* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Fab-7 1 + +
DNA vaccination wikipedia , lookup
Genetic engineering wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Primary transcript wikipedia , lookup
Genomic imprinting wikipedia , lookup
X-inactivation wikipedia , lookup
Genome evolution wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Point mutation wikipedia , lookup
Protein moonlighting wikipedia , lookup
Genome (book) wikipedia , lookup
Epigenetics wikipedia , lookup
Minimal genome wikipedia , lookup
Gene expression profiling wikipedia , lookup
Histone acetyltransferase wikipedia , lookup
Genome editing wikipedia , lookup
Microevolution wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Designer baby wikipedia , lookup
Helitron (biology) wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
History of genetic engineering wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Epigenomics wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
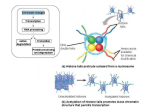
Introduction in: Epigenetic regulation by Polycomb and trithorax group proteins Giacomo CAVALLI. Montpellier, December 2006 Cavalli Lab The chromatin Yin / Yang Heterochromatin - Heavily condensed - Gene poor - Silent genes - Late replicating - DNA hypermethylated - Rich in histone H1 - Histones have repressive post translational marks Euchrochromatin - Less condensed - Gene rich - Active genes - Early replicating - DNA hypomethylated - Poor in histone H1 - Histones have specific, activating post translational marks Cavalli Lab Where on the chromosomes are condensed and open chromatin located? Condensed chromatin Constitutive Heterochromatin Telomeres Centromeres Open chromatin Typical of the active gene loci along the chromosomal arms Condensed chromatin is also found at several silent genes located in the chromosomal arms Polycomb group and trithorax group genes are important regulators of chromatin along the chromosomal arms Polycomb (PcG) and trithorax (trxG) group proteins: epigenetic regulators of genome function • Originally discovered in Drosophila as regulators of Homeotic genes, responsible for specification of the body plan, they also regulate many other targets involved in cell differentiation and proliferation • PcG proteins silence genes, trxG proteins activate them • Conserved throughout evolution • In human, they maintain the fates of differentiated cells, but they are also required for cell proliferation and the maintenance of several types of stem cells including ES cells. Finally, they regulate X-inactivation in females as well as genomic imprinting • Mutations in PcG and trxG genes induce many different cancers PcG, trxG, and maintenance of gene expression Early development Establishment of patterns OFF Ubx ON Maternal, Gap, Pair-rule, Segment polarity OFF Polycomb-Group OFF ON Maintenance phase Transmission of pattern after disappearance of early factors trithorax-Group ON PcG and trxG proteins associate to multiple genomic loci DAPI Merge PH Polytene chromosome staining shows around 100 bands for each PcG protein PcG proteins bind to specific DNA elements, named PREs Bound by PcG proteins in vivo (in polytene chromosomes and by cross-linking experiments) Binding leads to maintenance of PcG-dependent repression of reporter genes Repression is enhanced by the presence of multiple PRE copies Members of the PcG and of the trxG PcG Gene protein motifs PhoRC Complex Pleiohomeotic (pho) Zinc-finger (DNA binding) Enhancer of zeste (E(z)) SET (H3K27MTase) extra sex combs (esc) WD repeat Polycomb (Pc) chromo domain (Binding to H3 methyl K9 or K27) polyhomeotic (ph) Zinc finger, SAM domain Posterior sex combs (Psc) RING finger trxG Gene protein motifs FACT complex Trithorax-like (Trl) BTB/POZ (dimerization) Zinc finger (DNA binding) TAC1 complex trithorax (trx) SET (H3K4HMTase)/ PHD-finger Ash-1 SET (H3/H4HMTase)/ PHD-finger brahma (brm) bromo domain Esc/E(z) Complex PRC1 complex Brm complex (DNA dependent ATPase/helicase) Action of PcG and trxG complexes on chromatin Ac Histone acetylation and methylation (TAC1 and ASH1 complexes) Nucleosome remodeling (BRM complex) trxG ON Maintenance of active states (open chromatin) Me K4 H3 Target gene PRE Deacetylation and methylation (ESC-E(Z) complex) Me K27 H3 - Chromatin compaction - H2A Ubiquitination (PRC1 complex) PcG OFF Maintenance of repressed states (compact chromatin) Ub H2A Histone H3 K27 methylation and Polycomb Pc H3K27me3 Merge Data from: Ringrose et al. (2004) Mol. Cell 16, 641 There is a strong correlation between trimethylation of K27 (and K9) trimethylation and Polycomb