* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Full text - PAHdb - McGill University
Tay–Sachs disease wikipedia , lookup
Public health genomics wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Human genome wikipedia , lookup
Gene expression programming wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genome (book) wikipedia , lookup
Designer baby wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Genetic code wikipedia , lookup
Koinophilia wikipedia , lookup
Genetic drift wikipedia , lookup
Medical genetics wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Genome evolution wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Genome editing wikipedia , lookup
Microsatellite wikipedia , lookup
Human genetic variation wikipedia , lookup
Oncogenomics wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Population genetics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microevolution wikipedia , lookup
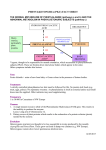
HUMAN MUTATION 21:333^344 (2003) DATABASES PAHdb 2003: What a Locus-Specific Knowledgebase Can Do Charles R. Scriver,1n Mélanie Hurtubise,1 David Konecki,2 Manyphong Phommarinh,1 Lynne Prevost,1 Heidi Erlandsen,3 Ray Stevens,3 Paula J. Waters,4 Shannon Ryan,1 David McDonald,5 and Christineh Sarkissian1 1 Departments of Human Genetics, Biology, and Pediatrics, McGill University Health Centre, Montreal, Canada; 2Medical Genetics Branch, NHGRI/NIH, Bethesda, Maryland; 3Department of Molecular Biology, The Scripps Research Institute, La Jolla, California; 4 Biochemical Genetics Laboratory Children’s and Women’s Health Center of British Columbia, Vancouver, British Columbia, Canada; 5 Department of Biological Sciences, Wichita State University, Wichita, Kansas For the PKU Special Issue PAHdb, a legacy of and resource in genetics, is a relational locus-specific database (http:// www.pahdb.mcgill.ca). It records and annotates both pathogenic alleles (n = 439, putative diseasecausing) and benign alleles (n = 41, putative untranslated polymorphisms) at the human phenylalanine hydroxylase locus (symbol PAH). Human alleles named by nucleotide number (systematic names) and their trivial names receive unique identifier numbers. The annotated gDNA sequence for PAH is typical for mammalian genes. An annotated gDNA sequence is numbered so that cDNA and gDNA sites are interconvertable. A site map for PAHdb leads to a large array of secondary data (attributes): source of the allele (submitter, publication, or population); polymorphic haplotype background; and effect of the allele as predicted by molecular modeling on the phenylalanine hydroxylase enzyme (EC 1.14.16.1) or by in vitro expression analysis. The majority (63%) of the putative pathogenic PAH alleles are point mutations causing missense in translation of which few have a primary effect on PAH enzyme kinetics. Most apparently have a secondary effect on its function through misfolding, aggregation, and intracellular degradation of the protein. Some point mutations create new splice sites. A subset of primary PAH mutations that are tetrahydrobiopterin-responsive is highlighted on a Curators’ Page. A clinical module describes the corresponding human clinical disorders (hyperphenylalaninemia [HPA] and phenylketonuria [PKU]), their inheritance, and their treatment. PAHdb contains data on the mouse gene (Pah) and on four orthologous mutant mouse models and their use (for example, in research on oral treatment of PKU with the enzyme phenylalanine ammonia lyase [EC 4.3.1.5]). Hum Mutat 21:333–344, 2003. r 2003 Wiley-Liss, Inc. KEY WORDS: Phenylalanine hydroxylase; phenylketonuria; PKU; database; genotype-phenotype; hyperphenylalaninemia; HPA DATABASES: PAH – OMIM: 261600; GenBank: NM_000277 (mRNA), U49897.1 (cDNA), AF404777 (gDNA), X51942 (mouse cDNA); Swiss-Prot: P00439 http://www.pahdb.mcgill.ca (PAHdb) INTRODUCTION So often neglected by the communities they serve, databases are a legacy in and of science [Maurer et al., 2000]. Science is an explanatory process. As a particular domain of science develops, said Stéphane Leduc cited in Keller [2002], there is first a stage of explanation in classification and nomenclature (taxonomy) of its entities. Next, there follows inquiry into mechanisms underlying the entities. The final stage is to recreate or synthesize the entities. The science of genetics recognizes mutation as both entity and mechanism. Mutation can also be created. Mutation r2003 WILEY-LISS, INC. n Correspondence to: Charles Scriver, Montreal Children’s Hospital Research Institute, McGill University Hospital Centre, A717, 2300 Tupper Street, Montreal, Quebec H3H 1P3, Canada. E-mail: [email protected] Grant sponsors: Robert McDonald Gift; Quebec Network of Applied Genetics/Fonds de la Recherche en sante¤ du Que¤ bec; Canadian Genetic Diseases Network/Networks of Centers of Excellence; Canadian Institutes for Health Research, formerly Medical Research Council of Canada. DOI 10.1002/humu.10200 Published online in Wiley InterScience (www.interscience.wiley. com). 334 SCRIVER ET AL. databases recapitulate the science in their various ways and they can recreate it in silico. Databases have thus become necessary resources in genetics. They are repositories of the vast wealth of data being gathered about individual genes and the genomes they inhabit. PAHdb is one such legacy and resource. PAHdb (http://www.pahdb.mcgill.ca) is an online relational locus-specific knowledgebase [Scriver et al., 2000] originating in, and still serving, the PAH Mutation Analysis Consortium and other communities. This report is written by co-curators who maintain PAHdb and work at five widely separated locations, yet the internet connects us. Additional databases related to the PAH gene are listed at the head of this chapter. Mutations at the human locus (symbol PAH; MIM# 261600) affect the phenylalanine hydroxylase enzyme (EC 1.14.16.1) and are a cause of the disease phenylketonuria (PKU) or related forms of hyperphenylalaninemia [Scriver and Kaufman, 2001]. PKU reflects paradigms of both transformational and translational knowledge. When the disease was first recognized in 1934, it became the fifth in a series of diseases known as ‘‘inborn errors of metabolism’’ [Garrod, 1908]. PKU thus consolidated an emerging (transformational) view that Mendelian inheritance could explain some forms of human disease. There has also emerged an understanding (translational) that the disease feature (mental retardation) of PKU was preventable through early diagnosis and treatment. As a result, our outlook on genetic disease in general began to change. With time, PKU has emerged as an explanatory prototype for human genetic disease that links gene, mutation, enzyme, metabolism, and disease effect [Scriver and Waters, 1999]. Accordingly, there has been an exceptional opportunity to sample PAH mutations in probands from populations around the world wherever newborn screening was practiced. As a result, PAH alleles and their attributes (polymorphic haplotypes and populations) have become widely known. PAHdb has thus emerged as a comprehensive and useful prototype of the locus-specific database [Claustres et al., 2002]. The origins, development, and design of PAHdb are