* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download When Cells Tell Their Neighbors Which Direction to Divide
Survey
Document related concepts
Signal transduction wikipedia , lookup
Tissue engineering wikipedia , lookup
Endomembrane system wikipedia , lookup
Biochemical switches in the cell cycle wikipedia , lookup
Extracellular matrix wikipedia , lookup
Cell encapsulation wikipedia , lookup
Programmed cell death wikipedia , lookup
Cell culture wikipedia , lookup
Organ-on-a-chip wikipedia , lookup
Cell growth wikipedia , lookup
Cellular differentiation wikipedia , lookup
Transcript
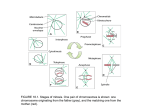
DEVELOPMENTAL DYNAMICS 218:23–29 (2000) REVIEWS A PEER REVIEWED FORUM When Cells Tell Their Neighbors Which Direction to Divide BOB GOLDSTEIN* Department of Biology, University of North Carolina at Chapel Hill, Chapel Hill, North Carolina ABSTRACT Many cells must divide in specific orientations, yet for only a handful of cases do we have some understanding of how cells choose division orientations. We know of only a few cases where division orientations are controlled by specific cell– cell interactions. These cases are of interest, because they tell us something new and seemingly fundamental about how cells can function during development. Here, the evidence that interactions control division orientation in some cells of the early C. elegans embryo is presented, and what is known about how contact can regulate division orientation is discussed. Whether contact-mediated division orientation is a peculiarity of C. elegans or whether it may be more widespread is addressed. Dev Dyn 2000;218:23–29. © 2000 Wiley-Liss, Inc. Key words: cell division orientation; cell interaction; cytoskeleton; mitotic spindle; Wnt pathway; C. elegans INTRODUCTION Many cells must choose their division orientations carefully. For example, a cell in a developing monolayer epithelium must divide in a particular plane for its daughters to remain in the monolayer epithelium, and a cell in which certain components are asymmetrically localized must orient its division axis if it is to partition such components to just one daughter cell (Fig. 1). Embryologists have been isolating embryonic cells for more than a century, and for scant few cases do we know that division orientations are affected, suggesting at first glance that cell contacts may be generally dispensable for proper division orientations. However, in most cases, the appropriate landmarks for proper division orientation are not apparent. Therefore, the question of whether division orientation requires specific cell contacts has remained unaddressed in most instances. We have known for some time that cell shape can play a role in division orientation, as mitotic spindles tend to shift into the longest axis of a cell as the growing asters run out of room (Hertwig, 1895). Hence, © 2000 WILEY-LISS, INC. cells, or physical barriers such as a vitelline envelope, can influence a cell’s division axis by altering cell shape. Only recently have we learned that cells also can specifically orient division axes of certain neighbors; this is the topic of this review. CELL CONTACTS ORIENT SOME CELL DIVISION AXES IN C. ELEGANS In the course of experiments using cultured C. elegans embryonic cells to investigate a potential role for cell interactions in specifying certain cell fates, some cells, when placed in contact with certain other cells, were found to divide in predictable orientations. After placing cells in contact in various orientations, one cell consistently divided in an axis that left its daughters in line with the contacting cell, suggesting that certain cells can induce specific division orientations in certain other cells (Goldstein, 1995). Which Cells Can Have Their Divisions Oriented by Cell Contacts? By making pairwise combinations of all cells of the four cell stage C. elegans embryo, one cell was found to be capable of inducing division orientation, and another cell was found to be capable of responding by orienting its division axis. The names of cells in early C. elegans development are shown in Figure 2; the cell capable of inducing is called P2, and the cell capable of responding is called EMS. As most of the ensuing experiments have involved these two cells, for simplicity P2 is often referred to here as the “inducing cell,” and EMS as the “responding cell.” There are, however, other cells which can induce and other cells which can respond: similar experiments performed with selected cells of the eight cell stage revealed that both daughters