* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download What is a genome?
Synthetic biology wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Gene desert wikipedia , lookup
Microevolution wikipedia , lookup
Epigenomics wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Human genetic variation wikipedia , lookup
Molecular Inversion Probe wikipedia , lookup
Adeno-associated virus wikipedia , lookup
Genetic engineering wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Oncogenomics wikipedia , lookup
Designer baby wikipedia , lookup
Copy-number variation wikipedia , lookup
Genome (book) wikipedia , lookup
Transposable element wikipedia , lookup
Microsatellite wikipedia , lookup
History of genetic engineering wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Public health genomics wikipedia , lookup
DNA sequencing wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Non-coding DNA wikipedia , lookup
Helitron (biology) wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Minimal genome wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Human genome wikipedia , lookup
Pathogenomics wikipedia , lookup
Craig Venter wikipedia , lookup
Genome editing wikipedia , lookup
Metagenomics wikipedia , lookup
Genome evolution wikipedia , lookup
Genomic library wikipedia , lookup
Whole genome sequencing wikipedia , lookup
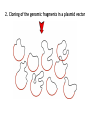
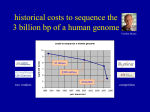
Bioinformatics for Genomics It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material. When I was young my Father used to tell me that the two most worthwhile pursuits in life were the pursuit of truth and of beauty and I believe that Alfred Nobel must have felt much the same when he gave these prizes for literature and the sciences. Sometime in the future, I am a hundred percent certain scientists will sit down at a computer terminal, design what they want the organism to do, and build it. What is a genome? What is a genome? The entirety of the inheritable information of an organism Being able to sequence the genome allows us to 'read' all this information of an organism (and hopefully understand it) The steps of the sequencing of the human genome 1953. Watson and Crick propose the double helix model for DNA 1977. Sanger proposes the sequencing method with terminators 1986. Dulbecco auspicates in Science sequencing of the human genome 1988. Watson becomes director of the project at the NIH 1991. Craig Venter (then NIH) publishes the first EST sequencing 1992. Watson leaves the direction of the genome project at NIH, enter Collins 1992. Venter leaves NIH and founds TIGR 1995. TIGR sequences from the first bacterial genome, with the shotgun method 1998. Venter founds Celera Genomics and PE; He announces the genome for 2001 The race for the human genome starts (Venter VS Collins) 26 June 2000. Bill Clinton announces the completion of the human genome sequencing (actually two 'proofs', published in early 2001) to sequence the genome we must first be able to sequence a gene How long is a gene? Bacterial gene Eukaryotic gene How long is a gene? 1000 bp for a bacterial gene 3000 bp for a human gene (much more variable) How long is a Sanger sequence? Ok, but how long is a genome? And how many genes does it contain? Ok, but how long is a genome? • • • • • Human genome: 3,12 billion bases, 3120 Megabases (Mb) Drosophila: 175 Megabases Small eukaryote (Saccharomyces cerevisiae): 12 Mb Big bacterial genome (Pseudomonas aeuriginosa): 6,4 Mb Small bacterial genome (Haemophilus inflenzae): 1,8 Mb And how many genes does it contain? • Human genome: 3120 Mb, 25.000-40.000 genes • Pseudomonas aeuriginosa: 6,4 Mb, 5570 genes • Haemophilus inflenzae: 1,8 Mb, 1727 genes So how do we sequence an entire genome? The strategy of the public consortium for the human genome Cut a chromosome into pieces with restriction enzymes, clone one piece in a vector, subclone into