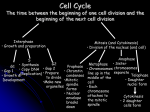
* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Here - Events
Protein phosphorylation wikipedia , lookup
Cell nucleus wikipedia , lookup
Cell encapsulation wikipedia , lookup
Endomembrane system wikipedia , lookup
Extracellular matrix wikipedia , lookup
Signal transduction wikipedia , lookup
Programmed cell death wikipedia , lookup
Cell culture wikipedia , lookup
Organ-on-a-chip wikipedia , lookup
Cellular differentiation wikipedia , lookup
Kinetochore wikipedia , lookup
Cell growth wikipedia , lookup
List of types of proteins wikipedia , lookup
Spindle checkpoint wikipedia , lookup
Cel
l
Cy
c
l
e
47S
ept
ember2015|Buda
pes
t
,
Hung
a
r
y
Da
nubi
usHea
l
t
hS
paRes
or
tMa
r
g
i
t
s
z
i
g
e
t
Or
g
a
ni
z
er
s
Bél
aNov
á
k
F
r
a
nkUhl
ma
nn
This meeting was funded by the European Molecular Biology Organization within the EMBO Workshop programme. Sponsors: Cover image kindly provided by Mark Petronczki 16th European Cell Cycle Workshop and EMBO Workshop on the Cell Cycle 4 – 7 September 2015 Danubius Health Spa Resort Margitsziget Budapest, Hungary Organizers: Béla Novák and Frank Uhlmann EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Programme overview
Friday,
4 September
2015
Saturday,
5 September
2015
Sunday,
6 September
2015
Monday,
7 September
2015
09:00
09:00 - 10:15
Meiosis
10:15
Star Auditorium
09:00 - 10:15
Cell Cycle and
Development
Star Auditorium
09:00 - 10:15
Mitotic Exit and
Cytokinesis
Star Auditorium
10:15
10:15 - 10:45
Coffee break
Jasmine corridor
10:15 - 10:45
Coffee break
Jasmine corridor
10:15 - 10:45
Coffee break
Jasmine corridor
10:45
10:45 - 12:00
Meiosis
12:00
Star Auditorium
10:45 - 12:00
Cell Cycle and
Development
Star Auditorium
10:45 - 12:00
Mitotic Exit and
Cytokinesis
Star Auditorium
12:00
12:00 - 13:00
Lunch
12:00 - 13:00
Lunch
12:00 - 12:15
Closing remarks
Star Auditorium
Platán Restaurant
Platán Restaurant
13:00 - 15:00
Poster session
Jasmine rooms
13:00 - 15:00
Poster Session
Jasmine rooms
15:00 - 16:15
S Phase
Star Auditorium
15:00 - 16:15
Mitosis
Star Auditorium
16:15 - 16:45
Coffee break
Jasmine corridor
16:15 - 16:45
Coffee break
Jasmine corridor
16:45 - 18:00
S Phase
16:45 - 18:00
Mitosis
Star Auditorium
Star Auditorium
Free time
Free time
19:00 - 22:00
19:00 - 22:00
Boat tour and dinner on the
Danube
Gala dinner
10:45
13:00
15:00
15:00
16:15
16:15
16:45
15:00 - 17:00
Arrival
and
Refreshment
Jasmine corridor
16:45
17:00
17:00 - 18:00
Keynote lecture
18:00 Star Auditorium
18:00
18:00 - 18:30
Coffee break
18:30 Jasmine corridor
19:00
19:00
20:00
20:00
22:00
18:30 - 20:00
Cell Size Control
Star Auditorium
20:00 - 22:00
Dinner and
Drinks
Platán
Restaurant
Széchenyi Terrace of
Grand Hotel
Boat Sirona
ii
12:15 - 13:15
Lunch
Platán Restaurant
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Programme
Programme
Friday, 4 September 2015
Arrival and refreshment - 15:00 - 17:00
Keynote lecture: Eric Wieschaus - 17:00 - 18:00
Jasmin corridor
Star Auditorium
Chair: Patrick O'Farrell (UCSF, San Francisco, United States)
17:00 Cell Cycle Control and the Initiation of Transcription in Drosophila Embryos
Shelby Blythe, Stefano Di Talia, Amir Momen Roknabadi, Eric Wieschaus
Molecular Biology Department, Howard Hughes Medical Institute, Princeton University
Coffee break - 18:00 - 18:30
Jasmin corridor
Cell Size Control - 18:30 - 20:00
Chair:
18:30
Star Auditorium
Bruce Edgar (University of Heidelberg, Germany)
Cell size homeostasis in mammalian cells
Clotilde Cadart, Sylvain Monnier, Matthieu Piel
Systems Biology of Cell Division and Cell Polarity, UMR 144 Institut Curie/CNRS
18:45
Dilution of the cell cycle inhibitor Whi5 controls budding yeast cell size
Kurt Schmoller, Jonathan Turner, Mardo Kõivomägi, Jan Skotheim
Department of Biology, Stanford University, Stanford
19:00
Molecular competition and cell size control
Eva Parisi, Galal Yahya, David Moreno, Carme Gallego, Attila Csikasz-Nagy, Marti Aldea
Molecular Biology Institute of Barcelona, CSIC, Barcelona, Catalonia, Spain
19:15
Regulation of cell size homeostasis in plants
Rossana Henriques, Csaba Papdi, Beátrix Horváth, Aladár Pettkó-Szandtner, Zoltán
Magyar, Laszlo Bögre
Royal Holloway, University of London, School of Biological Sciences
19:30
Mitotic control of the nuclear membrane
Snezhana Oliferenko, Maria Makarova, et al.
King's College, London
Welcome Reception and Dinner - 20:00 - 22:00
iii
Platán Restaurant
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Programme
Saturday, 5 September 2015
Meiosis - 09:00 – 12:00
Chair:
09:00
Star Auditorium
Angelika Amon (Koch Institute/HHMI/MIT, Cambridge, USA)
The final cut: control of chromosome segregation in meiosis II
Orlando Arguello-Miranda, Ievgeniia Zagoriy, Valentina Mengoli, Wolfgang Zachariae
Laboratory of Chromosome Biology, Max Planck Institute of Biochemistry, Germany
09:30
To activate, or not to activate cyclin B-Cdk1: that is the question of the
oocyte at G2/M-phase border
Daisaku Hiraoka, Takeo Kishimoto
Science & Education Center, Ochanomizu University, Tokyo, Japan
9:45
Chromosome orientation in meiosis and mitosis
Yoshinori Watanabe
University of Tokyo, Japan
Coffee break - 10:15 - 10:45
10:45
Jasmin corridor
Error-prone chromosome-mediated spindle assembly favors chromosome
segregation defects in human oocytes.
Zuzana Holubcova, Martyn Blayney, Kay Elder, Melina Schuh
MRC Laboratory of Molecular Biology, Cambridge, UK
11:15
How oocytes try to get it right: spindle checkpoint control in meiosis
Antoine Vallot, Ioanna Leontiou, Warif El Yakoubi, Damien Cladière, Eulalie Buffin,
Katja Wassmann
Institut de Biologie Paris Seine, Paris, France
11:30
Challenges of segregating chromosomes in oocytes
Martin Anger, et al.
11:45
How is bivalent cohesion maintained in oocytes arrested for months in
prophase I?
Sabrina Burkhardt, Mate Borsos, Jonathan Godwin, Takayuki Hirota, Mitinori Saitou, Paula
Cohen, Kikue Tachibana-Konwalski
Institute of Molecular Biotechnology of the Austrian Academy of Sciences, Vienna
Lunch - 12:00 - 13:00
Platán Restaurant
Poster session 1 - 13:00 - 15:00
Jasmine rooms
iv
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Programme
Saturday, 5 September 2015
S Phase - 15:00 - 18:00
Star Auditorium
Chair: Frank Uhlmann (The Francis Crick Institute, London, United Kingdom)
15:00 When DNA replication does not finish on time
Magda Magiera, Julie Petit, Nicolas Talarek, Elisabeth Gueydon, Etienne Schwob
Institute of Molecular Genetics, CNRS UMR5535 & University Montpellier, France
15:30
Visualizing developmentally programmed endoreplication in mammals and
drug-induced cell cycle modulation using ubiquitin oscillators
Atsushi Miyawaki
Riken, Japan
16:00
Developmental activation of the DNA damage checkpoint
Chames Kermi, Susana Prieto, Siem van der Laan, Nikolay Tsanov, Benedicte
Recolin, Domenico Maiorano
Institute of Human Genetics. CNRS-UPR1142. Montpellier. France
Coffee break - 16:15 - 16:45
16:45
Jasmine corridor
DNA replication control during the Mid-Blastula Transition in Xenopus
laevis
Philip Zegerman, Clara Collart, Jim Smith
Gurdon Institute, University of Cambridge, UK
17:15
A chromatin perspective of cell cycle progression in whole developing
organs
Crisanto Gutierrez
Centro de Biologia Molecular Severo Ochoa, CSIC-UAM
17:45
Entangling the function of the large number of plant CDKs - The plant
specific CYCLIN-DEPENDENT KINASE B1-CYCLIN B1 complex mediates
homologous recombination repair in Arabidopsis
Annika Weimer, Sascha Biedermann, Hirofumi Harashima, Peter Dörner, Naoki Takahashi,
Masaaki Umeda, Arp Schnittger
University of Hamburg, Department of Developmental Biology, Germany
Boat tour and dinner on the Danube - 19:00 - 22:00
v
Boat Sirona
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Programme
Sunday, 6 September 2015
Cell Cycle and Development - 09:00 - 12:00
Star Auditorium
Chair: Snezhana Oliferenko (King's College London, United Kingdom)
09:00 Growth-dependent G1/S control in Drosophila endocycles
Bruce Edgar, Norman Zielke, Jinyi Xiang, Xueyang Yu, Jonathan Bohlen
German Cancer Research Center (DKFZ), Heidelberg, Germany
09:30
Emergence of heterochromatin and prolongation of S phase in Drosophila
embryos.
Patrick H O'Farrell, Kai Yuan, Antony W Shermoen
Dept. of Biochem. and Biophys. UCSF, San Francisco, CA, US
10:00
S-phase waves of Cdk1 activity synchronize mitosis in the early Drosophila
embryo
Victoria Deneke, Anna Melbinger, Massimo Vergassola, Stefano Di Talia
Department of Cell Biology, Duke University Medical Center, Durham, USA
Coffee break - 10:15 - 10:45
10:45
Jasmin corridor
From clocks to dominoes: cell cycle remodeling during hES cell
differentiation
Silvia Santos
MRC-Imperial College, London, UK
11:15
Growth Regulation by the Hippo Pathway
Nicolas Tapon
The Francis Crick Institute, London, UK
11:45
Cytoskeletal Dynamics during Inter-nuclear Spacing in the Drosophila
Syncytial Embryo
Ojas Deshpande, Jorge de-Carvalho, Ivo Telley
Instituto Gulbenkian de Ciência, Oeiras, Portugal
Lunch - 12:00 - 13:00
Platán Restaurant
Poster session II - 13:00 - 15:00
Jasmin rooms
vi
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Programme
Sunday, 6 September 2015
Mitosis - 15:00 - 18:00
Star Auditorium
Chair: David Morgan (University of California, San Francisco, United States)
15:00 How is cohesin positioned in mammalian genomes?
Jan-Michael Peters
15:30
Pds5 proteins are key regulators of cohesin dynamics
Ana Losada, Miguel Ruiz-Torres, et al.
Chromosome Dynamics Group, Spanish National Cancer Research Centre, Madrid, Spain
16:00
Mitotic chromosomes acquire mechanical independence through Ki-67
Sara Cuylen, Claudia Blaukopf, Thomas Müller-Reichert, Beate Neumann, Ina Poser, Jan
Ellenberg, Anthony A. Hyman, Daniel W. Gerlich
Institute of Molecular Biotechnology of the Austrian Academy of Sciences, Vienna, Austria
Coffee break - 16:15 - 16:45
16:45
Jasmin corridor
Effects of aneuploidy on fitness and tumorigenesis
Angelika Amon, et al.
Koch Institute/HHMI/MIT, Cambridge, USA
17:15
Integration of microtubule-kinetochore attachment formation and spindle
checkpoint signalling by phosphatase complexes
Ulrike Gruneberg
University of Oxford, UK
17:45
Functions of Greatwall-PP2A/B55 pathway in the mammalian cell cycle
Mónica Alvarez-Fernández, María Sanz-Flores, Belén Sanz-Castillo, Marcos Malumbres
Cell Division and Cancer Group, CNIO, Madrid, Spain
Gala dinner - 19:00 - 22:00
Széchenyi Terrace of Grand Hotel
vii
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Programme
Monday, 7 September 2015
Mitotic Exit and Cytokinesis - 09:00 - 12:00
Star Auditorium
Chair: Silvia Santos (MRC-Imperial College, London, United Kingdom)
09:00 Mechanisms of mitotic regulation by the APC/C
Dan Lu, Juliet Girard, Arda Mizrak, David Morgan
University of California, San Francisco
09:30
Deciphering the role of the GWL/ARPP19-ENSA/PP2A pathway in the
control of the cell cycle
Anna Castro, et al.
CRBM-CNRS, Montpellier, France
10:00
PP2A-B55 is an NLS-directed Cdk-counteracting phosphatase coordinating
mitotic spindle reorganisation with nuclear envelope reformation and
cytokinesis.
Michael Cundell, Ricardo Nunes Bastos, Elena Poser, Shabaz Mohammed, Francis Barr
University of Oxford, Department of Biochemistry
Coffee break - 10:15 - 10:45
10:45
Jasmin corridor
Spindle pole body specification in budding yeast
Jette Lengefeld, Manuel Hotz, Yves Barral
Institute of Biochemistry, Department of Biology, ETH Zürich, Switzerland
11:15
A novel protein network stabilizes the yeast septin ring upon mechanical
stress
Laura Merlini, Maria Angeles Juanes, Alessio Bolognesi, Yves Barral, Simonetta Piatti
Centre de Recherche en Biochimie Macromoléculaire-CNRS, Montpellier, France
11:45
Spastin and ESCRT-III coordinate mitotic spindle disassembly and nuclear
envelope sealing.
Marina Vietri, et al.