of P2 (P3 and C) could orient the divisions of both daughters of EMS (E and MS). In the eight cell stage embryo, neither daughter of the inducing cell touches the MS cell, suggesting that in normal development MS has an unrealized potential to respond. These re- *Correspondence to: Bob Goldstein, Department of Biology, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599-3280. E-mail: [email protected] Received 10 January 2000; Accepted 8 February 2000 24 GOLDSTEIN Fig. 1. The consequences of alternative spindle orientations for a polarized cell (A) and for a cell in a monolayer epithelium (B). Black areas in (A) represent asymmetrically localized components. sults, along with cell isolation experiments in other cells (Goldstein, 1995) and results described below have led to the generalization that cell division orientations can be regulated in two ways in the early C. elegans embryo— by cell interactions in certain somatic cells, and cell-autonomously in germline precursor cells (Fig. 2). Fig. 2. The 1-, 2-, 4-, and 8-cell stages of C. elegans. Cell division orientations are regulated cell autonomously in the cells marked blue (Goldstein, 1995; Gönczy et al 1999b), and can be regulated by cell interactions in the cells marked yellow (Goldstein, 1995). The white cells are those in which cell division orientations are likely to be unregulated, as divisions follow a default orthogonal pattern (Hyman and White, 1987). HOW DO CELL CONTACTS CONTROL DIVISION ORIENTATION? As the plane of cell division is determined by the positions of the mitotic asters (Rappaport, 1961), which are nucleated by centrosomes, division orientation by cell contact was suspected to work by affecting the positions of the centrosomes. Hyman and White (1987; Hyman, 1989) have shown that centrosomes are positioned in certain cells of the early C. elegans embryo by a mechanism involving capture of astral microtubules at a specific site in the cortex. This site is referred to here as the “cortical site.” The microtubule tether between the centrosome and the cortical site shortens, resulting in rotation of the centrosome-nucleus complex and, consequently, alignment of the division axis with the cortical site (Fig. 3). Evidence which suggested this model included the findings that depolymerizing microtubules prevented rotation of the centrosome-nucleus complex, and that laser-ablating the proposed cortical site, the leading centrosome, or at sites along the proposed tether could prevent rotation, whereas ablating other sites in the cell had no such effect. Additional evidence for a pulling force generated by the microtubule tether was the Fig. 3. Model for cell-autonomous spindle orientation by a cortical site in the P1 cell of the two cell stage (Hyman, 1989; Skop and White, 1998; Waddle et al., 1994). The black spot represents the cortical site, and the arrow shows the direction of rotation of the centrosome-nucleus complex. The plus and minus ends of the tethered microtubule(s) are marked. observation of a dimple forming around the cortical site during rotation (Hyman, 1989; Keating and White, 1998). Cortical sites for orienting mitotic spindles have also been reported in budding yeast (Palmer et al., INDUCED CELL DIVISION ORIENTATION 1992) and in embryos of sea urchins (Dan, 1979), clams (Dan and Ito, 1984), flies (Kraut et al., 1996), ascidians (Hibino et al., 1998; Nishikata et al., 1999), and fucoid algae (Fowler and Quatrano, 1997), as well as for meiotic spindles in annelids (Lutz et al., 1988), suggesting they represent a common theme in eukaryotic cells which divide asymmetrically. Does contact-induced division orientation work by inducing such cortical sites? To test this, centrosome positions were followed live in isolated inducing-responding cell pairs (Goldstein, 1995). Rotation of the centrosome-nucleus complex occurred in the responding cell toward the site of cell– cell contact. Anti-microtubule immunofluorescence showed that one end of the mitotic spindle in the responding cell ended up close to the site of contact with the inducing cell, as in intact embryos (Hyman and White, 1987). These movements were dependent on cell contact, as in the absence of an inducing cell, no rotation occurred, and division always occurred in a plane perpendicular to the usual division plane (Goldstein, 1995). Placing the inducing cell near to but not touching the responding cell (about 1 cell diameter apart) also failed to cause rotation (BG, unpublished). As expected, contact-induced rotation could be prevented by treating cells with a