described in an earlier report [Scriver et al., 2000]. The database is built on four core elements: 1) a unique identifier for each allele; 2) the source of the information; 3) the context of the allele (e.g., the species and name of the gene); and 4) the name of the allele. PAHdb contains entities (mutations) and annotates them with attributes. The Tables of Mutations arise from these core elements. Additional tables in the database provide information, from in vitro expression analysis, on the functional effects of mutations on enzyme integrity and function. The database also visualizes in silico how mutations map onto the 3D protein structure. A Curators’ Page highlights unscheduled topics (e.g., discovery of BH4- responsive PAH alleles) and novel data not readily handled by the existing tables. Among other options, PAHdb introduces visitors to the mouse Pah gene and a mouse model of PKU. The clinical significance of human PAH mutations is the subject of another module. A counter logs visits (>30 hits/day) and records the most recent date of curation. Table 1 is a partial site map of PAHdb. The intellectual property of PAHdb is copyrighted. Because PAH alleles are named according to a convention now widely accepted and used [Antonarakis et al., 2001], PAHdb can also be searched using appropriate tools to retrieve and transfer its alleles to an experimental ‘‘WayStation.’’ The WayStation will be linked to a comprehensive data ‘‘Warehouse,’’ thus making a repository of human genomic allelic variation [Teebi et al., 2001]. When the WayStation is established, PAH-related data could be submitted either through it or directly to PAHdb. PAH NUCLEOTIDE SEQUENCES: cDNA AND gDNA Co-Curators: David Konecki, Mélanie Hurtubise, and Manyphong Phommarinh The human phenylalanine hydroxylase (PAH) gene is embedded in the chromosomal region 12q23.2, covering 1.5 Mbp, that contains PAH itself and five other genes of known or unknown function [International Human Genome Sequencing Consortium, 2001; Venter et al., 2001]. The cDNA and full-length genomic sequences are both visible online in PAHdb. cDNA sequences have long been available [Kwok et al., 1985; Konecki et al., 1992] but the genomic sequence is recent (Konecki, D.S., unpublished; deposited in PAHdb Nov. 2001). The genomic sequence of the PAH gene and its flanking regions spans 171,266 bp with ~27 kbp of 50 untranslated region (50 UTR) upstream from the translation initiation site and ~64.5 kbp of 30 sequence downstream from the poly(A) site in the last exon (exon 13). By convention in GenBank, the first nucleotide of any sequence is numbered from its 50 end. However, to have gDNA nucleotides in register with the older cDNA sequence (which has long served PAH mutation nomenclature), the PAH gDNA has been renumbered in PAHdb: the +1 nucleotide is the adenine of the translation initiation site (ATG) in exon 1. Thus, gDNA exons, introns, and the 30 UTR have positive numbers. The 50 UTR has negative numbers. Table 2 lists the 13 exons in the PAH gene and compares gDNA with cDNA positions. Amino acid residues of the PAH protein that are conserved in mouse, rat, and human are shown on a protein sequence page directly under the cDNA heading. The 50 UTR of the human PAH gene contains cis control elements [Konecki et al., 1992]. They are PAHDB KNOWLEDGEBASE TABLE 1. A Partial Site Map of PAHdb (http://www.pahdb.mcgill.ca) Page name (alphabetic order) About Consortium Contact Clinical Haplotype InVitro analysis Molecular Mouse Mutation map Other PAHdbNewsletter Search Sequence Submission Survey 335 Content Abstract; curators’ page; list of curators List of members Information PKU for families, Link to GeneReviews Polymorphic markers in PAH, Link toALFRED See search page Structural genomics of the PAH enzyme; link to Scripps Site The mouse PKU/HPA models; mouse gene (link) Mutations displayed on the PAH gene sequence Copyright information;Various links (n = 18) Print version of PAHdb (Dec. 2001) The key page in PAHdb leading to: In vitro expression analysis Mutations Mutations (statistics) Mutation Associations Clinical Authors Genotype^phenotype correlations CpG sites and mutability pro¢le cDNA and genomic reference sequences, CpG sites, Alu sites exon^intron junctions User registration, data submission Hits counter, comments, etc. shown on both the cDNA and gDNA sequences in PAHdb. Single nucleotide polymorphic (SNP) and restriction fragment length polymorphic (RFLP) sites, currently used to create PAH polymorphic haplotypes, are annotated on the genomic sequence. Other sites at the PAH locus can be identified by using tools such as the NEBcutter (www.neb.com). Exonic regions in the human PAH gene are ~2.88% of the genomic sequence between the 50 +1 position down to the 30 poly (A) tract. Amplicon primer sequences for all 13 exons are provided on the cDNA sequence page. The shortest and longest exons are 57 bp (exon 9) and 892 bp (exon 13), respectively. The mean exon size is 170 bp. Three polyadenylation signals [AATAAA] in exon 13 are annotated on the gDNA sequence. The third site is used most frequently. The shortest and longest introns are 556 bp (intron 10) and 17,874 bp (intron 2), respectively. Intron 3 is 17,187 bp in length. The mean intron size is 6390 bp. These dimensions are typical for mammalian genes. The PAH genomic sequence consists of 40.7% GC, slightly above the modal value (37–38%) for human genes. RepeatMasker analysis shows the density of interspersed repeats to be 42.2% in the PAH gene, a value typical for a mammalian gene. RepeatMasker data and a table of DNA variations, mostly SNPs, are shown in PAHdb. Repetitive DNA is often the cause of large genomic deletions and duplications [Antonarakis et al., 2001]. A search for Alu repeat elements in the PAH gene was performed using NCBI BLAST (www.ncbi.nlm.nih.gov/blast/). Alu repeat elements are annotated on the gDNA sequence. Intron 2 has a 99% nucleotide identity with the Alu repeat element between bp 17,273 and bp 17,546, which could possibly coincide with the 50 deletion found in a PKU family from Scotland [Sullivan et al., 1985]. Putative Alu repeats are highlighted on the PAH genomic sequence and CpG dinucleotide sites (n = 1198) are annotated since these are potential sites for mutation in the gene. ALLELIC VARIATION Co-Curators: Lynne Prevost, Manyphong Phommarinh, and Charles Scriver PAHdb lists 439 human mutations classified as potentially disease-causing [Cotton and Scriver, 1998]. Another 25 alleles are considered to be non-pathogenic polymorphisms (but see K274E in Expectations Confounded section, below) and there are at least 16 alleles in the known STR and VNTR regions of the gene. The PAH genomic sequence is likely to contain many as yet unrecognized polymorphisms. Pathogenic alleles These are either displayed on a cartoon of the gene, listed as entities by name in sequential order (50 to 30 ) in a core table, or listed in two expanded tables containing attributes (e.g., unique identifier, associations with polymorphic haplotypes and populations, or source of the data). Polymorphic alleles These are either annotated on the DNA sequences, shown in a pictorial configuration on a cartoon of the gene, or tabulated as configurations