Department of Molecular Cell Biology, Oslo University Hospital, Norway
Closing Remarks - 12:00 - 12:15
12:00
Star Auditorium
Closing Remarks
Tim Hunt
Lunch - 12:15 - 13:15
Platán restaurant
viii
ABSTRACTS
OF
ORAL PRESENTATIONS
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Keynote lecture
Cell Cycle Control and the Initiation of Transcription in
Drosophila Embryos
Shelby Blythe1, Stefano Di Talia1, 2, Amir Momen Roknabadi1, Eric Wieschaus1
1
2
Molecular Biology Department, Howard Hughes Medical Institute, Princeton University
Department of Cell Biology, Duke University School of Medicine
Following fertilization in Drosophila, the embryo undergoes 13 synchronous nuclear mitoses. It then
pauses cell cycle progression to partition the nuclei into individual cells and to assign those cells
distinct developmental fates. The pause in cell cycle is analogous to the midblastula transition
observed in many organisms and is associated with the degradation of a subset of maternally
supplied mRNAs and large-scale transcriptional activation of the genome. In my talk, I will describe
experiments that investigate the remodeling of the rapid maternally controlled cell cycle into the
extended cell cycle characteristic of later stages, the role of the DNA replication checkpoint in this
process, and the relationship between this cell cycle transition and the restructuring of chromatin
associated with the transcriptional activation.
Using interphase duration as an assay for checkpoint activity, we have shown that activation of the
checkpoint depends not on total DNA content, but instead on the dosage of DNA that is
transcriptionally engaged. We have extended these observations using ChIP-seq to reconstruct the
genome-wide distribution of RNA Polymerase II (Pol II). We find that large-scale de novo recruitment
of poised Pol II to thousands of sites coincides with activation of the MBT checkpoint, and that the
checkpoint itself is not required for this initial recruitment. Instead we see that loci occupied by Pol II
at the MBT recruit a sensor of replication stress generally thought to be upstream of checkpoint
activation (Rp-A70). Reducing Pol II recruitment in zelda mutants reduces recruitment of Rp-A70 and
partially rescues the lethality associated with checkpoint mutants. These results suggest a model
where checkpoint activation and timing of the MBT is triggered by the large-scale zygotic genome
activation.
The cell cycle pause at cycle 14 is associated with a downregulation of the Drosophila homologues of
the cdc25 phosphatase, String and Twine. Re-initiation of cell division occurs in a defined temporal
sequence at gastulation and requires renewed zygotic transcription of cdc25-String. I will describe
genetic screens designed to identify the global and region specific transacting elements that time the
re-initiation of these divisions.
1
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Cell Size Control
Cell size homeostasis in mammalian cells
Clotilde Cadart1, 2, Sylvain Monnier1, Matthieu Piel1
1
2
Systems Biology of Cell Division and Cell Polarity, UMR 144 Institut Curie/CNRS, 26 rue d’Ulm 75248 Paris Cedex 05 France
Univ Paris-Sud
The way mammalian cells coordinate cell growth and cell cycle progression in order to correct for
potential mistakes at division and maintain cell size homeostasis is not yet known. Although often
hypothesized, the requirement of a size checkpoint similar to that in S. Pombe has not been
confirmed experimentally.
To monitor cell growth over several cell cycles we developed two volume measurement methods.
The first method is based on fluorescence-exclusion and enables precise growth rate measurement
throughout the cell cycle. The second method uses confinement inside micro-channels and enables
inducing asymmetric divisions to assess potential size-correction mechanisms during the following
cell cycle. The combination of these two methods allowed characterizing the homeostasis process in
different cell types. We show that, in HeLa, MDCK and HT29 cells, there is no apparent cell size
checkpoint and that cells instead maintain size homeostasis through a constant volume increase at
each cell division cycle, independent of their volume at birth. This process, similar to what was
recently observed in bacteria (1) enables the progressive reduction of errors in volume segregation.
In one other cell type, Raji cells, the homeostasis mechanism appears less efficient and the amount
of volume gained is partly dependent on initial size, with big cells gaining more volume than smaller
ones.
In HeLa, MDCK and HT29 cells, the central mechanism allowing homeostasis seems to rely on the
fact that cells with smaller growth rate (which is often the case of smaller cells) have longer cell
division cycle timing and thus eventually grow as much than cells with larger growth rates. In Raji
cells, the same inverse correlation between volume at birth and cell cycle length is found but it is not
sufficient to allow small cells to gain the same amount of volume than big cells.
We are now combining volume measurement with complementary techniques to assess cell dry
mass (quantitative phase) and cell cycle phase. This allows us to measure all the key parameters of
cell growth. These measures are then performed on cells grown in various conditions such as
confinement, osmotic shocks, rapamycin treatment or nutrient depletion to perturb cell growth or
cyclin inhibitors to perturb cell cycle progression.
With this approach we will be able to understand the coupling between initial cell size, growth rate
and cell cycle duration that leads to size homeostasis in cultured cells. We also aim at testing the
robustness of this homeostasis mechanism in various environments and identify conditions in which
size is not maintained.
Reference:
(1) Campos, M., Surovtsev, I. V., Kato, S., Paintdakhi, A., Beltran, B., Ebmeier, S. E., & Jacobs-Wagner,
C. (2014). A Constant Size Extension Drives Bacterial Cell Size Homeostasis. Cell, 159(6), 1433–1446.
doi:10.1016/j.cell.2014.11.022
2
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Cell Size Control
Dilution of the cell cycle inhibitor Whi5 controls budding yeast
cell size
Kurt Schmoller, Jonathan Turner, Mardo Kõivomägi, Jan Skotheim
1Department of Biology, Stanford University, Stanford, CA 94305, USA
Cell size fundamentally affects all biosynthetic processes by determining the scale of organelles and
influencing surface transport. Although extensive studies have identified many mutations affecting
cell size, the molecular mechanisms underlying size control have remained elusive. In budding yeast,
size control occurs in G1 phase prior to Start, the point of irreversible commitment to cell division. It
was previously thought that activity of the G1 cyclin Cln3 increased with cell size to trigger Start by
initiating the inhibition of the transcriptional inhibitor Whi5. Our work overturns this model. While
Cln3 concentration does modulate the rate at which cells pass Start, its synthesis increases in
proportion to cell size so that its total concentration is constant during pre-Start G1. Rather than
increasing Cln3 activity, we identify decreasing Whi5 activity — due to the dilution of Whi5 by cell
growth — as a molecular mechanism through which cell size controls proliferation. Whi5 is
synthesized in S/G2/M phases of the cell cycle in a largely size-independent manner. This results in
smaller daughter cells being born with higher Whi5 concentrations that extend their pre-Start G1
phase. Thus, at its most fundamental level, budding yeast size control results from the differential
scaling of Cln3 and Whi5 synthesis rates with cell size. More generally, our work shows that
differential size-dependency of protein synthesis can provide an elegant mechanism to coordinate
cellular functions with growth.
3
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Cell Size Control
Molecular competition and cell size control
Eva Parisi1, Galal Yahya1, David Moreno1, Carme Gallego1, Attila Csikasz-Nagy2, Marti
Aldea1
1
2
Molecular Biology Institute of Barcelona, CSIC, Barcelona, Catalonia, Spain
Institute for Mathematical and Molecular Biomedicine, King's College London, London, UK
Coordination of cell growth and DNA replication ensures size adaptation and homeostasis. Cells are
able to adapt their size to growth rate both at population and single-cell levels, which suggests that
growth is intimately linked to the molecular mechanisms that govern the cell cycle. Budding yeast
cells, as most eukaryotic cells, exert this coordination essentially during G1, where a critical size must
be attained before cells trigger Start. The most upstream activator of cell cycle entry in budding
yeast is Cln3, a cyclin that critically depends on molecular chaperones to accumulate in the nucleus
in late G1. On the other hand, chaperones are massively involved in key growth processes, and we
have investigated the possibility that coordination between proliferation and growth relies on the
competition for limiting stocks of shared chaperones. As deduced from mathematical modeling, a
molecular competition device would be able to (1) accumulate Cln3 in the nucleus in a non-linear
manner during G1, and (2) trigger Start at a cell size that is proportional to growth rate. We have
used different experimental approaches to test predictions of the model, and found that availability
of chaperones is negatively correlated with growth rate. More important, chaperone availability
increases during G1 as cells grow, thus emerging as a new strong candidate for the sizer mechanism.
Our data support the notion that competition for molecular chaperones would provide Start with an
upstream switch-like mechanism, and adjust the critical size to the individual growth potential.
4
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Cell Size Control
Regulation of cell size homeostasis in plants
Rossana Henriques1, Csaba Papdi2, Beátrix Horváth2, Aladár Pettkó-Szandtner3, Zoltán
Magyar3, Laszlo Bogre2
1
Center for Research in Agricultural Genomics (CRAG), Consortium CSIC-IRTA-UAB-UB, Parc de Recerca UAB, Edifici CRAG,
Campus UAB, Bellaterra (Cerdanyola del Vallés), 08193 Barcelona, Spain.
2 Royal Holloway, University of London, School of Biological Sciences, Egham Hill, Egham, TW20 0EX, UK
3 Institute of Plant Biology, Biological Research Centre, Temesvári kru. 62., POB 521, H-6701, Szeged, Hungary
How cell growth and proliferation are coupled in meristematic plant cells is not well understood yet
it is pivotal to determine organ growth. Cell growth, which relies on protein translation, constrains
cell proliferation. The hypothesis is that translational control of specific mRNAs provides important
regulatory links to couple cell growth and cell proliferation. Key pieces of evidence in plants are that
silencing or overexpression of translation/ribosome associated protein EBP1 constrains or promote
cell proliferation, respectively. We propose that translation and cell proliferation are further
integrated with each other by growth regulatory signalling pathways; TOR and S6K. Emerging data
indicate that these pathways are known to affect translation as well as the commitment to DNA
replication and mitosis. We have shown that the Arabidopsis S6 Kinase1 inhibits cell proliferation
through the RBR1-E2FB complex. S6K1 interacts with RBR1 via its N-terminal RBR1 binding motif,
promotes its nuclear localization and consequent RBR1-dependent repression of cell cycle genes
through E2FB. S6K1 and E2FB are in a mutually antagonistic relationship both in their protein
abundance and in their activity. S6K1 activity to phosphorylate RBR1 is increased when E2FB is
silenced. Conversely, transcriptional activation of cell cycle genes by E2FB is elevated when S6K1 is
silenced. The same antagonism holds for S6K1 and E2FB protein abundance; S6K1 is stabilised when
E2FB is silenced, while E2FB is stabilised when S6K1 is silenced. The double inhibitory regulatory
connection between S6K1 and E2FB might constitute a regulatory switch to determine whether cells
divide or grow.
5
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Cell Size Control
Mitotic control of the nuclear membrane
Snezhana Oliferenko, Maria Makarova, et al.
King's College London
Eukaryotes remodel the nucleus during mitosis using a variety of mechanisms that differ in the
timing and the extent of nuclear envelope (NE) breakdown. We probe the principles enabling this
functional diversity by exploiting the natural divergence in NE management strategies between the
related fission yeasts Schizosaccharomyces pombe and Schizosaccharomyces japonicus. Using these
two systems, we have shown that the mitotic control of the nuclear surface area may determine the
choice between the nuclear envelope breakdown and a fully closed division. I will present our recent
work on the mechanistic basis of this variability and argue that comparative cell biology studies using
two fission yeast species could provide unique insights into physiology and evolution of mitosis.
6
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Meiosis
The final cut: control of chromosome segregation in meiosis II
Orlando Arguello-Miranda, Ievgeniia Zagoriy, Valentina Mengoli, Wolfgang Zachariae
Laboratory of Chromosome Biology, Max Planck Institute of Biochemistry, 82152 Martinsried, Germany
The generation of haploid gametes from diploid germ cells during meiosis depends on DNA
replication being followed by two rounds of chromosome segregation, called meiosis I and –II.
Whereas the mechanisms governing the segregation of homologous chromosomes in meiosis I have
been studied extensively, comparatively little is known about the control of chromatid segregation in
meiosis II. This is in part due to the perception of meiosis II as a “mitosis-like division”. However,
meiosis II differs from mitosis in fundamental aspects: in contrast to mitosis, meiosis II is preceded
not by an S-phase but by an M-phase (meiosis I) and it is not followed by another S-phase but by a
differentiation program that generates gametes. Furthermore, meiosis II chromosomes are unique
because they consist of recombined chromatids held together solely at their centromeres by cohesin
that has been protected from cleavage by the separase protease in meiosis I. Cleavage of cohesin at
entry into anaphase I requires phosphorylation of its meiosis-specific Rec8 subunit. At centromeres,
this phosphorylation is prevented by the phosphatase PP2A, which is recruited to kinetochores by
the shugoshin protein. It is currently unclear whether cleavage of centromeric cohesin in meiosis II
requires phosphorylation of Rec8 and whether this depends on the inactivation or removal from
kinetochores of PP2A. A detailed understanding of meiosis II has been hampered by technical
challenges. Experimental manipulation of meiosis II has to be performed without perturbing meiosis
I, which requires conditional methods for protein inactivation that work in the short interval
between meiosis I and –II. We use budding yeast to investigate how centromeric cohesin is cleaved
at the onset of anaphase II and how this event is coordinated with exit from meiosis II and spore
formation, the yeast equivalent of gametogenesis.
7
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Meiosis
To activate, or not to activate cyclin B-Cdk1: that is the
question of the oocyte at G2/M-phase border
Daisaku Hiraoka, Takeo Kishimoto
Science & Education Center, Ochanomizu University, Tokyo 112-8610, Japan
Some cells, particularly non-mammalian oocytes at the G2/M-phase border lack the checkpoint
control. Nevertheless, they properly respond only to physiologically relevant extracellular stimuli,
implying a mechanism that prevents unexpected cell cycle transition in response to noise stimuli.
However, it remains unclear how these cells cancel noise stimuli. Here we investigated how noise
stimuli are processed using a simple, highly synchronized binary response system, the starfish
oocyte, in which the maturation-inducing hormone 1-methyladenine (1-MeAde) causes meiotic
G2/M-phase transition via the Gβγ-PI3K-Akt pathway with no requirement of new protein synthesis;
Akt directly phosphorylates both Cdc25 and Myt1 to reverse the balance of their activities, resulting
in activation of cyclin B-Cdk1. We found that a subthreshold noise stimulus by 1-MeAde can initiate
cyclin B-Cdk1 activation. However, cyclin B-Cdk1-dependent negative feedback (Cdk-NF) causes
dephosphorylation of Akt substrates, thereby canceling noise signaling and preventing full activation
of cyclin B-Cdk1. By contrast, after suprathreshold stimuli, Gβγ activates an additional, novel
pathway that is distinct from the PI3K-Akt pathway but promotes the PI3K-Akt-dependent
phosphorylation, thereby overriding Cdk-NF and fully activating cyclin B-Cdk1. These findings
demonstrate that noise canceling is a self-regulatory decision process mediated by the final effector
of the signaling pathway.