smaller vectors, and into smaller vectors YACs (yeast artificial chromosome), inserts from 0.5-1 Mb BACs (bacterial artificial chromosomes), inserts of 0.1-0.2 Mb Cosmids, inserts under 50 Kb Plasmid vectors, few kb Merging the fragments of the genome cloned in the different vectors Here comes Venter, with his rebellion against NIH Just 1% of the human genome translated into proteins Why not start to sequence just mRNA? Venter generates cDNA from RNA (reverse transcription) → cDNA cloning → random sequencing of clones → production of EST EST = Expressed Sequence Tags 1991: Venter publishes 337 ESTs corresponding to genes expressed in the brain The publication in Science was preceded by the presentation of a patent application for the sequenced ESTs Patent rejected, he leaves NIH to give birth to the TIGR, financed by Healthcare Investment, with the goal to compete with the public consortium to sequence the human genome 1993 – A novel idea for the first genome End of 1993. Hamilton Smith (Nobel in 1978 for the description of the first restriction enzyme) enters the Scientific Committee of the TIGR. Smith proposes the idea of the shotgun sequencing Chosen organism: Haemophilus influenzae Why shotgun? NOT A NOVEL TECHNOLOGY IT IS A NOVEL EXPERIMENTAL METHOD It was invented to work with Sanger technology... … but the principle is still used with nextgen 1. Random fragmentation of many copies of genomic DNA Sonication, hydroshearing... 2. Cloning of the genomic fragments in a plasmid vector 3. Sequencing of the inserts based on PCR primers located on the plasmid (known sequence) Each sequence we obtain will be called a sequencing READ 3. Sequencing of the inserts based on PCR primers located on the plasmid (known sequence) An excess of sequences is generated If the genome is x long, at least 5x bases are sequenced The obtained sequences will cover the genome, randomly, with different 'depth', or 'coverage' Genome coverage: The average depth of sequencing coverage is theoretically LN/G L is the read length N is the number of reads G is the genome length LN= total number of bases generated Genome coverage: The average depth of sequencing coverage is theoretically LN/G L is the read length N is the number of reads G is the genome length LN= total number of bases generated EXAMPLE: 1 Megabase genome 100 nt reads 200,000 reads The coverage is 200,000x100/1,000,000=20 Genome coverage: The average depth of sequencing coverage is theoretically LN/G HOWEVER: genome coverage of 1 does not mean that each base in my genome has been sequenced, because I am sequencing randomly P=1-e-m The probability that a base has been sequenced (P) is equal to 1 minus e (Euler's number = 2.71828) elevated to -m, where m is the coverage Genome coverage A coverage of 5x leads to sequencing of 99.33% of the bases IN THEORY Possible problems? Genome coverage Uneven sequencing Can be due to Technical biases Difficult stretches So in the times of Sanger sequencing 10x was considered good Now with NextGen you want to go 30x, better if 50x 4. use bioinformatic techniques to ASSEMBLE the genome Most bases will be covered on both strands, a few gaps will be present Example: I want to sequence the following genome This lecture is very very interesting I fragment, clone and sequence This l is lect ure is e is v ry ver teres ecture s very v ry inte esting But I do not know the order of the fragments Thanks to the partial overlapping I can reconstruct the sequence This l is lect ecture ure is e is v s very v ry ver ry inte teres esting Advantages of shotgun sequencing -No Need for genomic mapping prior to sequencing -High level of automation -rapidità the sequencing phase -reducing costs Issues of shot-gun sequencing - Highly purified genomic DNA - Bionformatic support - Difficulty in presence of repeated sequences in the genome Let's try to sequence the sentence below It's a fair bet that if it's fair