microtubule-depolymerizing drug. The results suggest that the interaction affects spindle orientation by inducing a cortical site which captures astral microtubules, resulting in realignment of the mitotic spindle. Rotation of the centrosome-nucleus complex is induced by cell interactions in only certain cells of the early C. elegans embryo. In other cells, rotation occurs cell-autonomously (i.e., it occurs even when these cells are cultured in isolation), and in yet other cells, rotation does not occur at all and divisions occur in successively orthogonal planes (Fig. 2). To test whether cell contact induces a cortical site de novo, or alternatively whether each responding cell might contain only a single site that translocates to the area of contact with the inducing cell, two inducing cells were placed on one side of a responding cell (Goldstein, 1995). Each of the two centrosomes was found to move toward one of the two inducing cells, pulling the nucleus to an eccentric position in the cell. The result indicates that a responding cell is capable of forming new cortical sites in response to cell contacts. Contact Is Required Only Briefly, and the Response Does Not Require New Transcription Cell contact was found to be required at a specific time, 10 min prior to cytokinesis in the responding cell. Recent experiments by A. Schlesinger (personal communication) using cells of disparate ages have revealed that this is a property of the responding cell: the inducing cell could induce at any time during its cell cycle, but the responding cell required contact at the critical time. Whether the responding cell requires contact only at this time, or alternatively also for some time afterward, is not yet known. 25 Schlesinger et al. (1999) have tested whether transcription is required for contact-mediated spindle orientation in EMS by treating cell pairs with actinomycin D, a chemical inhibitor of transcription, and by inactivating RNA polymerase II by RNA interference (RNAi; a method of inactivating gene function by injection of double-stranded RNA; Fire et al., 1998). When either or both of these treatments were used, a GFP transgene used to monitor transcription level was undetectable by fluorescence microscopy, yet the spindle in the responding cell still rotated toward the site of contact with the inducing cell, suggesting that the interaction affects the responding cell’s cytoskeleton directly, without requiring new transcription. The Molecular Basis for Contact-Induced Division Orientation A recent article has identified for the first time some of the genes required for contact-induced division orientation— homologs of Wnt pathway components (Schlesinger et al., 1999). The first Wnt gene was discovered in 1982 as a mouse proto-oncogene, and mutations in most of the Wnt pathway genes have since been implicated in cancers. Wnt pathway genes function in many developmental signaling pathways in diverse organisms (see Wodarz and Nusse, 1998 for review). Wnt pathway components were previously known to be required in C. elegans for P2 to induce EMS to produce a gut cell lineage (see Han, 1997 for review). Schlesinger et al. (1999) isolated the cell (called P1) which produces inducing and responding cells from embryos in which each of nine genes in the gut induction pathway were inactivated by mutation or RNAi. Each P1 cell was allowed to divide to form the inducing and responding cells, and then the division plane of the responding cell was followed. In wild-type embryos, this manipulation results in the responding cell dividing in a single division plane with respect to the plane of cell contact, whereas in several of the mutants and RNAi-affected embryos, responding cells divided in various orientations. Isolated cells were also combined from mutant and wild-type embryos to determine in which cell each gene is required. Genes found to be required in the signaling cell included mom-1 (a homolog of fly porcupine) and mom-3 (uncloned). Genes found to be required in the responding cell included gsk-3 (zeste-white3) and mom-5 (frizzled). Although inactivation of any of these genes affect spindle orientation in this assay, none of the relevant alleles are completely penetrant for loss of rotation, suggesting that either none of the alleles used are null alleles or that these genes might play partially redundant roles in spindle orientation. The signal for spindle orientation remains unidentified. One would expect a Wnt gene to be required, but surprisingly mom-2 (wingless), a proposed signal for gut induction, seems