recruitment at target loci. What does Polycomb do to chromatin ? Chromatin Condensation Recombinant PC-containing complexes can condense an array of 12 nucleosomes in vitro Condensation requires PSC (not PH) protein, and involves histones but does not necessitate histone tails Data from: Francis et al. (2004), Science 306, 1574 Features of PREs and TREs, and examples of how they are studied in Drosophila (No PREs characterized yet in vertebrates) 1. Example of PcG-dependent spatial specific silencing of homeotic genes bxd5.1 UbxlacZ reporter construct Bxd 5.1 PRE Silencing of a Ubx-lacZ reporter mimicking the wt behaviour of the Ubx gene, which is silenced in parasegments 1 to 5 Ubx prom LacZ mini-white PcG dependent derepression of a UbxlacZ reporter in embryonic territories where it is normally silenced Data from: Hodgson, J. W., Argiropoulos, B., and Brock, H. W. (2001). Site-specific recognition of a 70-base-pair element containing d(GA)(n) repeats mediates bithoraxoid polycomb group response element-dependent silencing. Mol Cell Biol 21, 4528-4543. 2. Silencing of a transgenic reporter gene depends on PcG and trxG proteins Fab-7 Fab-7 Pc -/+ Fab-7 Pc +/+ UAS-lacZ white Fab-7 trx -/+ Fab-7 trx +/+ 3. Recruitment of PcG and trxG proteins to PREs: analysis in Fab-7 by a combination of immunostaining and FISH in polytene chromosomes (immuno-FISH) Fab-7 UAS-lacZ white Transgene : transgene 24A 25E5 DAPI Immunostaining of PH protein FISH Immuno-FISH 4. Recruitment of PcG and trxG proteins to PREs: analysis by ChIP Recruitment of PcG proteins at PREs: chromatin analysis by chromatin immunoprecipitation (ChIP) and DNA microarrays (chips) Cross-link cells or embryos with formaldehyde to induce protein-DNA crosslinks Sonicate and purify chromatin (average size = 1 kb) Add antibody and purify antibody-chromatin complexes on Protein A Sepharose, purify DNA and amplify by Linker-mediated PCR Use amplified DNA as probe to hybridize DNA chips containing the genome. Extract the distribution profile of the proteins of interest ChIP on chip PH binding profiles at the embryonic developmental stage X sd X chromosome tip 2L Montpellier tiling paths Yale tiling paths Adh region 2R en/inv 3R 3L L82 Hh Bx-C 4 ci bi (1) noc ph esg mab2(3) gt/z 2D (2) 4C 5A 5D elB ct (4) 7B 34D 35B stc 35D 36A Negre N, Hennetin J, Sun LV, Lavrov S, Bellis M, White KP, Cavalli G. Chromosomal Distribution of PcG Proteins during Drosophila Development. PLoS Biol. 2006 Apr 20;4(6):e170 Example of a high-resolution description of PcG binding using ChIP and oligonucleotides arrays with one oligo every 100 bp H3K27me3 PC PH PcG proteins are required for the maintenance of ES cell fate, as well as for the fate of hematopoietic and neuronal stem cells and of differentiated cell types Genome-wide Chip on chip in ES cells Transcription factor family members occupied by the PRC2 protein SUZ12 in human ES cells Overlap between PRC1 and PRC2 targets in mouse ES cells Eed (831) PRC2 66 IRX BHLHB CDX PRC1 365 94 Suz12 (1271) HOX Rnf2 (1219) MEIS/EVX TBX 55 300 DLX 164 Phc (922) HES FOX ATOH 512 98 6 121 MYO 31 NEUROD EBF GATA POU RUNX 23 PAX SOX NKX Boyer et al. Nature. 2006 May 18;441(7091):349-53. Epub 2006 Apr 19 SIX LHX Lee et al. Cell. 2006 Apr 21;125(2):301-13 5. Cross-talk between multiple copies of PREs Pairing Sensitive Silencing: enhanced silencing of the mini-white reporter gene by multiple PRE copies P Fab-7 Transgenic Fab-7 heterozygous w mini-white Chromosome X P Transgenic Fab-7 homozygous w w ---> weak silencing of the mini-white reporter gene ---> strong mini-white silencing PcG-mediated silencing is enhanced by the presence of multiple copies of homologous PREs in the genome Fab-X; Fab71 Fab-X Fab-7 transgene Endogenous Fab-7 sd sd X X BX-C BX-C III III Endogenous Fab-7 deletion, named Fab-71 Data from: Bantignies, F., Grimaud, C., Lavrov, S., Gabut, M., and Cavalli, G. (2003). Inheritance of Polycomb-dependent chromosomal interactions in Drosophila. Genes Dev 17, 2406-2420. Homologous Fab-7 copies associate in the nucleus Dapi Sd BX-C Merge WT sd BX-C Fab-7 Fab-X sd Fab-7 BX-C Fab-7 25 % of interaction - 20 - 15 - 10 - 5 - 0 - Fab-X; Fab-71 sd Fab-7 BX-C WT 3mm Transgenic Fab-7 at sd (Chr.X) - Endogenous Fab-7 at the BX-C (Chr.III) + Fab-X Fab-X; Fab-71 + + + - Two identical Fab-7 can interact in the nucleus, leading to an increase of PcG-dependent silencing Nucleus in a Rabl configuration For more info… http://www.igh.cnrs.fr/equip/cavalli http://www.epigenome-noe.net In french… Inserm Actualités N. 201, Octobre 2006 (http://www.insermactualites.fr/index.php?id=562) [email protected]