tomorrow, then my fair wife and I will head to the Spring Fair, held in a fair sized park, in this fair city to win a prize, if everyone plays fair It's a fair bet that if it's fair tomorrow, then my fair wife and I will head to the Spring Fair, held in a fair sized park, in this fair city to win a prize, if everyone plays fair We will have a number of sequences that will read 'fair' and we would not know how to assemble them So repeats, and other issues, can lead to gaps in the assembly Our assembly will not consist of 1 or more chromosome, but multiple contigs CONTIG: a contiguous sequence generated by the overlap of sequencing reads 5. Finishing This is the step in which we try to close the gaps 5. Finishing This is the step in which we try to close the gaps Molecular biology methods can help: Cloning in vectors that can receive long inserts PCR Inverse PCR Bioinformatics can also help, with a number of approaches, but mainly with scaffolding PAIRED END READS These are reads that are generated when both ends of a fragment are sequenced We will thus have 2 reads, that represent 2 fragments of the genome which are distinct, but near PAIRED END READS Many technologies can generate paired end reads (Sanger, 454, Illumina) We can use the information from pairs to order contigs into a scaffold SCAFFOLD A sequence of a genome that is composed of contigs and gaps Contigs are ordered based on the information from paired ends The unknown bases are indicated with N SHOTGUN SEQUENCING PIPELINE 1. Fragmentation 2. Cloning 3. Random sequencing at high coverage 4. Bioinformatic assembly 5. Finishing Final result: a complete assembly (sometimes) So, does shotgun sequencing work? Sequencing of Haemophilus influenzae (1995) proved that it does Automatization of the preparation steps Improvement of Sanger sequencing technologies Evolution of powerful assembly bioinformatics algorithms Sequencing of D. melanogaster (2000) Sequencing of the human genome (1998-2003) "When we started to sequence the Drosophila, we had already halved the timing and sequencing of a genome assembly such as Haemophilus, which only three years ago represented a whole year of work. Today it would only take eight hours to sequence it and fifteen minutes to assemble it. " Craig Venter 5 November 2003 And the human genome In 1998, the public project, which has already costed1.9 billion dollars, is going slowly ... In January 1998, Mike Hunkapiller (PE) Shows Venter the PRISM 3700 (96 capillaries) Celera Genomics is founded (PE funding, direction by Venter): 300 PRISM 3700, 100 robots, and computers for 80 million dollars Celera announcees the sequencing of the human genome in two years, with a shotgun approach to cost 200-500 million dollars The race continues, with strong ethical implications 26 June 2000. Bill Clinton announces the completion of the human genome sequencing (actually two 'proofs', published in early 2001). And now the bioinformatics! The assembly starts when we generate the output of the sequencing machines This output can differ, but most of the time it will be a big FASTQ file maybe you know what a FASTA file is? a FASTA file >PoolPRRSnew_1/1 CCCCGGGTCAAGGGCTGTTGTTTTATTGTTCACCTTCATTATGACAGTGCGAGGTGGTTT ATACGGGATTTGCTGCAACAC >PoolPRRSnew_2/1 CTTTCATGTGGTACGCAAGGGCGGCCAGGATCCGGTCACGATTGGGAACTAACTGCCGC CCTGCTTCAATTTTGCAGCCGA Starts with a > Followed by the name of the sequence On the second line we have the nucleotides The FASTQ FORMAT Specific file format for DNA sequences @PoolPRRSnew_1/1 CCCCGGGTCAAGGGCTGTTGTTTTATTGTTCACCTTCATTATGACAGTGCGAGGTGGT + 1D1>CDF1AAAEABFGGHHGDGFAGC2EEAG11FBF2GHFHHHBBEGCEFFE/AEEEDGGE @PoolPRRSnew_2/1 CTTTCATGTGGTACGCAAGGGCGGCCAGGATCCGGTCACGATTGGGAACTAACTGCC + ABBBBFFFFFBCFGGGGGGGGGGGGGGHHAHHHFGGGGHGEGHFHGHHGHHHHHHH It is a plain text file that contains all the information regarding the reads and their quality THE FASTQ FORMAT Each sequencing read is