Reference
Kishimoto, T. (2015). Entry into mitosis: a solution to the decades-long enigma of MPF (review).
Chromosoma, 124, DOI 10.1007/s00412-015-0508-y, in press.
8
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Meiosis
Chromosome orientation in meiosis and mitosis
Yoshinori Watanabe
Laboratory of Chromosome Dynamics, Institute of Molecular and Cellular Biosciences, University of Tokyo
During mitosis, replicated chromosomes (sister chromatids) become attached at the kinetochore by
spindle microtubules emanating from opposite poles and segregate equationally. In the first division
of meiosis, however, sister chromatids become attached from the same pole and co-segregate,
while homologous chromosomes connected by chiasmata segregate to opposite poles. Recent our
studies together with others have elucidated the molecular mechanisms determining chromosome
orientation, and consequently segregation, in meiosis. Comparative studies of meiosis and mitosis
lead to the general principle that kinetochore geometry and tension exerted by microtubules
synergestically generate chromosome orientation. Disorder of this regulation may lead to birth
defects and tumorigenesis in humans.
9
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Meiosis
Error-prone chromosome-mediated spindle assembly favors
chromosome segregation defects in human oocytes.
Zuzana Holubcova1, Martyn Blayney2, Kay Elder2, Melina Schuh1
1
2
MRC Laboratory of Molecular Biology
Bourn Hall Clinic
Aneuploidy in human eggs is the leading cause of pregnancy loss and several genetic disorders such
as Down syndrome. Most aneuploidy results from chromosome segregation errors during the
meiotic divisions of an oocyte, the egg's progenitor cell. The basis for particularly error-prone
chromosome segregation in human oocytes is not known. We analyzed meiosis in more than 100
live human oocytes and identified an error-prone chromosome-mediated spindle assembly
mechanism as a major contributor to chromosome segregation defects. Human oocytes assembled a
meiotic spindle independently of either centrosomes or other microtubule organizing centers.
Instead, spindle assembly was mediated by chromosomes and the small guanosine triphosphatase
Ran in a process requiring ~16 hours. This unusually long spindle assembly period was marked by
intrinsic spindle instability and abnormal kinetochore-microtubule attachments, which favor
chromosome segregation errors and provide a possible explanation for high rates of aneuploidy in
human eggs.
10
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Meiosis
How oocytes try to get it right: spindle checkpoint control in
meiosis
Antoine Vallot1, 2, Ioanna Leontiou1, 2, Warif El Yakoubi1, 2, Damien Cladière1, 2, Eulalie
Buffin1, 2, Katja Wassmann1, 2
1
2
Paris Sorbonne Universities, Institut de Biologie Paris Seine (IBPS), Paris, 75005 France
CNRS UMR7622 Developmental Biology Lab, Paris, 75005 France
In meiosis I, homologous chromosomes consisting of two sister chromatids each are segregated to
opposite poles of the bipolar spindle. This requires the orientation of sister kinetochores to the same
pole through monopolar attachment to the meiotic spindle. Because chromosomes are held
together by chiasmata, tension-generating attachments can be established. Correct attachment of
chromosomes to the bipolar spindle is verified by the spindle assembly checkpoint (SAC), as in
mitosis. In mitosis, the checkpoint not only verifies whether all sister kinetochores are attached, but
also whether those attachments are under tension. Tension is established not only between sister
kinetochores, but also in the kinetochore itself ("intra-kinetochore tension"). In the absence of
tension, an error-correction mechanism is activated. Missing tension allows the kinase Aurora B to
phosphorylate substrates that destabilize kinetochore-microtubule attachments. Tension-dependent
relocalization of Aurora B and recruitment of phosphatases to the kinetochore counterbalance
Aurora B substrate phosphorylation, leading to the stabilization of correctly attached kinetochores.
In meiosis I, the tension applied on bivalents does not lead to tension between mono-oriented sister
kinetochores, and it has been suggested that missing inter-kinetochore tension (now between pairs
of sister kinetochores) of wrongly attached chromosomes is not recognized by the meiotic SAC.
Stabilization of attachments which is dependent on BubR1-dependent PP2A localization to
kinetochores in mitosis, requires BubR1 function independent of kinetochore localization in meiosis
(Touati et al, 2015). BubR1 levels decrease with age in oocytes, therefore it is important to clarify
whether the meiotic SAC can recognize unstable attachments that are not under tension.To clarify
whether the meiotic SAC is sensitive to missing tension in mouse oocyte meiosis I, we analyzed
oocytes with shorter spindles and reduced tension, and localization of different checkpoint proteins
at tension-less kinetochores, as well as checkpoint response in oocytes without a functional SAC. Our
data show that the SAC is activated in meiosis when kinetochores are properly attached, but not
under tension. SAC activation leads to a metaphase delay and prevents timely Securin degradation
and Polar body extrusion. Bub1 but not Mad2 is recruited to tensionless kinetochores. Inactivation
of the SAC abolishes checkpoint response to loss of tension, showing that even though attachments
are monopolar, the meiotic SAC can distinguish between kinetochores that are attached, and
kinetochores that are attached and under tension.
Touati, S.A., Buffin, E., Cladiere, D., Hached, K., Rachez, C., van Deursen, J.M., and Wassmann, K.
(2015). Nat Commun 6, 6946.
11
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Meiosis
Challenges of segregating chromosomes in oocytes
Martin Anger, et al.
Department of Histology and Embryology
Faculty of Medicine, Masaryk University
Kamenice 3, building A1
62500 Brno, Czech Republic
&
Central European Institute of Technology - CEITEC
Veterinary Research Institute
Hudcova 70
Brno 621 00
Czech Republic
Mammalian oocytes and embryos are frequently affected by aneuploidy arising during chromosome
segregation. The reason why chromosome segregation errors are more frequent in germ cells and
embryos in comparison to somatic cells is unknown. We believe that the problem lies in less
stringent chromosome segregation control mechanisms operating in these cells. In our study we
focused on a functional analysis of a surveillance checkpoint mechanism called Spindle Assembly
Checkpoint (SAC) in live oocytes. Using micromanipulation and live cell confocal microscopy, we
have tested whether the SAC is capable to prevent metaphase to anaphase transition in oocytes in
which the chromosomes are not congressed properly or in which single chromatids are present in
meiosis I. Similar approach was used for monitoring the activity of SAC on individual kinetochores
and for correlation of this activity with chromosome movements, spindle formation, onset of
Anaphase Promoting Complex (APC) activation and polar body extrusion (PBE) at various time points
during the first meiotic division. Our results show important and unexpected differences in SAC
function in oocytes compared to somatic cells. In contrast to the somatic cells, oocytes were unable
to mount functional SAC response and to prevent anaphase onset in presence of unaligned
chromosomes or single chromatids. Our results indicate that the checkpoint mechanisms involved in
monitoring chromosome segregation, are in oocytes unable to detect or respond to errors in this
process, which might explain a high incidence of aneuploidy in these cells.
Project support: This work was supported by Czech science foundation projects P502/12/2201 and
15-04844S and by MEYS projects ED1.1.00/02.0068—CEITEC and LH 13072—Kontakt II.
12
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Meiosis
How is bivalent cohesion maintained in oocytes arrested for
months in prophase I?
Sabrina Burkhardt1, Mate Borsos1, Jonathan Godwin2, Takayuki Hirota3, Mitinori
Saitou3, Paula Cohen4, Kikue Tachibana-Konwalski1
1
IMBA, Institute of Molecular Biotechnology of the Austrian Academy of Sciences
University of Oxford
3 Kyoto University
4 Cornell University
2
It is presumed that sister chromatid cohesion is established during DNA replication in embryonic
oocytes and maintained during a prolonged prophase I arrest until its destruction at the meiotic
divisions in adult female mammals. Rec8-containing cohesin complexes, which may contain Smc1β,
maintain bivalent cohesion that is essential for chromosome segregation. Unlike Rec8, Smc1β is
nonessential for cohesion establishment but its absence causes premature cohesion loss and
infertility. Conditional deletion of Smc1β using Gdf9-iCre thought to be active after birth does not
affect fertility, suggesting its transcription is not required for cohesion maintenance. Whether new
cohesive structures strengthen meiotic cohesion in oocytes arrested for months or decades at the
dictyate stage of meiosis I remains unknown. Since both DNA synthesis-dependent and independent mechanisms can generate cohesive structures entrapping sister chromatids, cohesion
might be maintained with or without building additional structures in arrested oocytes. To
distinguish between these, the ability of Rec8 activated at different meiotic stages to generate
functional cohesion is tested by TEV protease cleavage and live-cell imaging. Genetic activation of
Rec8 using Cre recombinase under the Gdf9 or Spo11 promoter results in functional cohesion, likely
due to cohesin loading during DNA replication. In contrast, timely controlled Rec8 activation in
arrested oocytes before growth using tamoxifen-inducible Dppa3-Mer-Cre-Mer consistently does
not rescue cohesion despite new cohesin transcription. We conclude that cohesion is maintained
after birth without building new Rec8-cohesive structures in arrested mouse oocytes. Our results
imply that women’s fertility depends on the longevity of cohesin complexes that become cohesive
on chromosomes in oocytes in utero.
13
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
S phase
When DNA replication does not finish on time
Magda Magiera, Julie Petit, Nicolas Talarek, Elisabeth Gueydon, Etienne Schwob
Institute of Molecular Genetics, CNRS UMR5535 & University Montpellier, France
It was long assumed that cells would not enter mitosis until chromosome replication is completed.
This concept stems almost entirely from studies in which the progression of replication forks was
affected by chemical or enzymatic impediments. It is now well established that, in such case, long
stretches of single-stranded DNA are generated, which activate the Mec1/ATR checkpoint pathways
that delay mitosis until the problem is fixed.
We devised two strategies to address what happens when S phase is still under progress, without
fork perturbation, when cells are programmed to enter mitosis. In the first strategy we postponed S
phase to a later point in the cell cycle, so that its completion temporally overlaps with mitosis. This
indicated that physiological DNA replication can delay mitosis through activation of both Mec1/ATR
and Mad2/SAC checkpoint mechanisms, thus preserving genome integrity. Interestingly Mec1 and
Mad2 become together essential in cells where S phase and mitosis are triggered by the same set of
cyclin-Cdk, revealing an evolutionary driving force for checkpoint emergence.
In the second strategy we extended S phase by lowering the number of active origins. Such cells
enter mitosis with under-replicated DNA, undetected by checkpoints, and become highly genetically
unstable. The DNA damage response (DDR) is activated only when cells are allowed to progress past
the G2/M transition, irrespective of the amount of unreplicated DNA, indicating that there is no
checkpoint monitoring the completion of S phase. We will discuss the mechanism by which DNA
replication is completed during mitosis and how this leads to genome instability.
Altogether our work underscores the key role of origin licensing and replication fork density in the
maintenance of genome integrity. It challenges the dogma that cells cannot enter mitosis before S
phase is finished and provides a mechanism for the breakage of late-replicating fragile sites. The
mechanisms we uncovered in yeast offer a conceptual framework to explain the high chromosomal
instability of cancer cells with altered replication programmes.
14
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
S phase
Visualizing developmentally programmed endoreplication in
mammals and drug-induced cell cycle modulation using
ubiquitin oscillators
Atsushi Miyawaki
RIKEN Brain Science Institute
RIKEN Center for Advanced Photonics
We used transgenic mice expressing Fucci (Fluorescent Ubiquitination-based Cell Cycle Indicator)
probes that report the activity of APCCdh1 and SCFSkp2, to visualize spatio-temporal dynamics of
altered cell-cycle progression. For example, trophoblast giant cells (TGCs) in the placenta become
polyploid through endoreduplication (bypassed mitosis), and megakaryocytes (MKCs) in the bone
marrow become polyploid through endomitosis (abortive mitosis). By performing long-term, high
temporal-resolution Fucci imaging, we were able to visualize reciprocal activation of APCCdh1 and
SCFSkp2 in differentiating TGCs and MKCs grown in our custom-designed culture wells. We found
that TGCs and MKCs both skip cytokinesis, but in different ways, and that the reciprocal activation of
the ubiquitin oscillators in MKCs is varied with the polyploidy level. We also obtained threedimensional reconstructions of highly polyploid TGCs in whole, fixed mouse placentas.
We also used population and time-lapse imaging analyses of cultured cancer cells expressing Fucci,
to find great diversity in the cell-cycle alterations induced by anticancer drugs. Drug-induced cell
cycle modulation varied not only between different cell types or following treatment with different
drugs, but also between cells treated with different concentrations of the same drug or following
drug addition during different phases of the cell cycle. By combining cytometry analysis with the
Fucci probe, we have developed a novel assay that fully integrates the complexity of cell cycle
regulation into drug discovery screens. This assay system will represent a powerful drug-discovery
tool for the development of the next generation of anti-cancer therapies.
15
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
S phase
Developmental activation of the DNA damage checkpoint
Chames Kermi, Susana Prieto, Siem van der Laan, Nikolay Tsanov, Benedicte
Recolin, Domenico Maiorano
Institute of Human Genetics. CNRS-UPR1142. Montpellier. FRANCE
The cell cycles of early embryos are very rapid due to contraction of gap phases and reduced S phase
length. In this highly contracted cell cycle the DNA damage checkpoints that restrain proliferation in
the presence of DNA damage are inefficient and may constitute an adaptation to rapid proliferation.
We have previously shown that in mouse embryonic stem cells inhibition of the G1/S checkpoint is
untimely linked to maintenance of pluripotency through developmental regulation of protein
phosphatase Cdc25A stability. We have also shown that cell cycle remodelling is essential for the
execution of a viable differentiation programme (van der Laan et al., 2013). Here we have
investigated the molecular grounds of checkpoint inhibition in the early embryonic cleavages of the
vertebrate Xenopus laevis. Previous studies have suggested that checkpoint activation is triggered by
the increase of the nucleus-to-cytoplasmic (N/C) ratio of the embryo due to absence of cell growth,
which leads to titration of maternal-limiting factors, whose identity has been so far mysterious. We
report that the Rad18 ubiquitin ligase, a master regulator of DNA damage tolerance that promotes
monoubiquitylation of the replication factor PCNA (mUb), is a critical maternal checkpoint silencing
factor in Xenopus embryos. We have observed high levels of Rad18 and PCNAmUb in Xenopus
embryos strongly decreasing prior to MBT, at a developmental stage that coincides with acquisition
of the competence to sense DNA damage. Using Xenopus cell-free extracts, we also show that Rad18
and not PCNA is depleted from egg cytoplasm and titrated on chromatin by increasing the N/C ratio.