describing core PAH haplotypes. There is a link to the ALFRED database (http://alfred.med.yale.edu) where known 336 SCRIVER ET AL. TABLE 2. ConversionTable of cDNA Exon 1 2 3 4 5 6 7 8 9 10 11 12 13a Exon Annotations to Genomic Nucleotide Numbers cDNA gDNA Intron Sequence 1^60 61^168 169^352 353^441 442^509 510^706 707^842 843^912 913^969 970^1065 1066^1199 1200^1315 1316^1359 1^60 4233^4340 22215^22398 39586^39674 50550^50617 61890^62086 64272^64407 65466^65535 70273^70329 72793^72888 73445^73578 76709^76824 78006^78897 1 2 3 4 5 6 7 8 9 10 11 12 61^4232 4341^22214 22399^39585 39675^50549 50618^61889 62087^64271 64408^65465 65536^70272 70330^72792 72889^73444 73579^76708 76825^78005 a Exon 13 cDNA covers only the translated and termination codons; exon 13 gDNA extends 30 to include polyadenylation signals. PAH polymorphisms are lodged for the study of modern human populations [Kidd et al., 2000]. Haplotypes Because two RFLP sites (EcoRI and EcoRV) have yet to yield to PCR, the use of mini-haplotypes (STR and VNTR alleles only) has been recommended [Zschocke et al., 1995]. A more complex and informative new set of haplotype configurations has also been developed [Zschocke and Hoffmann, 1999]. The polymorphic PAH haplotypes are useful because they facilitate the population genetics of PAH. They also help to delineate identity, either by descent or by state, of particular alleles (e.g., R408W [John et al., 1990; Byck et al., 1994]). Haplotype markers have also contributed to the discovery of large deletions in the PAH gene [Zschocke et al., 1999; Gable et al., 2003]. MutationTypes Pathogenic PAH alleles, as predicted from the DNA sequence, comprise point mutations causing missense in translation (63%), most of which are likely to affect protein folding and assembly leading to secondary effects on enzyme function (see In Vitro Expression Analysis, below). Other PAH mutation types include: small deletions, 13%; confirmed splice, 11%; silent (putative), 7%; nonsense, 5%; small insertions, 1%; large deletions are rare. Some PAH point mutations, previously classified as missense or silent, generate alternative splice sequences (see Expectations Confounded section, below). A web site (http://exon.cshl.edu/ESE/) provides tools to assist in the recognition of new splice donor or acceptor nucleotide sequences generated by such point mutations. Frequencies The absolute aggregate frequency of pathogenic PAH alleles rarely exceeds 0.01 in human populations and only rarely does the absolute frequency of any particular allele exceed 0.01 (see Table in Zschocke [2003] for source data). The relative frequencies of PAH alleles follow a recognizable pattern for human locus-specific mutations [Weiss, 1996]. Regardless of the number of pathogenic alleles at the locus, only a few (o10) represent the major portion (75%) of those in the population. The majority of probands (~75%) have compound mutant PAH genotypes. Population Associations The history of human populations contains the history of their alleles and vice versa. PAH alleles have contributed to an understanding of the genetic configurations typical of European populations [Cavalli-Sforza and Piazza, 1993], for example: Germany [Zschocke and Hoffmann, 1999]; Denmark [Guldberg et al., 1993a]; Sicily [Guldberg et al., 1993b]; and, possibly, for their ancient origins in some populations [Zschocke et al., 1997]. The global European profiles of those PAH mutations (n=29) that exceed 3% individual relative frequency in at least two different populations is now known [Zschocke, 2003]. The effects of range expansion, genetic drift, and founders on genetic structure of overseas populations is illustrated in detail by alleles at the PAH locus, for example: Iceland [Guldberg et al., 1997]; Quebec [Carter et al., 1998; Scriver, 2001]; and the USA [Guldberg et al., 1996]. Identity by State or by Descent? Several PAH mutations, identical by state in their nucleotide change, occur on two or more different haplotype backgrounds. For example R408W, the most prevalent pathogenic PAH allele, occurs prominently on haplotypes 1 and 2 in European-derived populations [John et al., 1990; Treacy et al., 1993; Eisensmith et al., 1995; Tighe et al., 2003]. Because a CpG dinucleotide is involved, R408W has the potential to be a recurrent mutation and identical only by state on haplotypes 1 and 2 [Byck et al., 1994; Tighe et al., 2003]. R408W also appears on haplotypes PAHDB KNOWLEDGEBASE of the filter paper screen blood spots [Altland et al., 1982], that the PAH locus is unusually mutable or will serve as a sentinel for human genome mutability. PAH c.1066-11 g->a (IVS10nt-11)*: II % 2 Haplotype 10 * mutation site - + - + + - * - - IBD + + + - * - - IBD + + + - * - - IBD • 30-90 6 + - v.rare 39 - + • 25 + - - + 36 34 + - + - 2 (Italy) 9 + + - + + - * - - (IBD) (Spanish Gypsies) rare + + - * * - + 337 (IBD) ? A scheme to explain the presence of a major PAH mutation (c.1066^11g4a IVS10nt-11, PKU-causing) on six di¡erent haplotypes. Relative frequencies of the di¡erent mutant haplotypes are shown in the left hand column. With the possible exception of a recurrent mutation, the putative mechanisms are indicated on haplotype 9. All other forms are identical by descent from the haplotype 6 con¢guration. Intragenic recombination leads to the mutation on H39 or H10 and RFLP site mutations put it on H34 or H36. Figure from Scriver and Kaufman [2001], with permission. FIGURE 1. other than 1 and 2. In these instances, the R408W allele could be identical by descent because mutation at a polymorphic PAH marker is the more likely explanation for the variant haplotype association. Eleven different PAH mutations in Germany, each occurring on more than one haplotype, have been examined for evidence of identity by state, recurrence, or an alternative explanation [Zschocke and Hoffmann, 1999]. Figure 1 illustrates possible genealogies (mutation lineages) for the IVS10nt-11 mutation (c.1066–11g>a) which is prevalent in southern European populations. Some pathogenic PAH mutations, including R408W and IVS10nt-11, occur at quite high frequencies in particular populations. Selective advantage in the heterozygote is one possible explanation and the case for this hypothesis is being argued elsewhere in this issue [Krawczak and Zschocke, 2003]. Recurrent mutation and founder effect/genetic drift are alternative explanations. De Novo Alleles These are rare occurrences in the PKU and related phenotypes. M1I and IVS3nt-6 have each been reported once as de novo alleles, once from Norway [Eiken et al., 1996] and E76G has been reported once from Taiwan [Chen et al., 2002]. Another occurrence (unnamed) is reported once from Southern Germany [Aulehla-Scholz and Heilbronner, 2003]. The apparent rarity of de novo alleles could be a feature of ascertainment. Such alleles will not be recognized if the parental alleles have not also been analyzed, a practice that is not uniform. Accordingly, one cannot say that these few de novo alleles have revealed mutation hotspots. Nor does it seem, from the monitoring Insertion/Deletion Mutations Small deletions or insertions occur in the PAH gene. Large deletions affecting coding regions are rare. Some deletions involve a single exon. More frequently, multiple exons are affected. (For a literature review, see Gable et al. [2003]). The relative frequency of large PAH gene deletions is apparently low (o1%) but this may reflect poor ascertainment. Most of the large recognized deletions were discovered when a disease-causing point mutation could not be identified by current methods of mutation analysis or when a haplotype configuration lacked one or more of its polymorphic alleles. Large insertions or gene duplications have not yet been reported. 