* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download RER - Botanik in Bonn
G protein–coupled receptor wikipedia , lookup
Extracellular matrix wikipedia , lookup
Magnesium transporter wikipedia , lookup
Cell nucleus wikipedia , lookup
Cell membrane wikipedia , lookup
Protein moonlighting wikipedia , lookup
Protein phosphorylation wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
SNARE (protein) wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Cytoplasmic streaming wikipedia , lookup
Signal transduction wikipedia , lookup
Organ-on-a-chip wikipedia , lookup
Proteolysis wikipedia , lookup
Western blot wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Cytokinesis wikipedia , lookup
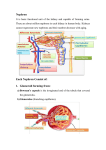
ENDOPLASMIC RETICULUM INTRODUCTION • ER is found in Eukaryotes. • In plant cells, ER occurs as - system of membrane tubules and sheets (cisternae) -distributed throughout the cytoplasm • Two different types, ER Smooth ER Rough ER Mostly tubules Cisternae, sheets • Seen as a continuous network with nuclear envelope. • Connected with the Golgi bodies. ROLES SER RER • • Protein synthesis • Post transitional modification of proteins PROTEINS • • Metabolic processes . Lipids - Steroids - Phospholipids Metabolizes the carbohydrate Detoxifying the drugs cytoplasm RER both has ribosomes end up in nucleus secreted into CM, mitochondria, integrated in peroxisomes, cytoplasm, ER, Stay back. golgi bodies, lysosomes. ER Dynamics and Mobility ER tubular extension 1) Sliding dynamics(SD) 2) Tip Attachment Complex(TAC) SD : By sliding of ER tubules along the microtubule (MT) cytoskeleton More predominant and faster. TAC : Physical interaction b/w ER Resident protein (STI M1) And MT plus end binding protein (EB1) STI M1 has the MT binding domain ( STI M1 --- ER tubule movement) ER-MT connections for ER remodeling. Rab 10- ER specific Rab GTPase – ER tubule extension along MT. used as a marker to find the new ER tubule growth. Shaping ER tubules • Reticulon (RTN) and DP1/Yop1p family members (RHD3) closely spaced TMD’s (Trans-membrane Domains) RTN and DP1/Yop1p are required for 1) Shaping of tubules (network formation) 2) Tubule formation in cos-7 cells 3) Responsible for curved edges of ER sheets and nuclear pores Overexpressed Induces constrictions results so from cisternae to tubules highly curvature membrane Three way junctions The ER network of COS-7 cells is labeled with mCherry-KDEL (red) and the junctions are labeled with Lnp1-GFP (green). Three way junction: • GTPase is the mediator for the formation of the network junctions. Sey 1p (yeast) and RHD 3 (Arabidopsis) plant homologue of atlastin Lnp1p (protein of lunapark family) – localises the ER network in yeast and mammalian cells. binds to Reticulon / Yop1p resides @ ER tubule junctions (stabilizes) ER movement in plants Actin – Myosin dependent XI-K XI-1, XI-2 a) b) ER PM APC actin ER myosin Tubule c) Fusion complex Cisternae a) Tubule growth or retraction b) Anchor points c) Tubule fusion and three way junction formation d) Tabulation versus cisternalisation (due to displacement of RTN’s) APC- Anchor Point Complex Fusion/tethering complex – RHD3 d) Vesicle formation • COP I and COP II – Vesicle coat protein occurs @ TER ER proteins –(COP II) Golgi Apparatus *TER– Transitional ER ERGIC- ER Golgi Intermediate complex TGN- Trans Golgi Network Cortical MT associated ER sites • Cortical ER network and its nearby MT’s intersect with each other. signal transduction , movement of molecules * ER membranes -- supported by Actin network – Myosin motors • This interaction is CMER’S (anchoring points) pausing them organelles meet, exchange of components MT MT CM PM ER a) b) Myosin Actin Golgi MT C ER Kinesin The ER membranes are associated with actin filaments that drive organelle movements along the membrane (e.g. Golgi; or other compartment carrying cargo (C)) with support of myosin motor proteins. Organelles may carry MTassociated proteins (e.g. kinesin) that cause pausing at C-MERs through binding to the MTs. Many plant viruses exploit C-MER’S for replication and PD targeting Summary • • • • • • • • Network of membranes Smooth and Rough ER Roles – metabolizes , protein synthesis, post transitional modification Tubule extension by sliding dynamics or TAC Reticulons and Dp1/YOP 1p involved for shaping of ER Three was junctions or network formations COP II protein involved for vesicle formation Cortical MT associated ER sites References • Pen E J , Heinlein M: Cortical microtubule-associated ER sites: organization centers of cell polarity and communication. Curr Opin Cell Biol 2013, 16:1–10. • Sparkes I, Hawes C, Frigerio L: FrontiERs: Movers and shapers of the higher plant cortical endoplasmic reticulum. Curr Opin Cell Biol 2011, 14:658–665. • Chen S, Novick P, Novick S F: ER structure and function. Curr Opin Cell Biol 2013, 25:1–6. THANK YOU



















![[pdf]](http://s1.studyres.com/store/data/008790868_1-5bdbc2c77d0606ba4b3bf1e2b9f0d1e7-150x150.png)




![[pdf]](http://s1.studyres.com/store/data/008789109_1-454f66405678d97a0b8542e58f570dae-150x150.png)