Moreover, we show that inactivation of Rad18 function in vitro and in vivo, leads to DNA damagedependent checkpoint activation, monitored by Chk1 phosphorylation upon UV irradiation. The
molecular mechanism involves inhibition of replication fork uncoupling, a critical determinant of
checkpoint activation. Strikingly, we have observed that this embryonic-like regulation can be
reactivated in somatic mammalian cells by ectopic Rad18 expression. Finally we report high Rad18
expression in cancer stem cells and show that it is relevant to resistance to DNA damaging agents.
Altogether these data propose Rad18 as a critical checkpoint-inhibiting factor and put forward
Rad18 as novel target in sensitizing cancer stem cells to DNA damaging agents.
Reference
van der Laan S, Tsanov N, Crozet C, Maiorano D. High Dub3 expression in mouse ESCs couples the
G1/S checkpoint to pluripotency. Mol Cell. 2013 Nov
7;52(3):366-79.
16
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
S phase
DNA replication control during the Mid-Blastula Transition in
Xenopus laevis
Philip Zegerman1, Clara Collart1, 2, Jim Smith2
1
2
Gurdon Institute, University of Cambridge, UK
MRC National Institute for Medical research, London, UK
Early embryonic divisions from flies to amphibia are extremely rapid, but at a point called the midblastula transition (MBT), multiple events are coordinated including the remodelling of the cell cycle,
the onset of bulk zygotic transcription and the activation of checkpoints. How these events are
coordinated is not known. We have shown in Xenopus laevis that four DNA replication factors are
limiting for replication initiation in vivo in MBT stage embryos and are required for the
developmental activation of the kinase Chk1.
While Chk1 activation is essential for gastrulation in Xenopus, we show that this is independent of
the roles of Chk1 in controlling cell cycle checkpoints. Another kinase, DDK, is also regulated during
embryogenesis. Cdc7-Drf1 is the predominant form of DDK in early embryos, but Drf1 is replaced by
the paralogue Dbf4 at around the time of gastrulation. We show that Drf1 down-regulation is Chk1dependent at the MBT in Xenopus. This is important because knockdown of Drf1 rescues the
gastrulation defect caused by inhibition of Chk1. This suggests that we have uncovered a new
essential role for Chk1-dependent control of DDK in metazoa. The mechanisms and physiological
consequences of the developmental regulation of replication kinases will be discussed.
17
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
S phase
A chromatin perspective of cell cycle progression in whole
developing organs
Crisanto Gutierrez
Centro de Biología Molecular Severo Ochoa, CSIC-UAM, Nicolás Cabrera 1, 28049 Madrid, Spain
Cellular homeostasis during organogenesis depends on the adequate balance between different cell
populations. In the case of plants organogenesis is postembryonic and leads to the formation of new
organs in a continuous manner during the entire life of the organism. Consequently, every growing
organ requires the activity of stem cells, the proliferation of their derivatives, the decrease in cell
division competence and the initiation of differentiation. The latter is frequently associated with the
switch to the endocycle in many cells. We are interested in understanding the crosstalk between
changes in chromatin organization and function, and cell proliferation control in the various cell
populations within a developing organ.
In the growing Arabidopsis root cell proliferation and differentiation are integrated radially and
longitudinally by gene expression patterns guided by patterning genes. We have found that after
chromatin replication in the S-phase an extensive chromatin reprogramming is at the basis of cell
population dynamics through the root compartments. Thus, the histone H3.1/H3.3 balance, which is
important for gene expression, defines a boundary within the proliferation compartment, where a
low ratio identifies cells in their last cycle before exit to differentiation. The spatial pattern of this
reprogramming process coincides with repression of genes at this proliferation boundary.
Furthermore, we have identified key patterning transcription factors that, together with other
transcription factors, regulate the coordinate expression of H3 and their chaperone genes.
Interestingly, a high H3.1/H3.3 ratio and H3.1 eviction pattern is reproduced also in the endocycling
cells before differentiation, revealing a common principle of chromatin reprogramming in
proliferating and endoreplicating cells.
The use of appropriate tools for in vivo monitoring of cell cycle progression together with highresolution gene expression data of individual cell types and screens for upstream regulators of cell
cycle are being instrumental in the identification of cellular factors that contribute to the
maintenance of an adequate balance of cell populations during organogenesis.
18
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
S phase
Entangling the function of the large number of plant CDKs - The
plant specific CYCLIN-DEPENDENT KINASE B1-CYCLIN B1
complex mediates homologous recombination repair in
Arabidopsis
Annika Weimer1, Sascha Biedermann1, Hirofumi Harashima1, Peter Dörner2, Naoki
Takahashi3, Masaaki Umeda3, Arp Schnittger1, 4
1
Department of Molecular Mechanisms of Phenotypic Plasticity, Institut de Biologie Moléculaire des Plantes du CNRS,
IBMP-CNRS - UPR2357, Université de Strasbourg, 12, Rue du Général Zimmer, 67084 Strasbourg Cedex, France.
2 University of Edinburgh, Mayfield Road, Edinburgh EH9 3JR, Scotland, UK
3 Graduate School of Biological Sciences; Nara Institute of Science and Technology; Takayama; Ikoma, Nara, Japan; JST;
CRESTl 8916-5 Takayama; Ikoma, Nara, Japan.
4 University of Hamburg, Biozentrum Klein Flottbek, Department of Developmental Biology, Ohnhorststr. 18 - 22609
Hamburg, Germany
A common response to DNA damage is the inhibition of cell division by inactivation of cyclindependent kinases (CDKs). Puzzlingly, recent evidence has revealed that CDK activity is required for
DNA repair pathways, especially for homologous recombination repair (HR), resulting in the
conundrum how mitotic arrest and effective repair can be reconciled. Here we show that the group
of so far ill-characterized plant specific B1-type CDKs (CDKB1) are major regulators of DNA damage
response in plants. We identify the group of B1-type cyclins (CYCB1) as the specific partners of B1type CDKs in HR and reveal that RADIATION SENSITIVE 51 (RAD51), a core component of HR, is a
substrate of CDKB1-CYCB1 complexes. Conversely, mutants in CDKB1 and all four CYCB1 genes fail to
recruit RAD51 to DNA after damage. CYCB1;1 is specifically activated under DNA damage condition
and we show that this activation is directly controlled by SUPRESSOR OF GRAMMA RADIATION 1
(SOG1), a plant specific transcription factor that acts similarly to p53 in animals. Thus, while the main
mitotic cell cycle activity is blocked under DNA damage conditions, CDKB1-CYCB1 complexes are
specifically activated to mediate HR.
19
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Cell Cycle and Development
Growth-dependent G1/S control in Drosophila endocycles
Bruce Edgar, Norman Zielke, Jinyi Xiang, Xueyang Yu, Jonathan Bohlen
German Cancer Research Center (DKFZ)-Zentrum für Molekulare Biologie der Universität Heidelberg (ZMBH) Alliance, Im
Neuenheimer Feld 282, 69120 Heidelberg, Germany
Endocycles are variant cell cycles comprised of DNA Synthesis (S)- and Gap (G)-phases but lacking
mitosis. Such cycles facilitate post-mitotic growth in many animal and plant cells. DNA replication in
endocycling Drosophila cells is triggered by Cyclin E/Cdk2 activity, but this kinase must be inactivated
during each G-phase to allow pre-Replication Complexe (preRC) assembly for the next S-phase. Using
genetic tests in parallel with computational modeling, we recently demonstrated that Drosophila’s
endocycles are driven by a molecular oscillator in which the E2F1 transcription factor promotes CycE
expression and S-phase initiation, S-phase then activates the CRL4Cdt2 ubiquitin ligase, and this in
turn mediates E2F1 destruction. We propose that it is the transient loss of E2F1 during S-phases that
creates the window of low Cdk activity required for preRC formation. In support of this model, overexpressed E2F1 accelerates endocycling, whereas a stabilized variant of E2F1 blocks endocycling by
de-regulating target genes including CycE, Cdk1 and mitotic Cyclins. Since rates of E2F1 accumulation
can control the frequency of endocycle S phases, understanding how E2F1 expression is affected by
cell growth and growth factor signaling is key to understanding the control of cell cycle progression
in these cells. In Drosophila salivary glands, we find that altering cell growth by changing nutrition or
TOR signaling impacts E2F1 translation, and that this connection makes endocycle progression
growth-dependent. Assays in S2 cells show that the 5’ UTR of E2F1 mRNA is important in mediating
its translational control, likely in response to TOR and other growth-regulatory signals. In midgut
enteroblasts, endocycling rates are also determined by E2F1 expression levels, but in this case a
critical input is EGFR/MAPK signaling. Many of the regulatory interactions essential to the growthand translation-dependent G1/S cell cycle oscillator we describe are conserved, suggesting that
elements of this mechanism may act in most growth-dependent cell cycles in animal and plant cells.
20
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Cell Cycle and Development
Emergence of heterochromatin and prolongation of S phase in
Drosophila embryos.
Patrick H O'Farrell, Kai Yuan, Antony W Shermoen
Dept. of Biochem. and Biophys. UCSF, San Francisco, CA, US
Emergence of heterochromatin and prolongation of S phase in Drosophila embryos.
Kai Yuan, Antony W. Shermoen and Patrick H. O’Farrell
The first event in the slowing of the cell cycle at the midblastual transition in Drosophila embryos is
not introduction of a gap phase but the prolongation of S phase. This prolongation is due to a change
in the replication from simultaneous initiation of replication all sequences to a sequential process
where each of a number of satellite sequences exhibits a stereotyped delay of initiation of
replication. Without the resulting extension of S phase in cycle 14, gastrulation is disturbed by a
premature mitosis. The onset of late replication is the result of downregulation of cyclin:Cdk1,
which, if active during interphase, drives early replication of otherwise late replicating sequences.
The late replicating satellite sequences are normally heterochromatic, but lack molecular hallmarks
of heterochromatin during the early cell cycles. We examined the onset of heterochromatin
formation and its coordination with replication using TALE-lights to tag the highly repetitive
sequence that characterizes each block of satellite sequence, and visualizing replication by the
transient recruitment of GFP-PCNA while following the onset of HP1-RFP binding. There were several
surprising and interesting outcomes.
- The initial delay in replication of the satellite sequence in cycle 14 s did not depend on prior
recruitment of HP1, which followed the slightly delayed replication of the 359 repeat and did not
occur at all during cycle 14 on the 1.686 satellite.
- The HP1 recruitment to 359 occurred only after the completion of its replication implying that the
timing of this recruitment is due to some other signal.
- Mutations introduced into HP1 showed that the early recruitment of HP1 to the 359 sequence is
independent of the chromo domain, but depends on C-terminal extension that includes a protein
interaction domain. Other foci of HP1 recruitment appeared to depend on the chromo domain
indicating diversity in the interactions that initiate heterochromatin formation.
- Absence of the 359 sequence in the mother resulted in failure to properly recruit HP1 to the
paternal 359 sequence showing that a sequence specific maternal signal promotes embryonic
establishment of heterochromatin on this satellite.
- In the absence of HP1 binding, replication timing of a satellite was advanced in cycle 15, and
reciprocally, it was delayed by artificial recruitment of HP1 using a TALE fusion, showing that HP1
binding impacts replication timing.
21
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Cell Cycle and Development
S-phase waves of Cdk1 activity synchronize mitosis in the early
Drosophila embryo
Victoria Deneke1, Anna Melbinger2, Massimo Vergassola2, Stefano Di Talia1
1
2
Department of Cell Biology, Duke University Medical Center, Durham, NC 27710, United States
Department of Physics, University of California San Diego, La Jolla, CA 92093, United States
Since the classical work of Fisher and Kolmogorov in 1930s, chemical waves have been proposed as a
mechanism to coordinate events in biological systems. Chemical waves have the potential of
transferring information across large spatial scales very rapidly. However, the importance of such
waves during embryonic development remains unclear. In the earliest phases of development of
most metazoans, mitoses are organized across spatial scales of about 1mm. It has been proposed
that trigger waves of Cdk1 activity might play a role in coordinating mitosis in embryos. However,
the physical properties of these waves and the molecular mechanisms regulating them remain
unclear. Using quantitative live imaging of a FRET biosensor for the activity of Cdk1, we provide a
direct visualization of traveling waves of Cdk1 activity in Drosophila embryos. We demonstrate that
the speed of the mitotic waves is not regulated by the activity of Cdk1 during mitosis, but by the
activity of Cdk1 during S-phase. Consistently, the progressive slowdown of the mitotic waves
observed during development is regulated by the activity of the DNA replication checkpoint through
the Chk1/Wee1 pathway. Mathematical modeling of Cdk1 and Chk1 activities and ablation
experiments indicate that a trigger wave in S-phase is followed by a phase wave that regulates entry
into and exit from mitosis. Theoretical arguments argue that the scaling of the speed with Cdk1
activity display universal signatures of the dynamic of a bistable system, in which waves are initiated
by stochastic fluctuations. Our findings demonstrate how the physical properties of chemical waves
can be modulated by the dynamics of signaling pathways to coordinate cellular events during
development.
22
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Cell Cycle and Development
From clocks to dominoes: cell cycle remodeling during hES cell
differentiation
Silvia Santos
MRC-Imperial College London
During cell decision-making signal transduction networks dynamically change in time and space in
response to cues, and thereby trigger different cellular states. The decision to divide is one of the
most fundamental cellular responses and the evolutionarily conserved networks that control cell
division adapt and remodel in a variety of biological contexts - during development and homeostasis,
infections and malignancy, in response to drugs and stresses. A striking example of this versatility
occurs during development where the same core regulators drive structurally different divisions.
Divisions in the embryo are clock-like, fast, short and synchronous with no checkpoints or gap
phases. With time, these divisions become longer and asynchronous. The resulting somatic like
cycles have checkpoint control and gap phases, and the initiation of events is dependent on
completion of early events, just like a falling domino. The question, thus, arises on how do the same
cell cycle regulators self-organize and remodel in time and space to generate structurally different
cell division cycles?