described in 4 line FIRST LINE: @ followed by the unique name of the sequence SECOND ROW: nucleotide sequence THIRD ROW: + FOURTH ROW: the quality of each base, in ASCII code @PoolPRRSnew_1/1 CCCCGGGTCAAGGGCTGTTGTTTTATTGTTCACCTTCATTATGACAGTGCGAGGTGGTTTATACGG + 1AAA?1D1>CDF1AAAEABFGGHHGDGFAGC2EEAG11FBF2GHFHHHBBEGCEFFE/AEEEDGGE?E? A BASE QUALITY Every sequence can contain errors Base quality: given a probability of error P, the quality is: Q = -10 log10P So the probability of descresce error in logarithmic scale with increasing quality Bases with a quality <20 are usually discarded: QUALITY CHECK THE ASSEMBLY The ASSEMBLER is a software that uses algorithms to combine the short sequences obtained in the process of sequencing in longer fragments (contigs) The goal is to find the shortest string (the genome) that includes all of the input strings (the reads) As the number of input strings grows, the difficulty of the problem grows exponentially (NP complete problem) We must use approximation algorithms ASSEMBLY ALGORITHMS An example of algorithm 'greedy': 1. pairwise comparison of the sequences 2. Choose the pair that aligns best 3. repeat steps 1 and 2 Greedy algorithms have issues with repeats ASSEMBLY ALGORITHMS Other methods are more refined and based on graphs – Overlap-layout-consensus – EULER assembler ASSEMBLY ALGORITHMS The difficulties that an assembler has to face are: amount of data (gigabytes of sequences) presence of repeated sequences presence of sequencing errors Tens of Assemblers have been developed, how to choose Assemblathon 2: evaluating de novo methods of genome assembly in three vertebrate species - GigaScience 2013, 2:10 doi:10.1186/2047-217X-2-10 Read the literature • Consider your specific biological system • See what others have done • Read the manuals Try different assemblers and evaluate your results ASSEMBLY RESULTS Your assembler will output a set of contigs, usually in a fasta file >contig19.1|size257_bb gctcatttataacgttcctcaaagcctaaatgacaggttcaggtgggttgccttgctccc gtcacctacccttcaaaatcaataatttcacccgctcctctatcaactgcaatcaatata aaggtgtactcccacttccattctttggtcttttttttatatatgtgcacagctcctcaa tctctaggacctcgatatccttaagtgactttgtgctgttgcagctcttatctaactttt cttgcacatagccgccc >contig20.1|size182_bb ccaaaaagtataaacttttcaacgcctttttgaaacaatttccaagcttataaaacttga taaggaaccggtcttccagccgcatcacgcactaaatcaccgttatcaaatttagctgct acataattgccatgcccaatttttccgcccttatacattacaggtttcacttcttttgca cc How good is my assembly? What parameters should we check? ASSEMBLY RESULTS Your assembler will output a set of contigs, usually in a fasta file >contig19.1|size257_bb gctcatttataacgttcctcaaagcctaaatgacaggttcaggtgggttgccttgctccc gtcacctacccttcaaaatcaataatttcacccgctcctctatcaactgcaatcaatata aaggtgtactcccacttccattctttggtcttttttttatatatgtgcacagctcctcaa tctctaggacctcgatatccttaagtgactttgtgctgttgcagctcttatctaactttt cttgcacatagccgccc >contig20.1|size182_bb ccaaaaagtataaacttttcaacgcctttttgaaacaatttccaagcttataaaacttga taaggaaccggtcttccagccgcatcacgcactaaatcaccgttatcaaatttagctgct acataattgccatgcccaatttttccgcccttatacattacaggtttcacttcttttgca cc To evaluate the assembly we use multiple parameters: the number of contigs the genome size N50: the length of the shortest contig that added to those of higher dimensions comprises at least 50% of the genome N50: the length of the shortest contig that added to those of higher dimensions comprises at least 50% of the genome EXAMPLE OF ASSEMBLY RESULTS Sequencing of 16 genomes of Klebsiella pneumoniae ASSEMBLY RESULTS Genome size: GOOD, always as expected N50: not bad, always above 100000nt Contigs number: acceptable, maybe not for KPCVA3 However, since we know our biological system, we know these genomes contain plasmids, which make the assembly quality lower ASSEMBLY RESULTS What is GOOD depends on what we aim for If we are sequencing a novel genome, we should try to CLOSE it If we want to compare similar isolates we can accept the above results