50 UTR Alleles Polymorphic alleles occur in the 50 UTR of the PAH gene that are without apparent phenotypic effect [Svensson et al., 1993]. A large deletion (3767bp, nt-4173 to –407 in the Konecki sequence) has been identified [Chen et al., 2002]. The deletion removes a liver-specific enhancer element harboring a major hepatocyte nuclear factor-1-binding site. It severely impairs PAH gene transcriptional activity conferring functional hemizygosity on the affected proband. Expectations Confounded Several PAH point mutations are mentioned here to show how they challenged easy interpretation. Experimental evidence defied the sequence-based predictions that initially categorized them as either disease-causing or silent. These mutations highlight the relevance of using a combination of RNA processing analysis, in vitro expression analysis, molecular modeling, and other means [Terp et al., 2002], to identify the mechanism of mutation effect. The prevalent Y204C (c.611A>G) allele in exon 6 is associated with PKU, yet when the protein is expressed conventionally in vitro, the mutation appears non-pathogenic. RT-PCR analysis of illegitimate transcripts, however, revealed the creation of a novel splice site by this allele causing loss of protein function. The mutation was therefore given a new trivial name: E6nt–96A>G [Ellingsen et al., 1997]. Several alleles, including Q304Q (c.912G>A) [Guldberg et al., 1996], T323T (c.969A>G), K398K (c.1194A>G) [Zschocke and Hoffmann, 1999], and V399V (c.1197A>T) [Chao et al., 2001], initially called silent, are either shown to, or are likely to, promote exon skipping. In each case the adjacent nucleotide sequence becomes an important context to help explain the mutation 338 SCRIVER ET AL. effect [Ellingsen et al., 1999]. Prediction of such effects from consensus sequence algorithms (viz http://exon.chsl.edu/ESE/) is still imperfect and experimental studies on RNA processing are still necessary. A growing awareness [Cartegni et al., 2002] that exonic alleles in human genes can affect pre-mRNA splicing clearly encompasses the PAH gene where P281L, R408Q, G272X, and Y356X (and perhaps many other apparent missense, nonsense, and silent mutations) are associated with exon skipping [Ellingsen et al., 1999]. The K274E (c.820A>G) allele [Gjetting et al., 2001a] initially appeared to be a cause of PKU. However, in vitro expression analysis showed normal enzyme function. Further genotype analysis at the PAH locus, followed by in vitro expression, revealed that a rare pathogenic allele (I318T, c.953T>C) was also being transmitted in cis with K274E. The latter is a polymorphism (frequency ~0.04) in the source population (African–American). PAHdb lists variant PAH alleles paired in cis (see cis Mutation Table in PAHdb). Three of those pairs, submitted earlier by C. Aulehla-Scholz directly to PAHdb, are now published [AulehlaScholz and Heilbronner, 2003]. BH4-responsive PAH Alleles Following the discovery of patients with a variant form of hyperphenylalaninemia responsive to pharmacological doses of tetrahydrobiopterin [Kure et al., 1999], at least 21 corresponding primary mutations in the PAH gene have been proposed as BH4-responsive. This new phenotype is listed in the Mutation Tables in PAHdb and illustrated on a Curators’ page showing where the putative alleles map on the protein. Detailed molecular explanations for BH4-responsiveness have been proposed [Erlandsen and Stevens, 2001]. MOLECULAR MODELING Co-Curators: Heidi Erlandsen and Ray Stevens PAHdb hosts a module (http://www.pahdb.mcgill. ca/molecular.html) for molecular modeling of the PAH protein. It shows the derived three-dimensional structure of the enzyme (PAH) (Fig. 2) and predicts the structural effects of some currently known PAH mutations. The human enzyme exists in a pH-dependent equilibrium of homo-tetramers and homo-dimers [Martinez et al., 1995] and like the two other human aromatic amino acid hydroxylases (tyrosine hydroxylase and tryptophan hydroxylase), it has three domains: an N-terminal regulatory domain (residues 1–142); a catalytic domain (residues 143–410); and a C-terminal tetramerization domain (residues 411– 452). X-ray crystallography has been used previously to determine the 3D structures of truncated forms of phenylalanine hydroxylase [Erlandsen et al., 1997; Fusetti et al., 1998; Kobe et al., 1999], but because it is difficult to crystallize the full-length PAH subunit, no primary full-length tetrameric PAH structure has yet been analyzed. However, two truncated forms have been characterized. These include a dimeric form containing regulatory and catalytic domains [Kobe et al., 1999] and a tetrameric form containing catalytic and tetramerization domains [Fusetti et al., 1998]. From these two structures, and from a higherresolution dimeric double-truncated form of the PAH enzyme [Erlandsen et al., 1997], it has been possible to derive a composite full-length structural model [Erlandsen and Stevens, 1999] (Fig. 2). The regulatory domain of PAH contains an a–b sandwich with an interlocking double bab motif. The N-terminal autoregulatory sequence (ARS, residues 19–33) extends over the active site in the catalytic domain [Erlandsen et al., 1997]. The tetramerization domain contains two b-strands forming a b-ribbon and a 40 Å long a-helix. The four a-helices (one from each monomer) pack into a tight anti-parallel coiledcoil motif in the center of the tetramer structure [Fusetti et al., 1998]. The region containing the catalytic domain has a basket-like arrangement with a total of 13 a-helices and 8 b-strands. The active site, located in the center of the catalytic domain, is a pocket that is 13 Å deep and 10 Å wide [Erlandsen et al., 1997]. Adjacent to the active site is a channel that is 16 Å long and 8 Å wide by which substrate may access the active site [Anderson and Flatmark, 2001]. As isolated, PAH crystals contain an active site Fe(III) iron atom [Martinez et al., 1991; Erlandsen et al., 1997] located 10 Å below the surface of the protein on the floor of the active site at the intersection of the channel and the active site pocket. The Fe(III) atom is coordinated to H 285, H 290, and one oxygen atom in E 330. The cofactor (6R)-L-erythro-5,6,7,8-tetrahydrobiopterin (BH4) and the L-Phenylalanine substrate bind close to the iron at the active site [Erlandsen et al., 2000; Anderson and Flatmark, 2001; Anderson et al., 2002]. A set of 269 PAH missense mutations (see The PAH Mutation Analysis Consortium Newsletter, December 2001, online in PAHdb), 23 nonsense mutations, and 10 silent mutations were used for molecular interpretation. Most missense mutations map onto the region between and including exon 5 (PAH residue 