In our lab we use human embryonic stem cells as a model for the embryonic cell cycle and monitor
the activities, concentrations and spatial distribution of key cell cycle regulators in single cells, during
ES cell differentiation. In my talk I will be discussing how combining single cell imaging and omics
approaches with mathematical modelling is allowing us to shed light into how cell cycle networks
remodel in time and space during embryonic to somatic transition.
23
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Cell Cycle and Development
Growth Regulation by the Hippo Pathway
Nicolas Tapon
Francis Crick Institute
Lincoln's Inn Fields Laboratory
44 Lincoln's Inn Fields
London WC2A 3LY
Genetic screens in Drosophila melanogaster have identified the Hippo (Hpo) pathway as one of the
major signalling pathways required for tissue size control. The Hpo pathway controls tissue and
organ size by both inhibiting cell proliferation and promoting apoptosis. Subsequent studies in
mammals have shown that this growth control function is conserved and that Hpo signalling is
dysregulated in many types of cancer. At the core of the Hippo pathway lies a kinase cascade
comprising the Ste20-related kinase Hpo and the Dbf2-related kinase Warts (Wts). Upon Hpo
activation, the downstream kinase Wts phosphorylates and inhibits the pro-growth transcriptional
co-activator Yorkie (Yki). Hpo signalling has been proposed to sense various local cues relating to cell
density (contact inhibition of growth and mechanical tissue properties) or patterning (morphogen
gradients), and translate these cues into a growth arrest signal once an individual tissue has reached
its appropriate size. I will discuss our latest findings on the relationship between tissue growth and
mechanics, as well as the control of the Hpo pathway by nutritional signals.
24
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Cell Cycle and Development
Cytoskeletal Dynamics during Inter-nuclear Spacing in the
Drosophila Syncytial Embryo
Ojas Deshpande, Jorge de-Carvalho, Ivo Telley
Instituto Gulbenkian de Ciência, Oeiras, Portugal
The positioning of the nucleus, the central organelle of the cell, is an active and regulated process
crucially linked to cell cycle, differentiation, migration, and polarity. Alterations in positioning have
been correlated with cell and tissue function deficiency and genetic or chemical manipulation of
nuclear position is embryonic lethal. Nuclear positioning plays a key role during early embryonic
developmental stages in insects, such as Drosophila, where the nuclei divide without cytokinesis and
spread evenly throughout the large syncytial embryo cell. Little mechanistic insight is presently
available due to the high opacity caused by yolk within eggs and early embryos, which has restricted
the microscopic study of nuclear positioning in pre-blastoderm embryos to chemically fixed samples
only.
We use a novel technique whereby we create cytoplasmic explants from pre-blastoderm stage
embryos of Drosophila that reconstitute nuclear divisions. Using time-lapse microscopy and image
processing we quantify the cytoskeletal dynamics and nuclear positioning and dissect the forcegenerating structures that lead to nuclear movement. It is intriguing that the daughter nuclei
originating from adjacent mitotic divisions remain separated, resulting in a uniform nuclear
arrangement in the absence of compartmentalisation. Astral microtubules play a key role in
separation of daughter nuclei during mitosis. We hypothesise that microtubule-mediated repulsion
maintains the minimum distance between the neighbouring nuclei, creating nuclear domains. Using
transgenic fly lines and high-resolution imaging we are investigating the localisation and function of
candidate microtubule interacting proteins (Feo) and molecular motors (Klp3a, Klp61f) that are
known to generate stable microtubule overlaps. We show that microtubule overlaps between asters
from neighbouring nuclei do occur and Klp61F localises along the aster microtubules. We are now
testing if these candidate proteins are preferentially enriched at these overlaps.
25
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitosis
How is cohesin positioned in mammalian genomes?
Jan-Michael Peters
Research Institute of Molecular Pathology (IMP), Vienna
During S-phase newly synthesized “sister” DNA molecules become physically connected. This sister
chromatid cohesion resists the pulling forces of the mitotic spindle and thereby enables the biorientation and subsequent symmetrical segregation of chromosomes. Cohesion is mediated by
ring-shaped cohesin complexes, which are thought to entrap sister DNA molecules topologically. In
mammalian cells, cohesin is loaded onto DNA at the end of mitosis by the NIPBL-MAU2 (Scc2-Scc4)
complex, becomes acetylated during S-phase, and is stably “locked” on DNA during S- and G2-phase
by sororin. Sororin stabilizes cohesin on DNA by inhibiting Wapl, which can otherwise release
cohesin from DNA again.
In addition to mediating cohesion, cohesin also has important roles in organizing higher-order
chromatin structures and in gene regulation. Cohesin performs the latter functions in both
proliferating and post-mitotic cells and mediates at least some of these together with the sequencespecific DNA-binding protein CTCF, which co-localizes with cohesin at many genomic sites. Cohesin is
required for long-range chromatin interactions, which can either support or suppress enhancerpromoter interactions, and is thought to contribute to the folding of chromosomes into topologically
associated domains (TADs).
Whereas cohesin might be able to mediate cohesion by connecting any region of sister chromatids,
it is important that cohesin associates with specific binding sites in the genome to be able to
contribute to higher-order chromatin structure and gene regulation. We therefore analyzed how
cohesin is positioned in the mouse and human genome. Our results indicate that cohesin is initially
recruited to DNA at NIPBL-MAU2 binding sites, which are distinct from cohesin’s final binding sites.
Wapl enables reversible interactions of cohesin with these sites to allow the passage of large protein
complexes that move along DNA processively, such as RNA polymerases. To obtain mechanistic
insight into how cohesin might translocate from loading to CTCF sites we have begun to reconstitute
these processes from purified components and visualized in real time how cohesin interacts with
DNA.
26
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitosis
Pds5 proteins are key regulators of cohesin dynamics
Ana Losada1, Miguel Ruiz-Torres1, et al.
1
Chromosome Dynamics Group, Spanish National Cancer Research Centre, Madrid, Spain
Bioinformatics Unit, Spanish National Cancer Research Centre, Madrid, Spain
3 Research Institute of Molecular Pathology (IMP), Vienna, Austria.
2
Cohesin is a ring-shaped complex that entraps the sister chromatids from the time they emerge from
the replication fork until their separation in anaphase, thereby ensuring both faithful chromosome
segregation and accurate DNA repair by homologous recombination. Cohesin may also promote
long-range DNA looping which contributes to transcriptional regulation and organization of DNA
replication factories. In vertebrate somatic cells, cohesin consists of Smc1, Smc3, Rad21 and either
SA1 or SA2. Cohesion factors Pds5, Wapl and Sororin bind cohesin and modulate its interaction with
chromatin. There are two versions of Pds5 in vertebrates, Pds5A and Pds5B, and in order to address
their functional specificity, we generated single and double knock out alleles for Pds5A and Pds5B.
Analyses of Mouse Embryonic Fibroblasts (MEFs) carrying these alleles showed that simultaneous
depletion of both Pds5 proteins negatively affects Wapl and Sororin binding to chromatin, therefore
altering cohesin dynamics and preventing cohesion establishment. To better characterize the role of
Pds5 proteins on cohesin’s association to chomatin, we have now performed Fluorescence Recovery
After Photobleaching (FRAP) in quiescent cells. Depletion of both Pds5 proteins clearly reduces the
mobility of cohesin, similar to the effect previously observed in MEFs lacking Wapl. Moreover, upon
serum stimulation, these cells have trouble to reenter the cell cycle and start DNA replication.
Consistent with this defect, we have observed that establishment of the pre-replication complex
(preRC) on chromatin is delayed. This delay is likely a consequence of impaired synthesis and
accumulation of preRC components including MCMs, Cdc6 and Orc1, rather than a recruitment
failure. These results underscore the importance of Pds5 in regulating cohesin dynamics and further
suggest that this dynamic behaviour is important for transcriptional regulation.
27
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitosis
Mitotic chromosomes acquire mechanical independence
through Ki-67
Sara Cuylen1, Claudia Blaukopf1, Thomas Müller-Reichert2, Beate Neumann3, Ina
Poser4, Jan Ellenberg3, Anthony A. Hyman4, Daniel Gerlich1
1
Institute of Molecular Biotechnology of the Austrian Academy of Sciences (IMBA), Vienna, Austria.
Experimental Center, Medical Faculty Carl Gustav Carus, Technische Universität Dresden, Dresden, Germany
3 Cell Biology and Biophysics Unit, European Molecular Biology Laboratory (EMBL), Heidelberg, Germany
4 Max Planck Institute of Molecular Cell Biology and Genetics, Dresden, Germany
2
Eukaryotic genomes are partitioned into chromosomes, which during cell division form spatially
separate compact bodies to move one copy of the replicated genome to each daughter cell. How
mitotic chromosomes acquire the biophysical properties enabling their independent motility is
poorly understood. Here, we report that the chromosome surface is cell cycle-regulated.
Chromosomes have low adhesion towards each other during early mitotic stages, whereas they
cluster during mitotic exit to shape one mass of chromatin prior to nuclear envelope reassembly. We
identified Ki-67 as a critical factor required to maintain mitotic chromosomes spatially separate,
which is necessary for their independent motility and timely segregation. Our study elucidates a
biomechanical role of the mitotic chromosome periphery and uncovers an important function of Ki67, a widely used cancer prognostic marker.
28
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitosis
Effects of aneuploidy on fitness and tumorigenesis
Angelika Amon, et al.
Koch Institute for Integrative Cancer Research
Howard Hughes Medical Institute
Massachusetts Institute of Technology
76-561
500 Main Street
Cambridge MA 02139
Aneuploidy, a karyotype that is not a multiple of the haploid complement, is a hallmark of cancer. 90
percent of all solid human tumors harbor an incorrect karyotype. Thus, determining how aneuploidy
arises and how it impacts cellular behavior is critical for our understanding of tumorigenesis.
We developed in vitro and in vivo mouse models to study the effects of aneuploidy on cellular
fitness and tumorigenesis. Using hematopoietic reconstitution assays we find that aneuploidy leads
to decreased fitness of stem cells in vivo. Their ability to to proliferate and reconstitute the
hematopoietic system in vivo is significantly reduced leading to macrocytopenia and bone marrow
failure. Our studies further indicate that aneuploidy also reduces the fitness of immortalized and
transformed cells. Thus, aneuploidy has a significant impact on fitness in all contexts. The
implications of these findings for tumorigenesis and ways in which aneuploidy could promote
tumorigenesis despite its anti-proliferative effects will be discussed.
29
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitosis
Integration of microtubule-kinetochore attachment formation
and spindle checkpoint signalling by phosphatase complexes
Ulrike Gruneberg
University of Oxford
Faithful chromosome segregation in eukaryotic cells requires the formation and maintenance of
stable microtubule-kinetochore attachments. The spindle assembly checkpoint (SAC) is an important
regulatory mechanism ensuring the presence of correctly attached chromosomes before anaphase
onset. Both timely activation as well as inactivation of the SAC are key prerequisites for the
successful orchestration of chromosome segregation. The Mps1 kinase plays a crucial role in
triggering and maintaining SAC signalling by phosphorylating the Knl1 kinetochore protein, thus
forming a binding platform for the key SAC proteins Bub1, Bub3 and BubR1. Prompt silencing of the
SAC upon bipolar attachment thus requires a phosphatase activity opposing Mps1, initiating the loss
of Bub1, Bub3 and BubR1 from the kinetochore. In yeast, PP1 has been shown to fulfil this function.
We have taken an unbiased approach to identify the phosphatase activity controlling Knl1
phosphorylation and BubR1 and Bub1 kinetochore localisation in spindle checkpoint arrested
mammalian cells, and have recently identified BubR1-tethered PP2A-B56 as a key phosphatase
opposing Mps1 activity towards Knl1. Consequently, knock down of this phosphatase complex
resulted in retention of phospho-Knl1 as well as Bub1 and BubR1 kinetochore localisation in the
absence of Mps1 activity. We suggest that PP2A-B56 triggers anaphase onset by eliciting the loss of
BubR1 and Bub1 from kinetochores when Mps1 activity drops at the metaphase to anaphase
transition. Our latest results indicate that the same pool of PP2A-B56 also regulates Mps1 activity
itself and may control the residence time of Mps1 at the kinetochore, thus contributing to the
integration of microtubule-kinetochore attachment formation and spindle checkpoint signaling.
30
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitosis
Functions of Greatwall-PP2A/B55 pathway in the mammalian
cell cycle
Mónica Alvarez-Fernández, María Sanz-Flores, Belén Sanz-Castillo, Marcos
Malumbres
Cell Division and Cancer Group, Spanish National Cancer Research Centre (CNIO), 28029 Madrid, Spain
Protein phosphorylation is an essential mechanism to control progression throughout the different
phases of the cell cycle. During the last years, the Greatwall-PP2A axis has emerged as a new
pathway implicated in control of mitosis. Greatwall kinase, also known as Mastl (microtubuleassociated serine/threonine kinase-like protein) in mammals, specifically inhibits PP2A complexes
containing the B55 family of regulatory subunits, which in mammals is composed of 4 different
isoforms (PPP2R2A-D). Inhibition of PP2A-B55 complexes by Greatwall occurs in a kinase-dependent
manner through the phosphorylation of two small proteins, α-endosulfine and Arpp19, the only two
Greatwall substrates identified to date.
We previously generated the first genetic model of Greatwall in mammals. Constitutive ablation of
Mastl in mice resulted in embryonic lethality, indicating that Greatwall activity is essential for
embryonic development. By using a Mastl conditional knockout, we also found that cells lacking
Greatwall display a mitotic collapse after nuclear envelope breakdown, characterized by
condensation defects and prometaphase arrest, accompanied by chromosome segregation defects
and a global reduction in the phosphorylation of Cdk substrates. Interestingly, these defects were
partially rescued by concomitant depletion of PP2A B55 regulatory subunits. In order to explore the
physiological relevance of PP2A/B55 complexes in mammals, we have now generated knockout
mouse models for two of those B55 regulatory subunits, PPP2R2A (B55 alpha) and PPP2R2D (B55
delta), which have been reported as the ubiquitous and cell-cycle related ones. Our preliminary data
indicates that, whereas B55 delta knockout mice are viable and do no present obvious phenotypes,
deletion of PPP2R2A (B55 alpha) compromises mouse survival. Phenotypes both at the organism and
cellular levels are currently being characterized.
