* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download DNA-guided genome editing using the
Oncogenomics wikipedia , lookup
Non-coding DNA wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Genomic imprinting wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Transposable element wikipedia , lookup
RNA interference wikipedia , lookup
Copy-number variation wikipedia , lookup
Gene expression programming wikipedia , lookup
RNA silencing wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Gene nomenclature wikipedia , lookup
Gene desert wikipedia , lookup
Public health genomics wikipedia , lookup
Gene therapy wikipedia , lookup
Human genome wikipedia , lookup
Microevolution wikipedia , lookup
Gene expression profiling wikipedia , lookup
Genetic engineering wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genome (book) wikipedia , lookup
Minimal genome wikipedia , lookup
Helitron (biology) wikipedia , lookup
Genomic library wikipedia , lookup
Pathogenomics wikipedia , lookup
Metagenomics wikipedia , lookup
Human Genome Project wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Designer baby wikipedia , lookup
History of genetic engineering wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genome evolution wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
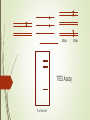
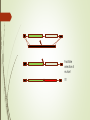
DNA-guided genome editing using the Natronobacterium gregoryi Argonaute Feng Gao, Xiao Z Shen, Feng Jiang, Yongqiang Wu & Chunyu Han Gene Editing Tools Overview https://en.wikipedia.org/wiki/Genome_editing In order to do a targeted genome editing, we need to make a double stranded breaks at specific sites within genome Existing technologies: Meganucleases TALEN ZFN CRISPR-Cas9 http://www.the-scientist.com/?articles.view/articleNo/45119/title/-Heroesof-CRISPR--Disputed/ http://www.nature.com/nature/journal/v534/n7605/full/534037a.html CRISPR https://biotechconnectla.files.wordpress.com/2015/06/crispr-cas9-figure-1.jpg Relevancy in Gene Editing https://www.neb.com/tools-and-resources/feature-articles/crispr-cas9and-targeted-genome-editing-a-new-era-in-molecular-biology CRISPR vs NgAgo-gDNA CRISPR - Template of ~20bp NgAgo-gDNA - Template of ~24bp - Requires gRNA, CAS 9 and - Template must be PAM phosphorylated at the 5’ end - 30-60% efficacy in KDs - Low % of off target mutants - 1-15% efficacy in KIs - Variable % off-target mutants Natronobacterium gregoryi Argonaute Natronobacterium gregori : An halophilic and alkiphilic bacteria that grows at 37°C. Argonaute : a family of endonucleases that require a 5’pSSDNA guide Natronobacterium gregoryi, DSM 3393 (EM from M. Rohde, HZI) Why N. gregoryi? Other species with endonuclease Argonaute protein: 1. Thermus thermophilus 2. Pyrococcus furiosus First things first: Does it work in vivo? +Proteinase K +Dnase I First things first: Does it work in vivo? - Cleavage at 37°C - Can only cleave with guide (either FW or RV), could not cleave with noncomplementary guide First things first: Does it work in vivo? Specificity Experiment -NgAgo can only use a 5’ phosphorylated single strand DNA guide. What about off-target cleavage? Are you sure it’s an endonuclease? Normally expresses green fluorescent protein Is it better than CRISPR? When using CRISPR, the guides are labeled sgRNA for “Short Guide RNA” Optimization Sequence Length Sequence 400bp T7E3 Assay % of mutant 100bp Figure 4a. Silencing human DYRK1A gene G5: 31% G6: 34% G10: 41% G12: 27% G13: 39% DYRK1A (Dual-Specificity Tyrosine-(Y)Phosphorylation Regulated Kinase 1A) gene. Diseases associated with DYRK1A include microcephaly and seizure disorder. Extensively used in CRISPR. Figure 4b. Silencing other human genes EMX1: 24.5% GRIN2B: 26.2% GATA4: 24.8% HBA2: 29% Figure 4c. Silencing DYRK1A gene in other human cell lines 293T: 20.3% MCF7: 13.7% K562: 24.8% HeLa: 11.2% Figure 4d. Determining critical NT in guide G10 Single mismatch of NT 8 completely abolish its function Single mismatch of NT 9-11 severely affects its function Three consecutive mismatches anywhere completely abolish its function Figure 4e. NgAgo is comparable to CRISPR in silencing DYRK1A gene NgAgo: 29.7-34.4% CRISPR: 31.2-33% Figure 4f. NgAgo is better than CRISPR in silencing HBA2 and GATA4 (GC rich) genes HBA2: NgAgo: 37.6% CRISPR: 0% GATA4: NgAgo: 31.5% CRISPR: 13.1% Facilitate selection of mutant ?? CRISPR vs NgAgo-gDNA CRISPR - Template of ~20bp NgAgo-gDNA - Template of ~24bp - Requires gRNA, CAS 9 and - Template must be PAM phosphorylated at the 5’ end - 30-60% efficacy in KDs - Low % of off target mutants - 1-15% efficacy in KIs - Variable % off-target mutants Research ethics Supplementary Figure 9 Full-length gel images (Unrelated lanes are marked with cross). a, for Fig 1a:Nucleic acids associated with NgAgo in E.coli. b, for Fig 1b: The in vitro plasmid cleavage assay(E.coli.-derived NgAgo). c, for Fig 1c: The in vitro plasmid cleavage assay(E.coli.-derived NgAgo, guides with or without 5' phosphorylation). d, for Fig 2a. e, for Fig 2b. f, for Fig 2c. g, for Fig 3a: The in vitro plasmid cleavage assay (293T cell-derived NgAgo). h, for Fig 3c: western blot (GFP,ACTIN). i, for Fig 3d: western blot (GFP,ACTIN). j, for Fig 4a: T7E1 (DYRK1A) . k, for Fig 4b: T7E1 (DYRK1A,EMX1,GRIN2B,GATA4,HBA2). Summary of Dal-iGEM project • 4 microbiome samples submitted for sequencing • 2-4 bacterial strains isolated from cellulose plates • 2nd trip to park for fecal samples from more animals and plants • a. Microbiome profiling • b. Metagenomic sequencing • Plating for more isolates that can utilize cellulose • Secondary screen of isolates that can detoxify sap • c. Genome sequencing of an ideal bacterial strain that could utilized cellulose and detoxify sap • Cloning of the genes that are responsible for the phenotypes • Cloning of NgAgo into a broad-range bacterial vector and deposit in BioBrick