148) and exon 12 (PAH residue 438): 57 of these mutations are located in the regulatory domain sequence (residues 1–142); 231 mutations target the catalytic domain (residues 143-410); and 14 are located in the tetramerization domain sequence (residues 411–452) (Fig. 3). A summary of genotype/ phenotype/structural interpretations for these mutations has been published in Erlandsen and Stevens [1999]. PAHDB KNOWLEDGEBASE 339 FIGURE 2. Three views of the tetrameric form of the composite model of phenylalanine hydroxylase. The regulatory domain (residues 19-142) is colored orange, the catalytic domain (residues 143^410) is colored gray, and the tetramerization domain is colored blue.The active site iron is shown as a yellow sphere. A: Front view; B: side view, seen in the plane of the paper along the x-axis as compared to A; C: side view, seen in the plane of the paper along y-axis as compared to A. Figure from Erlandsen and Stevens [2001], with permission. Ca-trace of one monomer of the full-length model of phenylalanine hydroxylase.The trace is colored yellow for residues not associated with any PKU mutations, and the active site iron is shown for reference as a green sphere.The region colored in red consists of residues that have PKU mutations associated with them. The regions colored blue have residues that show a high predicted frequency of mutation, and the regions colored green have residues that show a high calculated frequency of mutation (PKU database).The four residues colored purple are the residues with both the calculated and predicted frequency of mutation being high (Arg158, Arg252, Arg261, and Arg408). Figure from Erlandsen and Stevens [1999], with permission. FIGURE 3. The structural information for PAH has helped to formulate predictions [Erlandsen and Stevens, 1999; Kobe et al., 1999] about the likely effects of yet unclassified or newly discovered missense mutations. Furthermore, with the aid of recent crystal structure analysis using cofactor and substrate analogs bound at 340 SCRIVER ET AL. the active site [Erlandsen et al., 2000; Anderson and Flatmark, 2001; Anderson et al., 2002], the newly discovered BH4-responsive PKU/HPA genotypes can be mapped onto the PAH structure to seek molecular explanations for their BH4-dependent clinical responses [Erlandsen and Stevens, 2001]. lationships [Okano et al., 1991] and, in broad terms, such correlations exist [Kayaalp et al., 1997; Guldberg et al., 1998; Desviat et al., 1999]. There are, however, significant exceptions. Thus, IVE data are not always robust predictors of the in vivo phenotype and there are some partial explanations for the inconsistencies [Scriver and Waters, 1999]. IN VITRO EXPRESSION ANALYSIS Co-Curator: Paula J. Waters PAHdb contains data on in vitro expression analysis (IVE) of 81 different naturally occurring human mutations (227 individual records, see Table in PAHdb under IVE-Human). It also contains information on 12 artificially created mutations in the human nucleotide sequence (13 individual records, IVEArtificial) and on 41 artificially created mutations in the rat Pah gene (42 records, IVE-Rat). The search option on the Home Page leads to a search page containing all the relevant links. IVE-Human is listed under User-defined queries. IVE-Rat and IVE-Artificial are listed under pre-queried data. Another link on this page leads to IVE-Commentary that describes how to use and interpret the IVE Human table. There are three reasons to put IVE data on PAHdb: 1) to provide evidence that the mutation alters protein function; 2) to document the severity of mutation effect; and 3) to describe the mechanism of effect. Only 81 different human PAH mutations, from the more than 400 known mutations, have yet been studied by IVE. Two reviews [Waters et al., 1998; Waters, 2003] discuss the advantages and disadvantages of the different systems that are available for mutation analysis by IVE and highlight key overall findings from IVE. Prediction of mutation effects using IVE data is complemented by the analysis of PAH structure by both molecular modeling [Erlandsen and Stevens, 1999] and spectroscopy [Teigen et al., 1999]. Most PAH mutations are predicted from the DNA sequence to be missense alleles. The corresponding amino acid substitutions frequently affect folding and assembly of the protein, leading to decreased cellular PAH activity as a result of the enzyme’s degradation. Only a minority of the disease-causing mutations directly affect the kinetic properties of the enzyme (e.g., D143G [Knappskog et al., 1996]). Some mutations appear to have combined effects on both sutrate or cofactor binding [Leandro et al., 2000; Gjetting et al., 2001b] and on protein folding [Waters et al., 2000; Gamez et al., 2000]. The BH4-responsive PAH mutations await detailed analysis by IVE to determine whether they are kinetic mutants affecting BH4 binding or alleles that affect protein folding and assembly where BH4 at high concentration might act as a chemical chaperone. Expression analysis of PAH mutations also has the potential to shed light on genotype/phenotype re- CLINICAL RESOURCES Co-Curator: Shannon Ryan PAHdb provides visitors with a ‘‘Phenylketonuria Resource Booklet for Families’’ and an overview of the treatment for PKU. It is intended only as a supplementary resource for patients, family members, and other interested persons who wish to become familiar with PKU and the associated vocabulary. PAHdb does not replace necessary patient–care-giver relationships in optimal treatment of PKU. PAHdb also provides links to other sites containing information on PKU (e.g., www.espku.org). Access to resources in languages other than English is also provided (e.g., French, German, Spanish). Content of these sites varies. Some sites give details of everyday life with PKU and are intended for the lay person. Others are intended for the health-care professional. GeneClinics/GeneTests, a peer-reviewed internet resource (www.geneclinics.org), contains a document authored by the PAHdb co-curators. The latest version includes information, for example, on BH4responsive PAH phenotypes and on current diagnostic and molecular tests. MOUSE MODELS Co-Curators: Christineh N. Sarkissian and David McDonald PAHdb describes mouse lines carrying mutations in the orthologous Pah gene (GenBank accession # X51942, cDNA) and their use in the development of a potential new therapy for PKU with phenylalanine ammonia lyase enzyme (EC 4.3.1.5) [Shedlovsky et al., 1993; Sarkissian et al., 2000; Sarkissian et al., 1999]. Mouse and human PAH enzymes show homology and evolutionary divergence [Ledley et al., 1990]. Conserved amino acid residues (92% of the sequence) are indicated in the database. The PAHdb mouse page contains a list of published articles, mouse links, descriptions of ongoing research, and animal husbandry. Three mouse Pah mutations have been artificially created: Pahenu1 [McDonald et al., 1990]; Pahenu2; and Pahenu3 [Shedlovsky et al., 1993]. The Pahenu3 allele is now historic. With the currently existing mutant alleles, mouse phenotypes can be produced that range in severity from mild HPA (Pahenu1 homozygosity) to a more pronounced HPA (Pahenu1/Pahenu2 PAHDB KNOWLEDGEBASE compound