac operon
for physiological work. IPTG binds to repressor and inactivates it, but is not a substrate for β-galactosidase.
One advantage of IPTG for in vivo studies is that it cannot be metabolized by E. coli, therefore the growth
rate of cells (usually maintained with glycerol as the carbon and energy source), is not a variable
in the experiment.
In addition, IPTG is transported efficiently independent of whether the lacY gene
is functional. And since cells don't metabolize IPTG, its concentration doesn't change during the course
of an experiment.
Lactose analogues
1. X-gal (is metabolizide and gives a blue reaction product).
X-gal (5-bromo-4-chloro-3-indolyl-β-D-galactoside) turns colonies which produce
β-galactosidase blue. IPTG is used in what is called the white andblue screening.
http://en.wikipedia.org/wiki/Lac_operon
a negative repressible operon:
The trp operon encodes the genes for the synthesis of tryptophan.
This cluster of genes is regulated by a repressor that could bind to the operator sequence.
The activity of the trp repressor for binding the operator region is enhanced when it binds tryptophan:
tryptophan is known as a corepressor.
Since the activity of the trp repressor is enhanced in the presence of tryptophan
the rate of expression of the trp operon is graded in response to the level of tryptophan in the cell (attenuation
Element).
--------------------------------Cuestiones para el examen
1. Definir la palabra opéron en las bacterias
2. ¿ Que es que X-gal.
3. ¿ Que es que lacZ?
4. ¿ Que es que IPTG?
5. describir el opéron lac, trp…
Fine regulation of the tryptophane operon.
Expression of the trp operon is also regulated by attenuation (attenuator element).
The attenuator region is involved in controlling transcription from the operon after RNA polymerase
has initiated synthesis.
---------------------------------------------to stall: calar (autocalar)
Beginning of transcription
The RNA pol is a tetrameric enzyme. The factor sigma helps to recognize the promoters.
The genetic information contained in DNA is transcribed to RNA by a transcription complex
Including DNA-directed RNA polymerase (RNA Pol). A bacterial transcription initiation complex
comprised of the core RNA Pol enzyme and a (sigma) factor binds to a promoter and,
upon initiation of RNA synthesis, releases the s factor.
http://dnaresearch.oxfordjournals.org/cgi/content/short/dsi016v1
Institution: Bolivia: PNAS Sponsored
Published online before print March 27, 2006,
PNAS | April 4, 2006 | vol. 103 | no. 14 | 5332-5337
Biological Science / Biochemistry
Insights into transcriptional regulation and {sigma} competition from an
equilibrium model of RNA
polymerase binding to DNA
Irina L. Grigorova, Naum J. Phleger, Vivek K. Mutalik, and Carol A. Gros
In bacteria transcription is initiated by RNA polymerase (RNAP) holoenzyme (Es) which is formed
when core RNAP (E) binds the transcription initiation factor s (1). Es initially binds to promoter sites
in a closed complex, which then transits to an open complex, competent for transcription.
The number of intermediates between the closed and open complex is variable and promoter-dependent;
each step may be subject to regulation in vivo (2, 3). At least for some promoters, Es binding to promoters
is thought to be reversible on the time scale of transcription initiation in vivo (3); reversibility has
also been demonstrated in vitro for several promoters (3–6). Even binding to the strong lac UV5
promoter is reversible in vitro when tested under conditions that approximate the in vivo situation (6).
Recruitment of Es to promoters in vivo is thought to depend on the intrinsic binding affinity of the
promoter and is modulated by repressors that prevent and activators that stabilize interactions between
Es and the promoter (3). Based on in vitro studies of the mechanism of activator function, it is believed
that promoters that bind Es weakly require activators to recruit Es In addition, cells contain multiple s,
which direct E to various sets of promoters specific to the s factors (1). These s s are believed to compete
with each other for binding to E (7–10). By changing the relative levels of the {sigma}s, Escherichia coli is
thought to coordinate its transcriptional program with growth conditions (11–13). This view is based
on observations indicating that (i) overexpressing one s decreases expression of genes controlled
by another s (7), (ii) mutationally altering binding constants of one s for E, alters expression by another s(14),
and (iii) physiological effectors such as ppGpp may act by altering relative binding of s s to E (8–10).
In the present work, we use an equilibrium model of RNAP binding to DNA to explore in vivo scenarios
that permit transcription regulation by activator recruitment of RNAP and Es competition.
The promoter is recognized by RNA polymerase and the s factor. RNA pol then
initiates transcription. When the transcript is around 10 nt long, the s factor
quits the complex
-----------------------------Los promotores bacterianos poseen cuatro características comunes:
(1)
lel punto de comienzo de la transcripción, denominado +1,
(2)
la secuencia de la posición –10 llamada caja de Pribnow (TATAAT),
(3)
la secuencia –35 (TGTTGACA) y
(4)
la distancia entre las secuencias –10 y –35. E punto donde se inicia la transcripción es genralmente
una purina (A ou G).
the end of transcription: termination
Two termination mechanisms are well known:
Rho-independent termination involves terminator sequences within the
RNA that signal the RNA polymerase to stop. The terminator sequence is usually a palindromic sequence
that forms a stem-loop hairpin structure that leads to the dissociation of the RNA Pol from the DNA template.