Interestingly, our recent findings indicate that, beyond mitosis, Greatwall kinase is also important for
other cell cycle phases. Moreover, Greatwall is overexpressed in several human tumors and its
depletion leads to impaired proliferation of human tumor cell lines, although its relevance to cancer
is mostly unknown. On the other hand, PP2A is a well-known tumor suppressor, and its reactivation
has been proposed as a therapeutic strategy for cancer treatment. Therefore, we propose here that,
in addition to its antimitotic role, inhibition of Greatwall may also have therapeutic benefits through
the reactivation of PP2A-dependent pathways.
31
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitotic Exit and Cytokinesis
Mechanisms of mitotic regulation by the APC/C
Dan Lu, Juliet Girard, Arda Mizrak, David Morgan
University of California, San Francisco
The dramatic and beautiful events of chromosome segregation are governed by a multi-subunit
ubiquitin ligase called the anaphase-promoting complex or cyclosome (APC/C), which catalyzes the
ubiquitination and destruction of securin, cyclins, and other important regulatory proteins. The
APC/C is activated at specific cell-cycle stages by association with an activator subunit, Cdc20 or
Cdh1, which provides binding sites for specific substrate sequence motifs, or degrons. Like other
members of the RING family of E3s, the APC/C catalyzes direct ubiquitin transfer from an E2ubiquitin conjugate (E2-Ub) to lysine residues on the protein substrate.
We are using experimental and computational methods to dissect the mechanisms underlying the
ordered degradation of APC/C substrates. Our single-cell analyses of GFP-tagged APC/C substrates in
budding yeast suggest that the timing of substrate degradation is determined by changes in
substrate degrons and other mechanisms that govern substrate interaction with the APC/C.
Biochemical experiments show that degrons might not simply enhance substrate affinity for the
APC/C but also enhance the enzyme’s catalytic function: for example, mutation of a specific degron
motif in the S cyclin, Clb5, reduces maximal catalytic rate with that substrate. We are now using
computational modeling to develop a more complete understanding of how changes in substrate
affinity and catalytic rate can provide the precisely ordered degradation of different APC/CCdc20
substrates. These studies reveal that following APC/C activation, the multi-step ubiquitination
process provides an adjustable delay in the onset of substrate degradation. Small variations in
substrate affinity and/or catalytic rate can have a major impact on the timing of degradation onset,
and robust degradation timing can be achieved over a broad range of parameters. When two
substrates share the same pool of APC/CCdc20, their relative enzyme affinities and rates of turnover
can influence the partitioning of APC/CCdc20 among substrates, and under some conditions
competition between substrates can occur. However, increased expression of the early APC/CCdc20
substrate Clb5 does not delay the degradation of the later substrate securin, arguing against a role
for competition in establishing securin degradation timing. These studies provide a conceptual
framework for understanding the factors that determine the timing of substrate modification in the
cell cycle.
32
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitotic Exit and Cytokinesis
Deciphering the role of the GWL/ARPP19-ENSA/PP2A pathway
in the control of the cell cycle
Anna Castro, et al.
CRBM-CNRS
Greatwall has recently been identified as an essential kinase to promote mitotic entry and to
maintain the mitotic state. Studies in the xenopus egg extracts subsequently characterised the two
first substrates of Gwl, Arpp19-ENSA whose phosphorylation promote PP2A-B55 binding and
inhibition, and mitotic entry. We are now studying the the role of Arpp19 and ENSA on the control of
the cell cycle in mammalian cells and the function of Gwl in the control of normal and oncogenic cell
cycle.
33
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitotic Exit and Cytokinesis
PP2A-B55 is an NLS-directed Cdk-counteracting phosphatase
coordinating mitotic spindle reorganisation with nuclear
envelope reformation and cytokinesis.
Michael Cundell, Ricardo Nunes Bastos, Elena Poser, Shabaz Mohammed, Francis
Barr
University of Oxford, Department of Biochemistry
Prior to mitosis, many spindle assembly factors (SAFs) and cytokinesis-promoting factors (CYFs) are
targeted to the nucleus by polybasic nuclear localisation signals (NLS) leading to their effective
sequestration. In metazoan mitosis Cdk1:CycB mediated phosphorylation drives dissolution of the
nuclear envelope, activates liberated SAFs and inhibits CYFs, thereby simultaneously promoting
mitotic spindle formation whilst preventing precocious cytokinesis. Cdk1:CycB activity reinforces the
mitotic state by inhibiting PP2A-B55, one of the major Cdk-counteracting phosphatases, by the
Greatwall/ENSA pathway. Previously we have shown that regulation of PP2A-B55 by this pathway
during mitotic exit is crucial to explain how anaphase spindle formation and cytokinesis are initiated
later than sister chromatid separation. However, it has remained unclear how PP2A-B55 selects
specific substrates during mitotic exit, and if there are any general principles governing its action.
Using whole cell temporal profiling of protein phosphorylation we have identified 310 phosphosites
on 211 proteins that are specifically targeted by PP2A-B55 during anaphase and regulated by the
Greatwall/ENSA pathway. This work uncovers a general principle explaining substrate selection by
PP2A-B55. Analysis of specific substrates in vitro using purified components and in vivo shows that
PP2A-B55 requires a polybasic patch or NLS adjacent to the target phosphosite. This suggests that
the NLS is not only used for interphase nuclear sequestration, but also during mitotic exit to ensure
the order of cell-cycle events through timely and targeted dephosphorylation of a distinct pool of
substrates by PP2A-B55. PP2A-B55 inactivates SAFs such as TPX2, and activates CYFs including the
anaphase spindle protein PRC1 and RhoGEF ECT2 at the metaphase to anaphase transition. As a
consequence, PP2A-B55 coordinates remodelling of the metaphase spindle with anaphase central
spindle formation and cytokinesis. Moreover, PP2A-B55 dephosphorylates numerous components of
the nuclear pore complex and is necessary for timely nuclear envelope reformation. Removal of the
Greatwall/ENSA pathway therefore results in entrapment of the partly segregated chromatin and
central spindle within the confines of the nuclear envelope, as well as precocious anaphase spindle
formation and cytokinesis. Our results demonstrate that PP2A-B55 provides a means to coordinate
the temporal regulation of multiple proteins required for metazoan mitosis, by integrating nuclear
sequestration in G2, with regulation by Cdk1 in M-phase and dephosphorylation during mitotic exit.
34
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitotic Exit and Cytokinesis
Spindle pole body specification in budding yeast
Jette Lengefeld, Manuel Hotz, Yves Barral
Institute of Biochemistry, Department of Biology, ETH Zürich, 8093 Zurich, Switzerland
A number of cell types, including stem cells and budding yeast, segregate their centrosomes (or
spindle pole bodies – SPBs – in yeast) in a non-random manner at mitosis. For example, Drosophila
neuroblasts retain the young centrosome upon cell division while their differentiating daughters
inherit the old one. Likewise, budding yeast mother cells retain the new SPB and segregate the old
SPB to the bud. How these cells know which of the two centrosome/SPB is the old vs the young one,
and how they use this information to orient spindle poles appropriately along the division axis is
unknown. It is also unclear whether this process is a secondary consequence of other events, or the
result of dedicated pathways. We have started to dissect the molecular events specifying the relative
identity of the two SPBs of budding yeast through the cell cycle. These studies indicate that different
mechanisms are involved, depending on whether the old SPB of a considered cell was already the
old SPB in the previous cycle (established old SPB) or new (newly old SPB). In the second case,
phosphorylation of yeast centriolin by the wee1 kinase prior to SPB duplication is required to
instruct the “newly old” SPB to adopt the “old” identity. SPBs that have already went through a full
cell cycle as old SPB no-longer need instruction by wee1 but require activity of the NEK-related
kinase, Kin3 to maintain the old identity. Remarkably, we find that in both cases these pathways act
at least in part by promoting the recruitment of the bipartite GAP for the GTPase Tem1 to the old
SPB in early metaphase. Furthermore, ablation of the target modification sites did not cause any
other major effect than randomizing SPB inheritance, and particularly did not affect spindle
alignment with the division axis. Together, these data indicate that SPB specification and spindle
orientation is a dedicated process, the selective advantage of which remains to be determined. Our
data provide means to investigate what are the specific effects of SPB mis-orientation for cellular
physiology.
35
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitotic Exit and Cytokinesis
A novel protein network stabilizes the yeast septin ring upon
mechanical stress
Laura Merlini1, Maria Angeles Juanes1, Alessio Bolognesi2, Yves Barral2, Simonetta
Piatti1
1
2
Centre de Recherche en Biochimie Macromoléculaire-CNRS, Montpellier (France)
Institute of Biochemistry, ETH Zurich (Switzerland)
Septins are conserved GTP-binding proteins that play a critical role in cytokinesis, cell polarity and
membrane remodelling. In higher eukaryotes septin dysfunctions have been linked to tumorigenesis
and neurodegenerative diseases.
In many cell types septins assemble into filaments and rings at the neck of cellular appendages
and/or at the cleavage furrow to help compartmentalize the plasma membrane and support
cytokinesis. How septin ring assembly is coordinated with membrane remodelling and controlled by
mechanical stress at these sites is however unclear.
Through a genetic screen we uncovered an unanticipated link between the conserved Rho1 GTPase
(i.e. the yeast counterpart of metazoan RhoA) and its effector protein kinase C (Pkc1) with septin
ring stability in yeast. Both Rho1 and Pkc1 stabilize the septin ring, at least partly through
phosphorylation of the membrane-associated F-BAR protein Syp1, which colocalizes asymmetrically
with the septin ring at the bud neck. Syp1 is displaced from the bud neck upon Pkc1-dependent
phosphorylation at two serines, thereby impacting the rigidity of the new-forming septin ring. Our
data suggest that Syp1 may affect septin dynamics through the glucan synthase Fks1, which
remodels the cell wall. We propose that Rho1 and Pkc1 coordinate septin ring assembly with
membrane and cell wall remodelling partly by controlling Syp1 residence at the bud neck.
36
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Oral presentations
Mitotic Exit and Cytokinesis
Spastin and ESCRT-III coordinate mitotic spindle disassembly
and nuclear envelope sealing.
Marina Vietri, et al.
Centre for Cancer Biomedicine, Faculty of Medicine, University of Oslo, Montebello, N-0379 Oslo, Norway.
Department of Molecular Cell Biology, Institute for Cancer Research, Oslo University Hospital, Montebello, N-0379 Oslo,
Norway.
At the onset of metazoan cell division the nuclear envelope breaks down to enable capture of
chromosomes by the microtubule-containing spindle apparatus. During anaphase, when
chromosomes have separated, the nuclear envelope is reassembled around the forming daughter
nuclei. How the nuclear envelope is sealed, and how this is coordinated with spindle disassembly, is
largely unknown. Here we show that endosomal sorting complex required for transport (ESCRT)-III,
previously found to promote membrane constriction and sealing during receptor sorting, virus
budding, cytokinesis and plasma membrane repair, is transiently recruited to the reassembling
nuclear envelope during late anaphase. ESCRT-III and its regulatory AAA ATPase VPS4 are specifically
recruited by the ESCRT-III-like protein CHMP7 to sites where the reforming nuclear envelope engulfs
spindle microtubules. Subsequent association of another ESCRT-III-like protein, IST1, directly recruits
the AAA ATPase spastin to sever microtubules. Disrupting spastin function impairs spindle
disassembly and results in extended localization of ESCRT-III at the nuclear envelope. Interference
with ESCRT-III function in anaphase is accompanied by delayed microtubule disassembly,
compromised nuclear integrity and the appearance of DNA damage foci in subsequent interphase.
We propose that ESCRT-III, VPS4 and spastin cooperate to coordinate nuclear envelope sealing and
spindle disassembly at nuclear envelope-microtubule intersection sites during mitotic exit to ensure
nuclear integrity and genome safeguarding, with a striking mechanistic parallel to cytokinetic
abscission.
37
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
POSTER ABSTRACTS
Poster presentations
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 1
Deciphering the function of the CPC at the equatorial cortex in
anaphase
Ingrid Adriaans, Bas Ponsioen, Susanne Lens
Department of Molecular Cancer Research, Center for Molecular Medicine, University Medical Center Utrecht,
Universiteitsweg 100, 3584 CG Utrecht, the Netherlands
A divergent chromosome content, also known as aneuploidy, is a common feature of solid tumors.
Aneuploidy is the consequence of impaired fidelity of chromosome segregation in anaphase called
Chromosomal INstability (CIN). Tetraploidization is one of the causes of CIN and can arise due to cell
fusion, mitotic slippage or cytokinesis failure. In the following multipolar mitosis, a tetraploid cell has
an increased incidence of mis-segregating chromosomes1,2.
The Chromosomal Passenger Complex (CPC), consisting of INCENP, Survivin, Borealin and Aurora B
kinase, guards chromosomal stability through regulation of several important processes during cell
division, one of these being the proper execution of cytokinesis, thus preventing
tetraploidization3.While present at the inner centromere during prometaphase and metaphase, the
CPC relocates to the central spindle and the equatorial cortex during anaphase3. Global inhibition of
Aurora B kinase activity just before anaphase onset was shown to prevent cleavage furrow
ingression, whereas kinase inhibition in early anaphase resulted in regression of an initially fully
ingressed cleavage furrow4. How and which (i.e. central spindle or cortical) pool of the CPC controls
the onset of cytokinesis is not known. To investigate the function of the CPC at the equatorial
membrane we generated a membrane-localized FRET-based sensor of Aurora B kinase activity5.
Several membrane localization domains were tested and we found that fusion of the FRET-based
sensor to a TUBBY domain which binds PtdIns(4,5)P26, resulted in uniform localization at the cell
membrane during all stages of mitosis, including anaphase. Although Aurora B is active during
mitosis, we did not detect phosphorylation of the membrane-localized sensor in early mitosis.
Phosphorylation of the sensor at the cell membrane started in anaphase before visible membrane
ingression, and was most prominent in the equatorial furrow. Similar to FRET-based Aurora B
sensors that were targeted to chromatin, kinetochores or centromeres5,7, phosphorylation of the
membrane localized-sensor also rapidly diminished after inhibition of Aurora B. Importantly,
phosphorylation of the membrane-localized sensor was dependent on the presence of Mklp2, the
mitotic kinesin-like protein that promotes the relocation of Aurora B from centromeres to the
central spindle and equatorial cortex in anaphase.
These initial results demonstrate that the TUBBY-FRET-based Aurora B sensor is a specific tool for
detecting and measuring quantitative changes in Aurora B-dependent phosphorylation at the cell
membrane. It will allow us to elucidate how this local activity is regulated and how it is associated
with initiation of cytokinesis.