heterozygosity) to classical PKU (Pahenu2 homozygosity). The Pahenu1 allele is a c.364T>C missense mutation (V106A) in exon 3. The V106 residue is conserved among mammalian species [McDonald and Charlton, 1997]. The allele has no human counterpart. The mutation probably affects protein folding, stability, and assembly of the enzyme, as is the case with neighboring human missense alleles (A104D, S110L) in the regulatory domain of the PAH subunit [Erlandsen and Stevens, 1999]. The Pahenu2 allele (c.835T>C) is a missense mutation (F263S) in exon 7 [McDonald and Charlton, 1997]. This allele has a human counterpart (F263L). Residue F263 is located close to the Fe atom in the active site of PAH enzyme and a substitution here is most likely detrimental to activity (ligand binding) as well as proper folding of the protein [Anderson et al., 2002]. The substitution is also directly adjacent to amino acids involved in pterin-binding on the human enzyme [Erlandsen and Stevens, 1999; Erlandsen et al., 2000]. Although corresponding phenylalanine residues are affected in human and mouse, the cause of HPA would be different because of the different (serine and leucine) substitutions. Either substitution results in a severe clinical phenotype in the particular host, reaffirming the importance of this residue in the structure/function of the PAH protein. The Pahenu1/Pahenu2 compound heterozygote strain was developed [Sarkissian et al., 2000] as a model of most human PKU-causing genotypes which are heteroallelic compounds [Kayaalp et al., 1997]. The metabolic phenotype in this mouse is intermediate to that of Pahenu1 and Pahenu2 homozygous mice [Sarkissian et al., 2000]. Enzyme activity in the heteroallelic mouse is less than the predicted average activity of Pahenu1 and Pahenu2 homozygotes, suggesting that negative complementation may exist in the heteroallelic tetrameric enzyme. The archived Pahenu3 mutation (c.1126+2t>g transversion at the exon 11-intron 11 boundary) activates two cryptic splice donor sites and has a complicated effect [Haefele et al., 2001]. There is no known corresponding human allele. As described in PAHdb, these PKU mouse models have thus far proven their utility providing, for example, insights into: teratology among the progeny of mutant females; learning and behavior; brain metabolic pools; short- and long-term effects of neurochemical alterations; cerebral protein synthesis; brain myelination and CNS glial cell plasticity; cognitive defects; and experimental approaches to treatment of PKU. DISCUSSION The Curators of PAHdb welcome suggestions for improvements and notifications of errors or omissions 341 in its content. The current version of PAHdb has been transferred onto a COMPAQ server running Red Hat Linux version 7.3. This server also hosts four other locus-specific mutation databases and the catalogue for the Repository of Mutant Human Cell Strains. It also carries the WayStation [Teebi et al., 2001] for the project to create a repository of all human genomic allelic variation, notably its pathogenic alleles. Data can be submitted to PAHdb by registered users. The submission script, until recently difficult to use, has been improved. The web interface, running on Apache webserver, and the PAHdb software are also being improved. PAHdb can accommodate population-related mutation reports and the data from corresponding articles in this issue of the Journal will appear in PAHdb. Data in the two reports documenting BH4-responsive PAH alleles in this issue will also be accommodated. Their data will be linked to the BH4 database (www.bh4.org). Large deletions are rare in the PAH gene (see Gable et al. [2003] in this issue of the Journal). Whether an allele actually affects phenotype, or whether the phenotype can be predicted from the genotype, remains an important general inquiry—no less so at the PAH locus. The present guidelines [Cotton and Scriver, 1998], useful as they are, are no longer sufficient. We illustrate the problem here with several examples. Because in vitro expression analysis can provide data useful to this inquiry, PAHdb devotes a large module to them. However, such data are not available for the majority of PAH alleles. Accordingly, molecular modeling is another feature of PAHdb because it may provide insights on the likelihood of the mutation having an effect on the enzyme and thus on phenotype. In this initiative, PAHdb serves as a prototype for other Mendelian diseases. Data in PAHdb were used in another approach to the inquiry about causation [Terp et al., 2002], namely, an assessment of the functional significance of an amino acid substitution as it might be understood from its biophysical properties. The mechanisms by which mutations affect cellular function have always been of fundamental interest. PAHdb acknowledges investigations which reveal that many missense alleles seem to produce their effect through misfolding, causing instability and aggregation of the PAH monomer and/or aberrant oligomerisation (see Waters [2003] in this issue of the Journal). If misfolding is a prevalent pathogenic mechanism in Mendelian disease, might there be diffusable chemical chaperones to play therapeutic roles? Is tetrahydrobiopterin playing such a role for some of BH4responsive PAH alleles? PKU is a relatively prevalent variant Mendelian phenotype. Only a few PAH alleles achieve high relative frequencies in distinct populations and the mechanisms underlying prevalence and non-random 342 SCRIVER ET AL. distribution of PAH alleles are of abiding interest [Scriver et al., 1996]. The hypothesis of selective advantage in the heterozygote (over-dominant selection) has a certain appeal [Krawczak and Zschocke, 2003]. The debate for and against selective advantage continues to be aired and is not yet closed. ACKNOWLEDGEMENTS The March of Dimes, HUGO, and the Human Genome Variation Society have been supportive forces. The present curators thank colleagues active at earlier stages of PAHdb: David Côté, Ken Hechtman, Liem Hoang, Jaroslav Novak, Piotr Nowacki, Saeed Teebi, and Ziggy Zeng. The authors thank Johannes Zschocke and Saeed Teebi for their comments on the manuscript for this article. REFERENCES Altland K, Kaempher M, Forssbohm M, Werner W. 1982. Monitoring for changing mutations rates using blood samples submitted for PKU screening. In: Bonne-Tamir B, Cohen T, Goodman RM, editors. Human genetics, part A: the unfolding genome. New York: Alan R. Liss, Inc. pp 277–287. Anderson OA, Flatmark T. 2001. High resolution crystal structures of the catalytic domain of human phenylalanine hydroxylase in the catalytically active Fe(II) form and binary complex with tetrahydrobiopterin. J Mol Biol 314:279–291. Anderson OA, Flatmark T, Hough E. 2002. Crystal structure of the ternary complex of the catalytic domain of human phenylalanine hydroxylase with tetrahydrobiopterin and 3(2-thienyl)-L-alanine, and its implications for the mechanisms of catalysis and substrate activation. J Mol Biol 320:1095–1108. Antonarakis SE, Krawczak M, Cooper DN. 2001. The nature and mechanisms of human gene mutation. In: Scriver CR, Beaudet AL, Sly WS, Valle D, Childs B, Kinzler KW, Vogelstein B, editors. The metabolic and molecular bases of inherited disease. New