dispensable for spindle orientation, as it can only be shown to be defective in signaling 26 GOLDSTEIN in an experiment where the inducing cell is aged prior to being placed on a younger responding cell (Schlesinger et al., 1999). The C. elegans genome has only four other apparent wingless-related genes (cwn-1, cwn-2, lin-44, and egl-20; C. elegans Sequencing Consortium, 1998; Herman et al., 1995; Maloof et al., 1999; Shackleford et al., 1993); whether any of these signal for spindle orientation, or play a redundant role with mom-2, have yet to be tested. It will be interesting to see whether two signals are required from the inducing cell, as this possibility has been raised for the case of gut induction by the incomplete penetrance of all known mutations affecting the inducing cell’s ability to signal (Rocheleau et al., 1997; Thorpe et al., 1997). Genes in the Wnt pathway expected to act downstream of gsk-3, such as apr-1 (APC-related), wrm-1 (-catenin-related), pop-1 (pan/LEF-1) and mom-4 (TAK1) appear to be involved only in gut induction and not in spindle orientation, as mutations or RNAi for each of these genes have little if any effect on spindle orientation. apr-1;wrm-1 double RNAi also has no effect on spindle orientation. Another cell division in the early C. elegans embryo, the division of ABar, has a similar dependence on Wnt pathway genes as the EMS cell (Rocheleau et al., 1997; Thorpe et al., 1997), suggesting that its division orientation might also be induced by cell contacts. The results of Schlesinger et al. (1999) demonstrate that many of the Wnt pathway genes required for gut induction also play a role in spindle orientation, and that the pathway for spindle orientation splits following gsk-3. Especially in light of the fact that no new transcription is required for the response, it will be interesting to see in the future how this Wnt pathway links to the cytoskeleton. Together with the known roles of mutations in Wnt pathway genes in cancer, the results additionally raise the possibility that a failure of induced division orientation might play a role in the development of some cancers. Cells leave epithelia in an early step of many cancers, yet we do not know whether cells sometimes leave epithelia in these cases because of defects in spindle orientation. In fact, little is known about how cell divisions are properly oriented in monolayer epithelia (see Reinsch and Karsenti, 1994). Additional candidates for genes involved in contactmediated spindle orientation include the genes involved in cell-autonomous spindle orientation in the germline precursor cells. The growing list of genes required for cell-autonomous spindle orientations, or required to prevent rotation in cells in which rotation does not normally occur, includes the six par genes, pkc-3, let-99, nmy-2, pod-1, gpb-1, mes-1, ooc-3, ooc-5, stu-10, stu-11, lin-5, and rot-1,2,3 (see Rose and Kemphues, 1998 for review; Basham and Rose, 1999; Gönczy et al., 1999a; Lorson et al., 2000; O’Connell et al., 1998; Rappleye et al., 1999; Strome et al., 1995; Zwaal et al., 1996). Mutations in most of these genes affect early divisions dramatically, making it difficult to test their functions in cells with induced cell division orientations, as the appropriate cells do not form normally in the mutants. It is clear, however, that in normal development those PAR proteins which are asymmetrically localized in cells with autonomous cell division orientations are not asymmetrically localized in cells with induced cell division orientations. This result suggests that the asymmetrically localized PAR proteins do not function in these induced cell division orientations (see Kemphues and Strome, 1997 for review). Other genes which might be expected to be necessary for contact-mediated spindle orientation include genes required for the normal execution of the inducing cell’s fate; in fact, the pie-1 gene, which is necessary for normal execution of the P2 cell’s fate (Mello et al., 1992), is required for normal spindle orientation in P2-EMS cell pairs (BG, unpublished communication). What Is at the Cortical Site Data presented thus far suggests that contact between particular cells induces a site, in the cortex of the responding cell, which can capture astral microtubules. In cells with autonomously orienting spindles, several components have been found to be present at cortical sites. These include a concentration of actin (Waddle et al., 1994) and two dynactin complex members—actin capping protein (Waddle et al., 1994), and DNC-1 (a C. elegans homolog