References:
1. N.J. Ganem et al., Nature 460, 278-282 (2009)
2. Silkwort et al., PLOS one 4, e6564 (2009)
3. Van der Horst et al., Chromosoma 123, 25-42 (2014)
4. Guse et al., Current biology 15, 778-86 (2005)
5. Fuller et al., Nature 453, 1132-6 (2008)
6. Szentpetery et al., BMC Cell Biology, 10:67 (2009)
7. Liu et al., Science 323, 1350-3 (2009)
38
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 2
Temporal control of mitosis: the many roles of positive
feedback
Ana Rita Araujo, Rahuman Sheriff, Silvia DM Santos
MRC-Clinical Sciences Centre, Imperial College London, London, UK
The cell cycle is characterized by a sequence of events by which a cell grows, replicates its genetic
information and its components and gives rise to two identical daughter cells. Even though the
molecular machinery that drives cell division cycles is the same in all the tissues, within the same
organism, the cell cycle length varies amongst different cells. Preliminary quantification of the length
of cell cycle phases in single cells by live-cell imaging showed high variability in the dynamics of cell
cycle phases. An exception of this was seen in mitosis, where different cells seem to keep a constant
length of time to complete this phase. Surprisingly, there is no correlation between cell cycle length
and mitosis duration. In other words, it does not matter if a cell runs through the cell cycle fast or
slow, once it reaches mitosis it will complete mitosis in a short and synchronous manner. We and
others have shown that positive feedback regulation is crucial to keep mitotic events synchronized
(Holt et al 2008, Santos et al 2012). We hypothesized that positive feedback might be the molecular
mechanism that keeps the time of mitosis constant across different cell lines with variable cell cycle
lengths. To test this hypothesis we combined live cell imaging and computational modelling and
showed that when positive feedback is perturbed the switch-like activation of Cdk1 is compromised,
leading to a much more variable and sluggish completion of mitosis. Importantly, perturbing positive
feedback gave rise to a correlation between the length of the cell cycle and duration of mitosis. This
work shows that positive feedback, a recurrent motif in cell cycle control, may also be important to
keep mitosis short, synchronous and temporally insulated from earlier cell cycle events. We
hypothesized that positive feedback may be a recurrent strategy to help keep events modular in
other biological systems.
39
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 3
Meiosis II: a unique form of cell division
Orlando Argüello-Miranda, Ievgeniia Zagoriy, Valentina Mengoli, Wolfgang Zachariae
Laboratory of Chromosome Biology, Max Planck Institute of Biochemistry, 82152 Martinsried, Germany.
Meiosis, the basis for sexual reproduction, consists one round of chromosome replication followed
by two rounds of chromosome segregation, called meiosis I and –II. The second meiotic division
presents several features that set it apart from all other forms of cell division. (1) Unlike mitosis, it is
neither preceded nor followed by DNA replication. Instead, it occurs directly after the M phase of
meiosis I and prior to a differentiation process that creates the gametes. (2) Meiosis II segregates
chromatids that have undergone recombination and are held together by cohesin located solely
around their centromeres. While meiosis II is frequently called a mitosis-like division, it remains
unclear to what extend principles of mitotic chromosome segregation and cell cycle control apply to
meiosis II. The analysis of meiosis II has been hindered by the difficulty of manipulating meiosis IIspecific processes without affecting meiosis I. To address this problem, we have developed a
protocol that generated meiotic cultures of budding yeast that progress through meiosis II in a highly
synchronous manner. We find that meiosis II includes unique mechanisms that coordinate the
removal of centromeric cohesin with exit from M phase and spore formation, the yeast equivalent of
gametogenesis.
40
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 4
Regulation of the G1-to-S transcriptional wave in an
unperturbed cell cycle
Isabel Alves-Rodrigues, Angel Guerra-Moreno, Maribel Marquina, Elena Hidalgo, Jose
Ayte
Oxidative Stress and Cell Cycle, Universitat Pompeu Fabra, Barcelona, 08003, Spain.
Inactivation of the Retinoblastoma protein (pRB) leads to unregulated cell cycle progression
promoting uncontrolled cell growth, genomic instability and aneuploidy, hallmarks of tumor
progression. pRB tumor suppressor activity is achieved through binding and regulating the E2F family
of transcription factors. The fission yeast MBF complex, which is the functional homolog of
mammalian E2F/pRB, drives the G1-to-S wave of transcription controlling the expression of genes
that are directly or indirectly required for DNA synthesis. MBF has been found bound to its target
promoters throughout the cell cycle, implicating that regulation of MBF activity is not achieved by
simply modulating its DNA-binding activity. We have recently shown that the MBF complex is
regulated by both the DNA replication checkpoint and the DNA damage checkpoint, but in opposite
directions: while the former activates MBF-dependent transcription, the latter down-regulates MBF
activity by directly phosphorylating the core subunit of the MBF complex, Cdc10 (Gomez-Escoda et
al, 2011; Ivanova et al 2013). To characterize the regulation of MBF in an unperturbed cell cycle, we
have developed a fluorescence-based reporter strain that can measure MBF activity in vivo. Using
reverse genetics to introduce this reporter system in a fission yeast knock-out (KO) collection that
includes over 3000 non-essential mutants, we have performed a double High Throughput Screening
(HTS) on live cells by i) FACS on 96-well plates and ii) Cell Sorting of individual mutants from a pooled
population of the barcoded KO strains. With this double screening, we have been able to identify
individual regulators of MBF. Among the isolated genes, we will discuss the roles of the
COP9/Signalosome and the histone deacetylase (HDAC) Hst4 in the regulation of MBF activity.
Gómez-Escoda, B., Ivanova, T., Calvo, I.A., Alves-Rodrigues, I., Hidalgo, E. and Ayté, J. (2011) Yox1
links MBF-dependent transcription to completion of DNA synthesis. EMBO Reports 12:84-89.
Ivanova, T., Alves-Rodrigues, I., Gómez-Escoda, B., Dutta, C., DeCaprio, J.A., Rhind, N., Hidalgo, E. and
Ayté, J. (2013) The DNA damage and the DNA replication checkpoints converge at the MBF
transcription factor. Mol. Biol. Cell 24:3350-3357
41
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 5
Releasing the brake - the functional role of phosphatases in
spindle assembly checkpoint silencing.
Debora Bade1, Vincent Groenewold1, 2, Antoinette Teixeira1, Geert Kops1, 2, 3
1
Molecular Cancer Research, University Medical Center Utrecht, 3584 CG, Utrecht, The Netherlands.
Department of Medical Oncology, University Medical Center Utrecht, 3584 CG, Utrecht, The Netherlands.
3 Cancer Genomics Netherlands, University Medical Center Utrecht, 3584 CG, Utrecht, The Netherlands.
2
The spindle assembly checkpoint (SAC) plays an important role as proof-reading machinery during
cell division: it delays anaphase onset until all chromosomes are attached in a bipolar manner and
consequently directly protects the cell from aneuploidy. Although it is fairly established how the SAC
is initiated and maintained, the actions leading to its silencing and subsequently to mitotic exit are
poorly understood. Specifically the fact that many proteins change their phosphorylation state upon
SAC silencing has not been addressed systematically yet.
In this project, I assess the differential phosphorylation sate of the SAC proteome of mitotic cells
either displaying unattached kinetochores or bi-oriented chromosomes. I thereby specifically
concentrate on kinetochore complexes such as the KMN, MCC, APC/C, RZZ, and SKA. Furthermore, I
am currently attributing the (de)phosphorylation event(s) to specific kinases, such as MPS1, or
phosphatases, such as PP2A and PP1. In detail, I am using APEX and BioID purification experiments
to confirm suspected and to identify new substrates of SAC phosphatases/kinases.
Next, I am addressing the functional contributions of selected phosporylation/dephosphorylation
events to SAC activation/silencing. In conclusion, this project aims to draw a comprehensive picture
of the SAC phosphoregulation.
42
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 6
A dynamical framework for the all-or-none G1/S transition in
individual human cells: a role for Emi1 in the irreversible
transition
Alexis Barr2, Tongli Zhang2, Chris Bakal1, Bela Novak2
1
2
Institute of Cancer Research
Department of Biochemistry, University of Oxford
Whilst the irreversible and all-or-none nature of the G1/S transition is well understood in yeast, we
lack an understanding of the systems-level properties that control this transition in mammalian cells.
Here we use quantitative single cell imaging to track the expression of key G1/S regulators in human
cells to parametrise a new model of the G1/S transition. We show that a proteolytic double negative
feedback loop between Cdk2:Cyclin and the Cdk inhibitor, p27Kip1, drives a switch-like S-phase
entry. Furthermore, our model predicts that Emi1 levels are critical in maintaining an irreversible
G1/S transition, which we validate using Emi1 knockdown and live imaging of G1/S reporters. Thus
we provide a dynamical framework to understand the irreversible G1/S transition in human cells.
43
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 7
A 14-3-3 phospho docking protein stabilises bipolarity of the
acentrosomal meiotic spindle
Robin Beaven, Hiro Ohkura
Wellcome Trust Centre for Cell Biology, The University of Edinburgh, UK
To faithfully segregate chromosomes, spindles need to maintain their bipolar organisation.
Centrosomes are key to this function during mitosis, but it is unclear how bipolar spindles are
formed and stably maintained in female meiosis, where centrosomes are absent. Using Drosophila
oocytes we identified a new function for 14-3-3epsilon in this context. The 14-3-3s are phospho-S/T
docking proteins which bind and regulate a multitude of interactors, so acting at the core of diverse
signalling pathways. We found that loss of 14-3-3epsilon caused female sterility and frequent
tripolar spindles. Live imaging showed that bipolar spindles could form but were highly unstable.
Mass spectrometry was then used to identify binding partners of 14-3-3epsilon in oocytes. They
included the kinesin Ncd, loss of which is also known to result in highly unstable meiotic spindles.
We had previously shown that the pole localisation of the microtubule regulator Msps (XMAP215) is
lost in ncd mutants, and depletion of Msps results in tripolar spindles similar to 14-3-3epsilon
depletion. We hypothesise that 14-3-3epsilon could regulate Ncd’s ability to transport Msps to the
spindle poles, and indeed find Msps localisation to be disrupted in 14-3-3epsilon’s absence. We have
also identified a predicted 14-3-3 binding site and neighbouring Aurora phosphorylation site in the
tail of Ncd, and hypothesise that Aurora kinases act to regulate 14-3-3’s binding to Ncd. These
molecular insights are shedding light on a mechanism which the meiotic spindle utilises to overcome
its inherent instability, resulting from its lack of centrosomes.
44
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 8
Annexin A2 is required for the early steps of cytokinesis
Christelle Benaud, Gaelle Le Dez, Svetlana Mironov, David Reboutier, Claude Prigent
Université de Rennes I, Institut de Génétique et Développement, Centre National de la Recherche Scientifique, UMR 6290,
Rennes, France
Cytokinesis requires the formation of an actomyosin contractile ring between the two sets of sister
chromatids. Annexin A2 is a calcium and phospholipid binding protein implicated in cortical actin
remodeling. We report that Annexin A2 accumulates at the equatorial cortex at the onset of
cytokinesis and depletion of Annexin A2 results in cytokinetic failure, due to a defective cleavage
furrow assembly. In absence of Annexin A2, the small GTPase RhoA that regulates cortical
cytoskeletal rearrangement fails to form a compact ring at the equatorial plane. Furthermore,
Annexin A2 is required for cortical localization of the RhoGEF Ect2 and to maintain the association
between the equatorial cortex and the central spindle. Our results demonstrate that Annexin A2 is
necessary in the early phase of cytokinesis. We propose that annexin A2 participates in the central
spindle-equatorial plasma membrane communication.
45
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 9
Cdk1-mediated regulation of the protein phosphatase scaffold
RepoMan
Junbin Qian, Jin Huang, Sofie De Munter, Bart Lesage, Monique Buellens, Mathieu
Bollen
Laboratory of Biosignaling & Therapeutics, Department of Cellular and Molecular Medicine, University of Leuven, Belgium
RepoMan is a mitotic interactor of protein phosphatases PP1 and PP2A-B56. During prometaphase
PP2A-B56 promotes the chromosome targeting of RepoMan, whereas RepoMan-associated PP1
opposes the centromeric targeting of Aurora B through dephosphorylation of Histone H3 at Thr3.
Here, we show that the Cdk1-mediated phosphorylation of RepoMan reduces the binding of PP1 but
enhances the recruitment of PP2A-B56. Upon inactivation of Cdk1 at the beginning of anaphase
PP2A-B56 dissociates from RepoMan and the binding of PP1 is increased, culminating in the bulk
recruitment of RepoMan to the chromosomes. This phosphatase switch serves to make the
chromosome-binding of RepoMan Cdk1-independent and irreversible. We also show that the
reduced PP1-RepoMan interaction during prometaphase prevents the precocious dephosphorylation
of Aurora B and Importin beta which are substrates of RepoMan-associated PP1 in anaphase. Hence,
the Cdk1-mediated phosphorylation of RepoMan contributes to the ordered dephosphorylation of
mitotic-exit substrates.
46
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 10
The Drosophila protein Matrimony is a functional homolog of
kinetochore proteins Spo13, Moa1, and Meikin
Amanda Bonner1, R. Scott Hawley1, 2
1
2
Stowers Institute for Medical Research
Department of Molecular and Integrative Physiology, University of Kansas Medical Center
During meiosis homologous chromosomes must first pair and then segregate from each other in a
reductional division before going through a second equational, mitosis-like division in which sisters
separate to create haploid gametes. Proper segregation of homologs during the first meiotic division
depends on both the formation of chiasmata between homologs and the mono-orientation of sister
kinetochores towards the same pole. Meiosis-specific kinetochore proteins have been characterized
in budding and fission yeasts, and, very recently, in mammals, namely Spo13, Moa1, and Meikin,
respectively. While these proteins share no sequence homology, all three are expressed specifically
during meiosis I, co-precipitate with Polo-like kinases, and aid in the protection of cohesion between
sister chromatids at the centromere. Here we provide evidence that the Drosophila protein
Matrimony (Mtrm) plays a similar role during female meiosis. Mtrm, which was originally identified
genetically as a gene required for the proper segregation of achiasmate chromosomes, has also been
shown to be a strong interactor of Polo kinase. Also, despite the overall lack in sequence homology
among these four proteins, Mtrm contains an LxExxxN degron motif that was originally shown in
Spo13 to be required for both proteins to be recognized by the Anaphase Promoting Complex (APC).