York: McGraw-Hill. pp 343–377. Aulehla-Scholz C, Heilbronner H. 2003. Mutational spectrum in German patients with phenylalanine hydroxylase deficiency. Hum Mutat 21:399–400. Byck S, Morgan K, Tyfield L, Dworniczak B, Scriver CR. 1994. Evidence for origin, by recurrent mutation, of the phenylalanine hydroxylase R408W mutation on two haplotypes in European and Quebec populations. Hum Mol Genet 3:1675–1677. Cartegni L, Chew SL, Krainer AR. 2002. Listening to silence and understanding nonsense: exonic mutations that affect splicing. Nat Rev Genet 3:285–298. Carter KC, Byck S, Waters PJ, Richards B, Nowacki PM, Laframboise R, Lambert M, Treacy E, Scriver CR. 1998. Mutation at the phenylalanine hydroxylase gene (PAH) and its use to document population genetic variation: the Quebec experience. Eur J Hum Genet 6:61–70. Cavalli-Sforza LL, Piazza A. 1993. Human genomic diversity in Europe: a summary of recent research and prospects for the future. Eur J Hum Genet 1:3–18. Chao HK, Hsiao KJ, Su, TS. 2001. A silent mutation induces exon skipping in the phenylalanine hydroxylase gene in phenylketonuria. Hum Genet 108:14–19. Chen KJ, Chao HK, Hsiao KJ, Su TS. 2002. Identification and characterization of a novel liver-specific enhancer of the human phenylalanine hydroxylase gene. Hum Genet 110:235–243. Claustres M, Horaitis O, Vanevski M, Cotton RGH. 2002. Time for a unified system of mutation description and reporting: a review of locus-specific mutation databases. Genome Res 12:680–688. Cotton RGH, Scriver CR. 1998. Proof of ‘‘disease-causing’’ mutation. Hum Mutat 12:1–3. Desviat LR, Perez B, Gamez A, Sanchez A, Garcia MJ, Martinez-Padro M, Marchante C, Boveda D, Baldellou A, Arena J, Sanjurjo P, Fernandez A, Cabello ML, Ugarte M. 1999. Genetic and phenotypic aspects of phenylalanine hydroxylase deficiency in Spain: molecular survey by regions. Eur J Hum Genet 386–392. Eiken HG, Knappskog PM, Boman H, Thune KS, Kaada G, Motzfeldt K, Apold J. 1996. Relative frequency, heterogeneity and geographic clustering of PKU mutations in Norway. Eur J Hum Genet 4:205–213. Eisensmith RC, Goltzov AA, O’Neill C, Tyfield LA, Schwartz EI, Kuzmin AI, Baranovskaya SS, Tsukerman GL, Treacy E, Scriver CR, Guttler F, Gulberg P, Eiken HG, Apold J, Svensson E, Naughten E, Cahalane SF, Croke DT, Cockburn F, Woo SLC. 1995. Recurrence of the R408W mutation in the phenylalanine hydroxylase locus in Europeans. Am J Hum Genet 56:278–286. Ellingsen S, Knappskog PM, Eiken HG. 1997. Phenylketonuria splice mutation (EXON6nt–96A>g) masquerading as missense mutation (Y204C). Hum Mutat 9:88–90. Ellingsen SP, Knappskog PM, Apold J, Eiken HG. 1999. Diverse PAH transcripts in lymphocytes of PKU patients with putative nonsense (G272X, Y356X) and missense (P281L, R408Q) mutations. FEBS Lett 457:505–508. Erlandsen H, Fusetti F, Martinez A, Hough E, Flatmark T, Stevens RC. 1997. Crystal structure of the catalytic domain of human phenylalanine hydroxylase reveals the structural basis for phenylketonuria. Nat Struct Biol 4:995–1000. Erlandsen H, Stevens RC. 1999. The structural basis of phenylketonuria. Mol Genet Metab 68:103–125. Erlandsen H, Bjorgo E, Flatmark T, Stevens RC. 2000. Crystal structure and site-specific mutagenesis of pterin-bound human phenylalanine hydroxylase. Biochemistry 39: 2208–2217. Erlandsen H, Stevens RC. 2001. A structural hypothesis for BH4 responsiveness in patients with mild forms of hyperphenylalaninemia and phenylketonuria. J Inherit Metab Dis 24:213–230. Fusetti F, Erlandsen H, Flatmark T, Stevens RC. 1998. Structure of tetrameric human phenylalanine hydroxylase and its implications for phenylketonuria. J Biol Chem 273:16962–16967. Gable M, Williams M, Stephenson A, Okano Y, Ring S, Hurtubise M, Tyfield L. 2003. Comparative multiplex dosage analysis detects whole exon deletions at the phenylalanine hydroxylase locus. Hum Mutat 21:379–386. PAHDB KNOWLEDGEBASE Gamez A, Perez AB, Ugarte M, Desviat LR. 2000. Expression analysis of phenylketonuria mutations. J Biol Chem 275:29737–29742. Garrod AE. 1908. Inborn Errors of Metabolism. Oxford: Oxford University Press. Gjetting T, Romstad A, Haavik J, Knappskog PM, Acosta AX, Silvia WA Jr, Zago MA, Guldberg P, Guttler F. 2001a. A phenylalanine hydroxylase amino acid polymorphism with implications for molecular diagnostics. Mol Genet Metab 73:280–284. Gjetting T, Petersen M, Guldberg P, Guttler F. 2001b. Missense mutations in the N-terminal domain of human phenylalanine hydroxylase interfere with binding of regulatory phenylalanine. Am J Hum Genet 68:1353–1360. Guldberg P, Henriksen KF, Guttler F. 1993a. Molecular analysis of phenylketonuria in Denmark: 99% of the mutations detected by denaturing gradient gel electrophoresis. Genomics 17:141–146. Guldberg P, Romano V, Ceratto N, Bosco P, Ciuna M, Indelicato A, Mollica F, Meli C, Giovannini M, Riva E, Biasucci G, Henricksen KF, Guttler F. 1993b. Mutational spectrum of phenylalanine hydroxylase deficiency in Sicily - Implications for diagnosis of hyperphenylalaninemia in Southern Europe. Hum Mol Genet 2: 1703–1707. Guldberg P, Levy HL, Hanley WB, Koch R, Matalon R, Rouse BM, Trefz F, De la Cruz F, Henriksen KF, Guttler F. 1996. Phenylalanine hydroxylase gene mutations in the United States: report from the maternal PKU collaborative study. Am J Hum Genet 59:84–94. Guldberg P, Zschocke J, Dagbjartsson K, Friss Henrisksen K, Guttler F. 1997. A molecular survey of phenylketonuria in Iceland: identification of a founding mutation and evidence of predominant Norse settlement. Eur J Hum Genet 5: 376–381. Guldberg P, Rey F, Zschocke J, Romano C, Francois B, Michiels L, Ullrich K, Hoffman GF, Burgard P, Schmidt H, Meli C, Riva E, Dianzani I, Ponzone A, Rey J, Guttler F. 1998. A European multicenter study of phenylalanine hydroxylase deficiency: classification of 105 mutations and a general system for genotype-based prediction of metabolic phenotype. Am J Hum Genet 63:71–79. Haefele MJ, White G, McDonald JD. 2001. Characterization of the mouse phenylalanine hydroxylase mutation Pahenu3. Mol Genet Metab 72:27–30. International Human Genome Sequencing Consortium. 2001. Initial sequencing and analysis of the human genome. Nature 409:860–921. John SWM, Rozen R, Scriver CR, Laframboise R, Laberge C. 1990. Recurrent mutation, gene conversion, or recombination at the human phenylalanine hydroxylase locus: evidence in French-Canadians and a catalog of mutations. Am J Hum Genet 46:970–974. Kayaalp E, Treacy E, Waters PJ, Byck S, Nowacki P, Scriver CR. 1997. Human phenylalanine hydroxylase mutations and hyperphenylalaninemia phenotypes: a metanalysis of genotype-phenotype correlations. Am J Hum Genet 61: 1309–1317. 