of p150-Glued; Skop and White, 1998)— which assemble at the site of the midbody (the persistent remnant of cell division). These findings have led to the hypothesis that a dynactin complex links cortical actin at the midbody to a dynein motor, which reels in microtubules by walking in place toward the minus end of any astral microtubules that it encounters (Skop and White, 1998; Waddle et al., 1994). Data consistent with this model include Skop and White’s (1998) finding that RNA interference of dnc-1 or dnc-2 (a homolog of p50/Dynamitin, another dynactin complex member) can prevent rotation from occurring in P1, although Gönczy et al. (1999b) have suggested that this may be a result of eccentric placement of the cortical site rather than failure of rotation. Gönczy et al. (1999b) also found that a dynein homolog is present at the cortical site, although its distribution is not restricted to this site: DHC-1, a dynein heavy chain, is present throughout the cytoplasm and on nuclei, and is enriched in the cortex, particularly along cell boundaries. Although the hypothesis outlined here is at least consistent with current data, other hypotheses are clearly plausible as well. For example, it remains possible that dynein or another microtubule binding protein at the cortical site might cause rotation by simply holding onto the plus end of microtubules during microtubule catastrophe events (as in Lombillo et al., 1995), rather than by walking along microtubules. A motor for rotation might be present at the centrosome end of the microtubule tether rather than at the cortical site, as no INDUCED CELL DIVISION ORIENTATION 27 Fig. 4. Concentrations of actin and actin capping protein at areas of contact between specific cells (left). These concentrations are visible under only certain fixation conditions; these embryos were fixed in 3% formaldehyde and then 100% methanol. Note that for each disc of staining at the border of two cells, it is not clear whether the disc is in the cortex of just one or both cells. (Figure and methods kindly provided by Jim Waddle.) one has yet determined which end of the tethered microtubules shorten during rotation. Consistent with this possibility, a kinesin-like immunoreactivity has been found localized to both the cortical site and the centrosomes (S. Hird, personal communication). Less is known about the cortical site in cells with induced spindle orientation. What we do know is that these cells show a striking concentration of actin and actin capping protein at their boundaries, in a broad disc along the entire area of contact between P2 and EMS, and P3 and E (Fig. 4; J. Waddle, personal communication). A disc is also found at the area of contact between P4 and Ep, raising the possibility that these cells might interact similarly. DNC-1 is present at the P2-EMS border too, but as a spot localized to the midbody, rather than in a broad disc (Fig. 5; A. Skop, personal communication). FUTURE DIRECTIONS How Do Cell Contacts Affect Mitotic Spindle Orientation? The results cited here suggest a hypothesis for how cell contacts might affect cell division orientation: The wnt pathway may induce a dynactin complex to form in the cortex of a responding cell, either at the midbody or all along the area of contact with the inducing cell, and dynein might associate with the dynactin complex and pull astral microtubules, resulting in rotation of the centrosome-nucleus complex in line with the inducing cell. Clearly, many of the issues central to this hypothesis remain unresolved. For example, the signal presented by the inducing cells for spindle orientation is not yet known. We would expect a wingless homolog to be required in inducing cells, but none have yet been firmly implicated. How the Wnt pathway links to the cytoskeleton to affect division orientation is also an open question. The motor for rotation of the centrosome-nucleus complex has not yet been identified or localized, for either cell-autonomous or contact-induced spindle orientation, although the possibility that dy- Fig. 5. A spot of DNC-1 is present at the border of P2 and EMS. EMS is in prophase in the top figure, telophase in the bottom figure. Red is DNA, green is DNC-1 antibody, and arrows point to spots of DNC-1 at P2-EMS cell borders. (Figure kindly provided by Ahna Skop.) nein may pull microtubules toward the cortical site has been suggested (Gönczy et al., 1999b; Skop and White, 1998; Waddle et al., 1994). Whether any of the identified cortical site components are even induced by cell contact is still unclear; in fact, DNC-1 is still present at the P2-EMS border in mom-5 mutants (A. Skop, personal communication) and actin is still concentrated at the P2-EMS border in pie-1 mutants (BG, unpublished). If these suggestivelylocalized gene products in fact do play a role in contactmediated spindle orientation, then their localization must not be dependent on at least certain components of the P2-EMS signaling pathway. 