Rho-dependent termination uses a termination factor called r (rho) factor to stop RNA synthesis at specific sites.
This protein binds and runs along the mRNA towards the RNA Pol. When the r (rho) factor reaches the RNA Pol
it causes RNA Pol to dissociate from the DNA, terminating transcription.
Other termination mechanisms include where RNA Pol comes across a region with repetitious thymidine
residues in the DNA template. or where a GC-rich inverted repeat followed by 4 A residues
the inverted repeat forms a stable stem loop structure in the Rna, which causes the RNA to dissociate from
the DNA template.
Translation (genetic code)
Table of the codons and amino acids
(one letter code)
codon iniciador AUG
Stop: TAA, TAG, TGA
uuu
uuc
uua
uug
F
F
L
L
ucu
ucc
uca
ucg
S
S
S
S
uau
uac
uaa
uag
Y
Y
stop
stop
ugu
ugc
tga
ugg
C
C
stop
W
cuu
cuc
cua
cug
L
L
L
L
ccu
ccc
cca
ccg
P
P
P
P
cau
cac
caa
cag
H
H
Q
Q
cgu
cgc
cga
cgg
R
R
R
R
auu
auc
aua
atg
I
I
I
M
acu
acc
aca
acg
T
T
T
T
aau
aac
aaa
aag
N
N
K
K
agu
agc
aga
agg
S
R
R
R
guu
guc
gua
gug
V
V
V
V
gcu
gcc
gca
gcg
A
A
A
A
gau
gac
gaa
gag
D
D
E
E
ggu
ggc
gga
ggg
G
G
G
G
Acido Aspartico : Asp = D
Acido glutamico : Glu = E
Alanina : Ala = A
Arginina : Arg = R
Asparagina : Asp = N
Cisteina : Cys = C
fenilalana : Phe = F
Glicina : Gly = G
Glutamina : Gln = Q
Histidina : His = H
isoleucina :Ile = I
Leucina :Leu = L
Lisina : Lys = K
Metioniona : Met = M
Prolina : Pro = P
Serina : Ser = S
Tirosina : Tyr = Y
Treonina : Thr = T
Triptofana : Trp = W
Valina : Val = V
Free tRNAs are loaded with their specific amino-acyl by an
Aminoacyl-tRNA transferase.
They are now ready to participate to translation.
Free tRNA, are made of a short RNA sequence, in which they are anticodons
tRNA are made of a short RNA sequence, in which they are anticodons they can
Be loaded with an aùino-acyl.
In the ribosome (large subunit), two sites A and P can be loaded each with
aminoacyl-tRNA.
A peptide bond is the made between thes two aminoacyls.
Then, the tRNA molecule is expelled see the black arrow below.
It reached the place where it could be loaded with another aminoacyl.
The ribosome moves (scarlet arrows, the tRNA in the second site moves in the first one
And another aminoacyl-tRNA takes its place.
Comparaison between prokaryotes and eukaryotes:
fused proteins
pUR278
pMAL
non fused proteins
pET-3a
---------------------------question
explain the control of protein production by pET-3a
pQE (Quiagen), HIS tag
Exercises: recombinant insulin and growth hormone
Search for the insulin gene:
Database: Gene (1)
For: INS or INSULIN, both are okay.
The click on Go
You should first read the Summary…. And remember the significance of signal peptide, post
Translationnaly, disulfide bonds, glucose receptor and glucose uptake.
What’s a minus strand ?
Does this gene have an intron ?
On what chromosome stand the gene ?
What is the meaning of p in 11p5.5 ?
Now, click on NM_000207.1 (DEFINITION homo sapiens insulin (INS), mRNA.
recombinant human insulin rh-insulin
LOCUS
NM_000207
450 bp mRNA
DEFINITION Homo sapiens insulin (INS), mRNA.
1
61
121
181
241
301
361
421
gctgcatcag
gcctcctgcc
tgaaccaaca
gaggcttctt
tggagctggg
tgcagaagcg
agaactactg
agagagatgg
aagaggccat
cctgctggcg
cctgtgcggc
ctacacaccc
cgggggccct
tggcattgtg
caactagacg
aataaagccc
caagcacatc
ctgctggccc
tcacacctgg
aagacccgcc
ggtgcaggca
gaacaatgct
cagcccgcag
ttgaaccagc
linear PRI 29-JAN-2006
actgtccttc
tctggggacc
tggaagctct
gggaggcaga
gcctgcagcc
gtaccagcat
gcagcccccc
tgccatggcc
tgacccagcc
ctacctagtg
ggacctgcag
cttggccctg
ctgctccctc
acccgccgcc
ctgtggatgc
gcagcctttg
tgcggggaac
gtggggcagg
gaggggtccc
taccagctgg
tcctgcaccg
/translation="MALWMRLLPLLALLALWGPDPAAAFVNQHLCGSHLVEALYLVCERGFFYTPKTRREAEDLQVG
QVELGGGPGAGSLQPLALEGSLQKRGIVEQCCTSICSLYQLENYCN"
proprotein
45..374 (329 = 100%)
sig_peptide
45..116 (71/329= 0.215)
atggcc ctgtggatgc gcctcctgcc cctgctggcg ctgctggccc tctggggacc tgacccagcc gcagcc
mat_peptide 117..374 (
/product="proinsulin"
mat_peptide 117..206 (89/329= 0,27)
tttgtgaaccaaca cctgtgcggc tcacacctgg tggaagctct ctacctagtg tgcggggaac gaggcttctt
ctacacaccc aagac
/product="proinsulin peptide B"
mat_peptide 312..374 (62/329= 0.188)
ggcattgtg gaacaatgct gtaccagcat ctgctccctc taccagctgg agaactactg caac
/product="proinsulin peptide A”
In between 0.327 %
Summary:
After removal of the precursor signal peptide, proinsulin is post-translationally cleaved into two chains
(peptide A and peptide B) that are covalently linked via two disulfide bonds.