343 Keller M. 2002. Making sense of life. Explaining biological development with models, metaphors and machines. Cambridge, MA: Harvard University Press. Kidd JR, Pakstis AJ, Zhao H, Lu RB, Okonofua E, Odunsi A, Frigorenko E, Bonne-Tamir B, Friedlaender J, Schulz LO, Parnas J, Kidd KK. 2000. Haplotypes and linkage disequilibrium at the phenylalanine hydroxylase locus, PAH, in a global representation of populations. Am J Hum Genet 66:1882–1899. Knappskog PM, Eiken HG, Martinez A, Bruland O, Apold J, Flatmark T. 1996. PKU mutation (D143G) associated with an apparent high residual enzyme activity: expression of a kinetic variant form of phenylalanine hydroxylase in three different systems. Hum Mutat 8:236–246. Kobe B, Jennings IG, House CM, Mitchell BJ, Goodwill KE, Santarsiero BD, Stevens RC, Cotton RGH, Kemp BE. 1999. Structural basis of autoregulation of phenylalanine. Nat Struct Biol 6:442–448. Konecki DS, Wang Y, Trefz FK, Lichter-Konecki U, Woo SLC. 1992. Structural characterization of the 50 region of the human phenylalanine hydroxylase gene. Biochemistry 31:8363–8368. Krawczak M, Zschocke J. 2003. A role for overdominant selection in phenylketonuria? Evidence from molecular data. Hum Mutat 21:394–397. Kure S, Hou DC, Ohura T, Iwamoto H, Suzuki S, Sugiyama N, Sakamoto O, Fugii K, Matsubara Y, Narisawa K. 1999. Tetrahydrobiopterin-responsive phenylalanine hydroxylase deficiency. J Pediatr 135:375–378. Kwok SCM, Ledley FD, Dilella AG, Robson KJH, Woo SLC. 1985. Nucleotide sequence of a full-length complementary DNA clone and amino acid sequence of human phenylalanine hydroxylase. Biochemistry 24:556–561. Leandro P, Rivera I, Lechner MC, Tavares de Almeida I, Konecki D. 2000. The V388M mutation results in a kinetic variant form of phenylalanine hydroxylase. Mol Genet Metab 69:204–212. Ledley FD, Grenett HE, Dunbar BS, Woo SLC. 1990. Mouse phenylalanine hydroxylase. Homology and divergence from human phenylalanine hydroxylase. Biochem J 267: 399–405. Martinez A, Andersson KK, Haavik J, Flatmark T. 1991. EPR and 1H-NMR spectroscopic studies on the paramagnetic iron at the active site of phenylalanine hydroxylase and its interaction with substrates and inhibitors. Eur J Biochem 198:675–682. Martinez A, Knappskog PM, Olafsdottir S, Doskeland AP, Eiken HG, Svebak RM, Bozzini M, Apold J, Flatmark T. 1995. Expression of recombinant human phenylalanine hydroxylase as fusion protein in Escherichia coli circumvents proteolytic degradation by host cell proteases. Biochem J 306:589–597. Maurer SM, Firestone RB, Scriver CR. 2000. Science’s neglected legacy. Nature 405:117–120. McDonald JD, Bode V, Dove W, Shedlovsky A. 1990. Pahhph-5: a mouse mutant deficient in phenylalanine hydroxylase. Proc Natl Acad Sci USA 87:1965–1967. McDonald JD, Charlton CK. 1997. Characterization of mutations at the mouse phenylalanine hydroxylase locus. Genomics 39:402–405. 344 SCRIVER ET AL. Okano Y, Eisensmith RC, Guttler F, Lichter-Konecki U, Konecki DS, Trefz FK, Dasovich M, Wang T, Henriksen K, Lou H, Woo SLC. 1991. Molecular basis of phenotypic heterogeneity in phenylketonuria. N Engl J Med 324:1232– 1238. Sarkissian C, Shao Z, Blain F, Peevres R, Su H, Heft R, Chang TMS, Scriver CR. 1999. A different approach to treatment of phenylketonuria: phenylalanine degradation with recombinant phenylalanine ammonia lyase. Proc Natl Acad Sci USA 96:2339–2344. Sarkissian C, Boulais DM, McDonald JD, Scriver CR. 2000. A heteroallelic mutant mouse model: a new orthologue for human hyperphenylalaninemia. Mol Genet Metab 69:188– 194. Scriver CR. 2001. Human genetics: lessons from Quebec populations. Annu Rev Genomics Hum Genet 2:69–101. Scriver CR, Kaufman S. 2001. Hyperphenylalaninemia: phenylalanine hydroxylase deficiency. In: Scriver CR, Beaudet A, Sly WS, Valle D, Childs B, Kinzler KW, Vogelstein B, editors. The metabolic and molecular bases of inherited disease. New York: McGraw-Hill. pp 1667–1724. Scriver CR, Waters PJ. 1999. Monogenic traits are not simple. Lessons from phenylketonuria. Trends Genet 15:267–272. Scriver CR, Byck S, Prevost L, Hoang L, PAH Mutation Analysis Consortium. 1996. The phenylalanine hydroxylase locus: a marker for the history of phenylketonuria and human genetic diversity. In: Chadwick D, Cardew G, editors. Variation in the human genome. Ciba Foundation Symposium No. 197. Chichester: John Wiley. Scriver CR, Waters PJ, Sarkissian C, Ryan S, Prevost L, Cote D, Novak J, Teebi S, Nowacki PM. 2000. PAHdb: a locusspecific knowledgebase. Hum Mutat 15:99–104. Shedlovsky A, McDonald JD, Smyula D, Dove WF. 1993. Mouse models of human phenylketonuria. Genetics 134:1205–1210. Sullivan SE, Lidsky AS, Brayton K, Dilella AG, King M, Connor M, Cockburn F, Woo SLC. 1985. Phenylalanine hydroxylase deletion mutant from a patient with classical PKU. Amer J Hum Genet 37:A177 (Abstract). Svensson E, Wang Y, Eisensmith RC, Hagenfeldt L, Woo SLC. 1993. Three polymorphisms but no disease-causing mutations in the proximal part of the promotor of the phenylalanine hydroxylase gene. Eur J Hum Genet 1:306–313. Teebi SA, Porter CJ, Talbot CC Jr, Cuticchia AJ. 2001. The BiSC WayStation: a centralized structure for the collection of genetic variations. Am J Hum Genet 69:452 (Abstract 1581). Teigen K, Froystein NA, Martinez A. 1999. The structural basis of the recognition of phenylalanine and pterin cofactors by phenylalanine hydroxylase: implications for the catalytic mechanism. J Mol Biol 294:807–823. Terp BN, Cooper DN, Christensen IT, Jorgensen FS, Bross P, Gregersen N, Krawczak M. 2002. Assessing the relative importance of the biophysical properties of amino acid substitutions associated with human genetic diseease. Hum Mutat 20:98–109. Tighe O, Dunican D, O’Neill C, Bertorelle D, Beattie D, Graham C, Zschocke J, Cali F, Romano V, Hrabincova E, Kozak L, Nechyporenko M, Livshits L, Guldberg P, Jurkowska M, Zekanowski C, Perez B, Ruiz Desviat L, Ugarte M, Kucinskas V, Knappskog P, Treacy E, Naughten E, Tyfield L, Byck S, Scriver CR, Mayne PD, Croke DT. 2003. Genetic diversity within the R408W phenylketonuria mutation lineages in Europe. Hum Mutat 21:387–393. Treacy E, Byck S, Clow C, Scriver CR. 1993. ‘Celtic’ phenylketonuria chromosomes found? Evidence in two regions of Quebec province. Eur J Hum Genet 1: 220–228. Venter JC, Adams MD, Myers EW, et al. 2001. The sequence of the human genome. Science 291:1304–1351. Waters PJ. 2003. How PAH gene mutations cause hyperphenylalaninemia and why mechanism matters: insights from in vitro expression. Hum Mutat 21:357–369. Waters PJ, Parniak MA, Nowacki P, Scriver CR. 1998. In vitro expression analysis of mutations in phenylalanine hydroxylase: linking genotype to phenotype and structure to function. Hum Mutat 11:14–17. Waters PJ, Parniak MA, Akerman BR, Scriver CR. 2000. Characterization of phenylketonuria missense substitutions, distant from the phenylalanine hydroxylase active site, illustrates a paradigm for mechanism and potential modulation of phenotype. Mol Genet Metab 69:101–110. (Erratum in Mol Genet Metab 72:89.) Weiss KM. 1996. Is there a paradigm shift in genetics? Lessons from the study of human diseases. Mol Phylogenet Evol 5:259–265. Zschocke J. 2003. Phenylketonuria mutations in Europe. Hum Mutat 21:345–356. Zschocke J, Hoffmann GF. 1999. Phenylketonuria mutations in Germany. Hum Genet 104:390–398. Zschocke J, Graham CA, Carson DJ, Nevin NC. 1995. Phenylketonuria mutation analysis in Northern Ireland: a rapid stepwise approach. Am J Hum Genet 57: 1311–1317. Zschocke J, Mallory JP, Eiken HS, Nevin NC. 1997. Phenylketonuria and the peoples of Northern Ireland. Hum Genet 100:189–194. Zschocke J, Quak E, Knauer A, Fritz B, Asselan M, Hoffmann GF. 1999. Large heterozygous deletion masquerading as homozygous missense mutation: a pitfall in diagnostic mutation analysis. J Inherit Metab Dis 22:687–692.