28 GOLDSTEIN Is Contact-Mediated Spindle Orientation a Peculiarity of C. elegans or Is It More Widespread? One case which closely resembles the cases described in C. elegans is found in the oligochaete annelid Tubifex (Takahashi and Shimizu, 1997). In the Tubifex embryo, one cell (called CD) of the two cell stage divides unequally, as in C. elegans. As the CD cell goes through mitosis, one end of the spindle ends up close to the site of contact with the other cell. The aster at this end of the spindle has an altered morphology: the side of the aster close to the cortex appears flattened. Takahashi and Shimizu have shown that both the altered astral morphology and asymmetric division do not occur in the absence of cell contact early in the two-cell stage. Placing the other cell of the two cell stage back in contact with CD, or even placing cells of other stages in contact with CD, can rescue unequal division of CD, although curiously, only when the inducing cell is placed on the side of CD where the inducing cell originally lay. A bead the size of the inducing cell, used to mimic the alteration in cell shape caused by reassociating cells, could not rescue asymmetric division, suggesting that the cells are not affecting CD via a change in cell shape (altering cell shape in this way can in fact rescue normal division pattern in leech CD cells; Symes and Weisblat, 1992). These results suggest that cell contacts induce asymmetry of division and altered astral morphology in Tubifex. As in C. elegans, there is a concentration of actin at the area of contact between the two cells (Shimizu et al., 1998). Depolymerizing actin can prevent movement of the spindle toward the area of cell contact (Takahashi and Shimizu, 1997). Cell contacts do affect mitotic spindle orientations in organisms other than C. elegans and Tubifex, but generally by changing cell shape (Freeman, 1983; Meshcheryakov, 1978). For embryonic cells of marine shrimp, Wang et al. (1997) have made an alternative claim on the basis of observations of spindle orientations in partial embryos—that mitotic spindles in these cells are not orienting based on cell shape but instead are orienting away from areas of cell contact. It would be interesting to see whether this hypothesis holds up in controlled manipulations of cell contacts and cell shapes. Cell contacts have been proposed to orient cell divisions in plants as well, although the evidence here is not as strong. Stebbins and co-workers (Stebbins and Jain, 1960; Stebbins and Shah, 1960) have proposed that stomatal cell divisions in monocotyledonous plants are oriented by specific cell contacts, primarily on the basis that natural variation in cell positions can be correlated with variation in cell division orientations. Although contact-mediated division orientation has only rarely been shown, the more general phenomenon in which the cytoskeleton is oriented by external cues has been demonstrated in many types of cells. Examples include budding yeast, which orients polarized growth toward a source of mating pheromone (Maddox et al., 1999), the algae Fucus and Pelvetia, which orient growth of the zygote in response to unilateral light (Fowler and Quatrano, 1997), mouse early embryonic cells, which polarize components in response to cell contacts (Ziomek and Johnson, 1980), and cytotoxic and helper T cells, which orient their cytoskeletons with respect to the position of target cells (Geiger et al., 1982). Chemotaxing cells from a variety of organisms also orient their cytoskeletons with respect to a source of chemoattractant (Devreotes and Zigmond, 1988). In some of these cases, division plane is also affected, but as a secondary effect of orienting the cytoskeleton for directed secretion or migration; these cases may be valuable in suggesting candidates for genes involved in contact-mediated division orientation. In the vast majority of potential cases, whether cell divisions are oriented by cell contacts has not been tested, often because of the inaccessibility of cells to direct manipulation. Fortunately, as the number of genetic tools becomes larger, the number of cases where we can genetically alter cell