Binding of this mature form of insulin to the insulin receptor (INSR) stimulates glucose uptake.
Official Symbol: INS and Name: insulin [Homo sapiens]
Other Designations: proinsulin
Chromosome: 11; Location: 11p15.5
GeneID: 3630
The mRNA is translated in a protein which have four domains: The hydrophobic siganl peptide is cleaved in the ER
f the eukaryotic cell. The C peptide is cleaved in the blood by proteases.
At the end of this process, we are left with this structure:
The cDNA coding proinsulin is used to make protein fusion with B-gal. An atg is added at the beginnning.
The fusion protein is purified. The ß-gal beginning ic cleaved by cyanogen bromide. The c peptide is cleaved by
A protease.
121
181
241
301
361
atg +
tttg
tgaaccaaca
gaggcttctt
tggagctggg
tgcagaagcg
agaactactg
cctgtgcggc
ctacacaccc
cgggggccct
tggcattgtg
caactag
tcacacctgg
aagacccgcc
ggtgcaggca
gaacaatgct
tggaagctct
gggaggcaga
gcctgcagcc
gtaccagcat
ctacctagtg
ggacctgcag
cttggccctg
ctgctccctc
tgcggggaac
gtggggcagg
gaggggtccc
taccagctgg
human growth hormone (hGH)
Official Symbol: GH1 and Name: growth hormone 1 [Homo sapiens]
Other Aliases: GH, GH-N, GHN, hGH-N
Other Designations: pituitary growth hormone
Chromosome: 17; Location: 17q24.2
GeneID: 2688
The protein encoded by this gene is a member of the somatotropin/prolactin family of hormones
which play an important role in growth control. The gene, along with four other related genes,
is located at the growth hormone locus on chromosome 17 where they are interspersed in the
same transcriptional orientation; an arrangement which is thought to have evolved by a series
of gene duplications. The five genes share a remarkably high degree of sequence identity.
Alternative splicing generates additional isoforms of each of the five growth hormones,
leading to further diversity and potential for specialization. This particular family member is expressed
in the pituitary but not in placental tissue as is the case for the other four genes in the growth hormone
locus. Mutations in or deletions of the gene lead to growth hormone deficiency and short stature.
LOCUS
NM_022562
376 bp mRNA linear PRI 26-FEB-2006
DEFINITION Homo sapiens growth hormone 1 (GH1), transcript variant 5, mRNA.
ACCESSION NM_022562
VERSION NM_022562.2 GI:20809252
1
61
121
181
241
301
361
aggatcccaa
caatggctac
tacagcaagt
ctctactgct
cgctctgtgg
ccagtgcctc
aaattaagtt
ggcccaactc
agaggctgga
tcgacacaaa
tcaggaagga
agggcagctg
tcctggccct
gcatca
cccgaaccac
agatggcagc
ctcacacaac
catggacaag
tggcttctag
ggaagttgcc
tcagggtcct
ccccggactg
gatgacgcac
gtcgagacat
ctgcccgggt
actccagtgc
gtggacagct
ggcagatctt
tactcaagaa
tcctgcgcat
ggcatccctg
ccaccagcct
FEATURES
source
gene
Location/Qualifiers
1..376 /organism="Homo sapiens"
1..376
/gene="GH1"
/note="synonyms: GH, GHN, GH-N, hGH-N"
CDS
63..155
/gene="GH1"
/translation="MATEAGRWQPPDWADLQADLQQVRHKLTQR"
misc_feature 72^73
/gene="GH1"
/note="Region: location of alternate exons 2, 3 and 4"
STS
197..336 (STS= sequence tagged sites)
see http://cfern.bio.utk.edu/journal/Hamilton1998/THESIS_A1.htm
polyA_signal 357..362
polyA_site
376
atggcta agaggctgga agatggcagc ccccggactg ggcagatctt caagcagacc
tacagcaagt tcgacacaaa ctcacacaac gatga
cacctagctg
caagcagacc
ctacgggctg
cgtgcagtgc
tgacccctcc
tgtcctaata