fates and hence place types of cells in new positions will likely grow. This and the potential for directly manipulating cells in more systems should make it possible to test whether cells orienting their neighbors’ divisions is a common theme in multicellular organisms. ACKNOWLEDGMENTS I thank Bruce Bowerman, Paul Maddox, and Jim Waddle for comments on the manuscript; Laurie Smith for discussions of cell division orientation in plants; Steve Hird, Ann Schlesinger, Ahna Skop, and Jim Waddle for sharing unpublished results; Jim Waddle for Figure 4; and Ahna Skop for Figure 5. REFERENCES Basham SE, Rose LS. 1999. Mutations in ooc-5 and ooc-3 disrupt oocyte formation and the reestablishment of asymmetric par protein localization in two-cell Caenorhabditis elegans embryos. Dev Biol 215:253–263. C. elegans Sequencing Consortium. 1998. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282:2012–2018. Dan K. 1979. Studies on unequal cleavage in sea-urchins. 1. Migration of the nuclei to the vegetal pole development growth and differentiation 21:527–535. Dan K, Ito S. 1984. Studies of unequal cleavage in mollusks. 1. Nuclear behavior and anchorage of a spindle pole to cortex as revealed by isolation technique. Dev Growth Diff 26:249 –262. Devreotes PN, Zigmond SH. 1988. Chemotaxis in eukaryotic cells—a focus on leukocytes and dictyostelium. Ann Rev Cell Biol 4:649 – 686. Fire A, Xu SQ, Montgomery MK, Kostas SA, Driver SE, Mello CC. 1998. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806 – 811. Fowler JE, Quatrano RS. 1997. Plant cell morphogenesis: plasma membrane interactions with the cytoskeleton and cell wall. Ann Rev Cell Dev Biol 13:697–743. Freeman, G. 1983. The role of egg oganization in the generation of cleavage patterns. In: WR Jeffery, RA Raff, editors. Time, space and pattern in embryonic development. New York: Academic Press. p 171–196. INDUCED CELL DIVISION ORIENTATION Geiger B, Rosen D, Berke G. 1982. Spatial relationships of microtubule-organizing centers and the contact area of cytotoxic T lymphocytes and target cells. J Cell Biol 95:137–143. Goldstein B. 1995. Cell contacts orient some cell division axes in the Caenorhabditis elegans embryo. J Cell Biol 129:1071–1080. Gönczy P, Schnabel H, Kaletta T, Amores AD, Hyman T, Schnabel R. 1999a. Dissection of cell division processes in the one cell stage Caenorhabditis elegans embryo by mutational analysis. J Cell Biol 144:927–946. Gönczy P, Pichler S, Kirkham M, Hyman AA. 1999b. Cytoplasmic dynein is required for distinct aspects of MTOC positioning, including centrosome separation, in the one cell stage Caenorhabditis elegans embryo. J Cell Biol 147:135–150. Han M. 1997. Gut reaction to Wnt signaling in worms. Cell 90:581– 584. Herman MA, Vassilieva LL, Horvitz HR, Shaw JE, Herman RK. 1995. The C. elegans gene lin-44, which controls the polarity of certain asymmetric cell divisions, encodes a Wnt protein and acts cell nonautonomously. Cell 83:101–110. Hertwig, O. 1895. The cell. New York: Macmillan & Co. 368 p. Hibino T, Nishikata T, Nishida H. 1998. Centrosome-attracting body: a novel structure closely related to unequal cleavages in the ascidian embryo. Dev Growth Diff 40:85–95. Hyman AA. 1989. Centrosome movement in the early divisions of Caenorhabditis elegans—a cortical site determining centrosome position. J Cell Biol 109:1185–1193. Hyman AA, White JG. 1987. Determination of cell division axes in the early embryogenesis of Caenorhabditis elegans. J Cell Biol 105: 2123–2135. Keating HH, White JG. 1998. Centrosome dynamics in early embryos of Caenorhabditis elegans. J Cell Sci 111:3027–3033. Kemphues KJ, Strome S. 1997. Fertilization and establishment of polarity in the embryo. In: Riddle D, Blumenthal T, Meyer BJ, Priess JR, editors. C. elegans II. New York: Cold Spring Harbor Laboratory Press. p 335–359. Kraut R, Chia W, Jan LY, Jan YN, Knoblich JA. 1996. Role of inscuteable in orienting asymmetric cell divisions in Drosophila. Nature 383:50 –55. Lombillo VA, Stewart RJ, McIntosh JR. 1995. Minus-end-directed motion of kinesin-coated microspheres driven by microtubule depolymerization. Nature. 373:161–164. Lorson MA, Horvitz HR, van den Heuvel S. 2000. LIN-5 is a novel component of the spindle apparatus required for chromosome segregation and cleavage plane specification in Caenorhabditis elegans. J Cell Biol 148:73– 86. Lutz DA, Hamaguchi Y, Inoue S. 1988. Micromanipulation studies of the asymmetric positioning of the maturation spindle in Chaetopterus sp oocytes. 1. Anchorage of the spindle to the cortex and migration of a displaced spindle. Cell Motil Cytoskel 11:83–96. Maddox P, Chin E, Mallavarapu A, Yeh E, Salmon ED, Bloom K. 1999. Microtubule dynamics from mating through the first zygotic division in the budding yeast Saccharomyces cerevisiae. J Cell Biol 144:977–987. Maloof JN, Whangbo J, Harris JM, Jongeward GD, Kenyon C. 1999. A Wnt signaling pathway controls Hox gene expression and neuroblast migration in C. elegans. Development 126:37– 49. Mello CC, Draper BW, Krause M, Weintraub H, Priess JR. 1992. The pie-1 and mex-1 genes and maternal control of blastomere identity in early C. elegans embryos. Cell 70:163–176. Meshcheryakov VN. 1978. Orientation of cleavage spindles in pulmonate mollusks. I. Role of blastomere shape in orientation of second division spindles. Ontogenez 9:558 –566. Nishikata T, Hibino T, Nishida H. 1999. The centrosome-attracting body, microtubule system, and posterior egg cytoplasm are involved in positioning of cleavage planes in the ascidian embryo. Dev Biol 209:72– 85. 29 O’Connell KF, Leys CM, White JG. 1998. A genetic screen for temperature-sensitive cell-division mutants of Caenorhabditis elegans. Genetics 149:1303–1321. Palmer RE, Sullivan DS, Huffaker T, Koshland D. 1992. Role of astral microtubules and actin in spindle orientation and migration in the budding yeast, Saccharomyces cerevisiae. J Cell Biol 119:583–593. Rappaport, R. 1961. Experiments concerning the cleavage stimulus in sand dollar eggs. J Exp Zool 148:81– 89. Rappleye CA, Paredez AR, Smith CW, McDonald KL, Aroian RV. 1999. The coronin-like protein POD-1 is required for anterior-posterior axis formation and cellular architecture in the nematode Caenorhabditis elegans. Genes Dev 13:2838 –2851. Reinsch S, Karsenti E. 1994. Orientation of spindle axis and distribution of plasma membrane proteins during cell division in polarized MDCKII cells. J Cell Biol 126:1509 –1526. Rocheleau CE, Downs WD, Lin R, Wittmann C, Bei Y, Cha YH, Ali M, Priess JR, Mello CC. 1997. Wnt signaling and an APC-related gene specify endoderm in early C. elegans embryos. Cell 90:707–716. Rose LS, Kemphues KJ. 1998. Early patterning of the C. elegans embryo. Ann Rev Genet 32:521–545. Schlesinger A, Shelton CA, Maloof JN, Meneghini M, Bowerman B. 1999. Wnt pathway components orient a mitotic spindle in the early Caenorhabditis elegans embryo without requiring gene transcription in the responding cell. Genes Dev 13:2028 –2038. Shackleford GM, Shivakumar S, Shiue L, Mason J, Kenyon C, Varmus HE. 1993. Two wnt genes in Caenorhabditis elegans. Oncogene 8:1857–1864. Shimizu T, Ishii R, Takahashi H. 1998. Unequal cleavage in the early Tubifex embryo. Dev Growth Diff 40:257–266. Skop AR, White JG. 1998. The dynactin complex is required for cleavage plane specification in early Caenorhabditis elegans embryos. Curr Biol 8:1110 –1116. Stebbins GL, Jain SK. 1960. Developmental studies of cell differentiation in the epidermis of monocotyledons. I. Allium, Rhoeo, and Commelina. Dev Biol 2:409 – 426. Stebbins GL, Shah SS. 1960. Developmental studies of cell differentiation in the epidermis of monocotyledons. II. Cytological features of stomatal development in the Gramineae. Dev Biol 2:477–500. Strome S, Martin P, Schierenberg E, Paulsen J. 1995. Transformation of the germ-line into muscle in mes-1 mutant embryos of C. elegans. Development 121:2961–2972. Symes K, Weisblat DA. 1992. An investigation of the specification of unequal cleavages in leech embryos. Dev Biol 150:203–218. Takahashi H, Shimizu T. 1997. Role of intercellular contacts in generating an asymmetric mitotic apparatus in the Tubifex embryo. Dev Growth Diff 39:351–362. Thorpe CJ, Schlesinger A, Carter JC, Bowerman B. 1997. Wnt signaling polarizes an early C. elegans blastomere to distinguish endoderm from mesoderm. Cell 90:695–705. Waddle JA, Cooper JA, Waterston RH. 1994. Transient localized accumulation of actin in Caenorhabditis elegans blastomeres with oriented asymmetric divisions development. 120:2317–2328. Wang SW, Griffin FJ, Clark WH. 1997. Cell-cell association directed mitotic spindle orientation in the early development of the marine shrimp Sicyonia ingentis. Development 124:773–780. Wodarz A, Nusse R. 1998. Mechanisms of Wnt signaling in development Ann Rev Cell Dev Biol 14:59 – 88. Ziomek CA. Johnson MH. 1980. Cell surface interaction induces polarization of mouse 8-cell blastomeres at compaction. Cell 21:935– 942. Zwaal RR, Ahringer J, van Luenen HGAM, Rushforth A, Anderson P, Plasterk RHA. 1996. G proteins are required for spatial orientation of early cell cleavages in C. elegans embryos. Cell 86:619 – 629.