There is also a shared region of homology between Mtrm and Meikin that contains a canonical Polobox binding motif. We demonstrate here that, as is the case for mutants in Spo13, Moa1, and
Meikin, mutations in Mtrm cause precocious separation of sister chromatids and chromosome
missegregation defects, particularly when recombination between homologs is absent. These
findings help explain the curious segregational defects we see in a mtrm-/+ heterozygote, where
homologs that are connected by a chiasmata disjoin as expected, while achiasmate homologs
seemingly segregate at random. We also show that Mtrm can inhibit cell cycle progression when
expressed during mitosis, as has also been previously shown with Spo13. On the basis of these data,
we suggest that Mtrm joins a very newly described family of meiosis-specific kinetochore proteins
that are important for regulating Polo-like kinase’s activities during the first meiotic division.
47
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 11
Degradation of Ndd1 by APC/CCdh1 generates a feed forward
loop that times mitotic protein accumulation
Michael Brandeis1, Julia Sajman1, Drora Zenvirth1, Mor Nitzan2
1
The Department of Genetics, The Alexander Silberman Institute of Life Sciences,The Hebrew University of Jerusalem,
Jerusalem 9190401, Israel
2 The Racah Institute of Physics, The Hebrew University of Jerusalem, Jerusalem 9190401, Israel
Ndd1 activates the Mcm1–Fkh2 transcription factor to transcribe mitotic regulators. The anaphasepromoting complex/cyclosome activated by Cdh1 (APC/C-Cdh1) mediates the degradation of
proteins throughout G1. Here we show that the APC/C-Cdh1 ubiquitinates Ndd1 and mediates its
degradation, and that APC/C-Cdh1 activity suppresses accumulation of Ndd1 targets. We confirm
putative Ndd1 targets and identify novel ones, many of them APC/C-Cdh1 substrates. The APC/CCdh1 thus regulates these proteins in a dual manner—both pretranscriptionally and posttranslationally, forming a multi-layered feedforward loop (FFL). We predict by mathematical
modelling and verify experimentally that this FFL introduces a lag between APC/C-Cdh1 inactivation
at the end of G1 and accumulation of genes transcribed by Ndd1 in G2. This regulation generates
two classes of APC/C-Cdh1 substrates, early ones that accumulate in S and late ones that accumulate
in G2. Our results show how the dual state APC/C-Cdh1 activity is converted into multiple outputs by
interactions between its substrates.
48
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 12
The double role of Cdc20 in the Spindle Assembly Checkpoint
Andrea Ciliberto1, Elena Chiroli1, Paolo Bonaiuti1, Fridolin Gross2
1
2
IFOM, Via Adamello 16, Milan
Institute for Theoretical Biology, Charite and Humboldt University, Invalidenstrasse 43, Berlin
The activation of the Anaphase Promoting Complex or Cyclosome (APC/C) drives cells from
metaphase to anaphase. APC/C ubiquitinates two key substrates – mitotic cyclins and securin –
whose degradation allows the progression of the cell cycle oscillator and the separation of sister
chromatids. To recognize its substrates, APC/C needs to be bound to its cofactor Cdc20.
Cdc20, however, is not simply an activator of APC/C, as it can also inhibit APC/C. More precisely, in
the presence of unattached kinetochores Cdc20 is part of a multi-protein complex, the Mitotic
Checkpoint Complex or MCC, which inhibits APC/C and arrests cells in metaphase. MCC is the final
output of a signaling pathway known as the Spindle Assembly Checkpoint which prevents cells from
becoming aneuploid.
The double effect of Cdc20 on APC/C prompted opposite interpretations regarding its role during a
mitotic arrest. Cdc20 degradation has been proposed to be required for timely APC/C activation (i.e.,
Cdc20 as an inhibitor) but also for proper checkpoint maintenance (i.e., Cdc20 as an activator). Here,
we use a combination of mathematical models and experiments to show that, during an arrest
induced by the SAC, Cdc20 is primarily an inhibitor of APC/C. It can only switch to an activator when
its synthesis becomes uncoupled from degradation.
49
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 13
Mitotic catastrophe: insights from synthetic yeast
Claude Gérard1, John Tyson2, Damien Coudreuse3, Bela Novak1
1
Oxford Centre for Integrative Systems Biology, University of Oxford, Oxford, UK
Department of Biological Sciences, Virginia Tech, Blacksburg, USA
3 Institute of Genetics and Development, CNRS UMR 6290, Rennes, France
2
Proper progression through the cell cycle is fundamental for cellular life. From the control of mitotic
entry to the regulation of the onset of DNA replication, the cyclin-dependent kinase (CDK) family is
the primary node of the circuit driving the eukaryotic cell division cycle. Modulation of CDK activity
relies on a host of inputs, highlighting the complexity of this critical process. However, discerning
essential controls from secondary regulation is a challenge that may limit our understanding of the
core engine behind cell cycle progression. Building on past studies demonstrating that a single cyclin
is sufficient for driving this process in fission yeast, we generate and analyze basic synthetic systems
providing controlled CDK/Cdc2 activity levels and bypassing a large part of the endogenous
regulatory network. Interestingly, the simplicity of the synthetic yeasts we have built makes them
particularly well-adapted for mathematical modeling and theoretical dissection of the organization
of cell cycle control. A combination of modeling and experimental approaches using fission yeast
cells operating with minimal cell cycle networks indeed allowed us to propose a novel mechanism
for the origin of mitotic catastrophe, a lethal result of major deregulation and overlap of cell cycle
phases. Our studies reveal a cumulative effect of deregulated G1/S cyclin-dependent CDK activity on
mitotic control in the absence of the Wee1/Cdc25 feedback loop. This work highlights the
importance of coupling classical genetics with synthetic biology and mathematical modeling for
understanding normal and pathological cell cycle events.
50
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 14
Single-cell monitoring of cell cycle progression in whole
developing organs.
Bénédicte Desvoyes, María Delgado-Barea, Sofía Otero, María Isabel Lopez, Crisanto
Gutierrez
Centro de Biología Molecular Severo Ochoa (CSIC-UAM), calle Nicolas Cabrera 1, 28049 Madrid, Spain.
Plant organ development is mainly a postembryonic process that occurs in the adult in a continuous
manner. Consequently, a strict balance between cell proliferation and differentiation is required. In
addition, many plant cells undergo one or more endoreplication cycles as part of their normal
differentiation program. Therefore, growing plant organs consist of populations of dividing,
endoreplicating and differentiated cells all derived from a few stem cells. The transitions between
the different cell pools are integrated and respond to developmental and environmental cues. In
fact, cell cycle control influences greatly developmental programs, e.g. increased or decreased
proliferation rates affect organ growth and shape.
To get insight into the coordination of the different processes that influence organ development it is
important to be able to identify cells progressing in vivo through the cell cycle in a non-invasive
manner. We have used fluorescently-labeled proteins that unambiguously identify cells in G1, S/G2
and G2 and generated plants that express them under their own promoters in a single plant. (1) As a
G1 marker, we used a CFP-labeled prereplication complex protein that is loaded in late M/early G1
and rapidly degraded soon after S-phase initiation in a proteasome-dependent manner. (2) S-phase
cells are followed by the incorporation of a canonical histone H3.1-mRFP, which is maintained
through mitosis in actively dividing cells and excluded late in G2 in cells undergoing their last mitotic
cycle. (3) Finally, G2 cells are labeled with CYCB1;1-GFP, with a maximum in late G2 and degraded by
the APC in anaphase. These markers are also useful to follow cells undergoing endoreplication
cycles. The combination of these three markers in a single organism allows monitoring all the cell
cycle phases and constitutes an important tool to study cell cycle regulation in the context of a
growing organism during normal growth or in response to internal and external signals.
51
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 15
Modeling the dynamics of spindle assembly checkpoint
signaling during mitotic exit progression
Amalie Dick1, Lukas Hutter2, Béla Novák2, Daniel Gerlich1
1
2
Institute of Molecular Biotechnology of the Austrian Academy of Sciences (IMBA), Vienna, Austria
Oxford Centre for Integrative Systems Biology, Department of Biochemistry, University of Oxford, Oxford, UK
The spindle assembly checkpoint contributes to faithful chromosome segregation by delaying
anaphase onset until all chromosomes have attached their sister kinetochores to opposing spindle
poles. Satisfaction of the spindle assembly checkpoint leads to rapid APC/C activation, which
promotes mitotic exit by targeting cyclin B1 for ubiquitin/proteasome-mediated degradation. Cdk1
inactivation by cyclin B1 degradation in turn contributes to permanent spindle assembly checkpoint
silencing, which has been proposed as a mechanism to warrant unidirectional progression through
mitotic exit.
Here, we performed quantitative live cell imaging and mathematical modeling to better understand
the kinetics of APC/C activation and the regulation of spindle assembly checkpoint signaling at the
metaphase-to-anaphase transition. By acutely detaching chromosomes from the spindle using laser
microsurgery or nocodazole, we found that the spindle assembly checkpoint remained proficient
until the very end of metaphase, whereas it was unresponsive already during very early stages of
anaphase. To investigate how the spindle assembly checkpoint rapidly switches from a responsive to
a non-responsive state, we established live cell assays probing for the effect of partial Cdk1
inactivation.
The quantitative data were integrated into a mathematical model that describes the kinetics of the
regulatory system in terms of its key components. Our model recapitulates key features of spindle
assembly checkpoint signaling from prometaphase until mitotic exit. This helps us to understand
how the steady states of the system determine its response to lack of attachment. Our analysis
shows how thresholds for inactivation and re-activation of spindle assembly checkpoint signaling
arise at the systems-level, and establish a “point of no return” at the metaphase-to-anaphase
transition as Cdk1-activity drops.
52
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 16
Roberts Syndrome Esco2-E450del Mutant Partially Rescues
Cohesion at Pericentric Regions of Esco2 Deficient MEFs.
Peter Ditte, Gregor Eichele
Max Planck Institute for Biophysical Chemistry, Göttingen, Germany
Although most of known ESCO2 mutations in Roberts Syndrome (RBS) lead to nonsense-mediated
decay or to severely truncated proteins, there are three mutations located in acetyltransferase
domain that stand out because they do lead to a protein of normal size. Among them p.E453del
corresponds to in-frame deletion of glutamic acid at position 453 and despite the fact that it causes
RBS, the protein possesses full auto-acetylation activity. No cytogenetic or in vivo biochemical data is
available since only one case of stillborn fetus with E453del mutation was reported.
Using BAC transgenomics we prepared MEF cell line expressing murine Esco2-E450del in Esco2
deficient background. The expression profile of Esco2-E450del is similar to wild type protein with
peak of expression in mid S-phase. The level of Smc3 acetylation in Esco2-E450del MEFs is decreased
during S-phase compared to wild type Esco2 and comparable to Esco2 null cells. The data shows that
Esco2-E450del is unable to effectively acetylate Smc3 in vivo. As a consequence of impaired sister
chromatid cohesion at pericentric regions, mitotic chromosomes of Esco2 deficient MEFs typically
show railroad track appearance, what can be evaluated by distance between centromeres. In Esco2E450del MEFs, the average distance between centromeres is clearly shorter than in Esco2 null cells
albeit still significantly longer than in Wt cells, suggesting that Esco2-E450del MEFs partially establish
sister chromatid cohesion at centromeres.
Altogether, our data provide the first biochemical evidence that E450del mutation has severe impact
on ability of Esco2 to acetylate Smc3 in vivo, yet showing milder cohesion defects at centromeric
regions of Esco2 deficient MEFs. Furthermore, the case of Esco2-E450del indicates that at least some
RBS mutations may not be null.
53
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 17
Rab35 GTPase couples lumen formation to cell division during
polarity establishment in 3D renal cysts
Kerstin Klinkert, Arnaud Echard
Membrane Traffic and Cell Division Lab
Institut Pasteur - CNRS UMR3691
25-28 rue du Docteur Roux, 75015, Paris, France
Most of the human cancers arise from epithelial cells and the loss of epithelial apico-basal polarity is
a hallmark in tumor progression. It has been shown that the initiation of epithelial polarity is tightly
connected to cell division and polarized membrane traffic, but how this is coordinated at a molecular
level remains unknown. We addressed this question in renal MDCK cells, which develop 3D polarized
cysts with an open apical lumen from a single cell when cultured in Matrigel. Unpolarized renal
MDCK cells establish an apical membrane already at the first cell-cell interface during the two-cell
stage, and this is coupled to cytokinesis by a unknown mechanism. Unexpectedly, we found that the
Rab35 GTPase directly interacts with the cytoplasmic tail of the apical transmembrane protein
Podocalyxin (PODXL, aka GP135), a classical apical marker essential for epithelial polarity and lumen
formation. Interestingly, Rab35 was enriched at the future apical membrane during the first cell
division before PODXL or any other tested apical proteins could be detected. Depletion of Rab35 or
replacement of endogenous PODXL by a mutant unable to interact with Rab35, prevented PODXL
and other apical proteins to become enriched at the future apical membrane during the first cell
division. Consequently, a complete inversion of apico-basal polarity was observed in these
conditions. Furthermore, the experimental delocalization of Rab35 to mitochondrial membranes
induced a striking accumulation of vesicles containing PODXL, Crb3, Cdc42 and aPKC around the
mitochondria, leading to a loss of apico-basal polarity. We propose that Rab35 acts as a direct tether
for transcytosed PODXL-containing vesicles at the future apical membrane, and thereby plays a
crucial role in triggering the establishment of apical polarity. In addition, this work provides a
molecular mechanism for coupling the site of cytokinesis to the initiation of polarity and lumen
localization in 3D structures.
54
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 18
Kinase-Phosphatase interplay regulates Kar9 function in spindle
positioning
Ana-Maria Farcas, Mathias Bayer, Yves Barral
Institute of Biochemistry, ETH Zurich
Faithful alignment of the mitotic spindle with the polarity axis of the cell drives asymmetric division
of polarized cells. In constitutively asymmetric dividing cells, such as budding yeast, this process is
also essential in order to ensure that each daughter inherits a full copy of the genome. In
Saccharomyces cerevisiae the protein Kar9 mediates the alignment of the spindle with the motherbud axis of the metaphase cell. Kar9 localizes to aster microtubules in an asymmetric fashion, almost
exclusively to the plus-end of microtubules emanating from the old spindle pole body (SPB, the yeast
equivalent of the centrosome). Interaction of Kar9 with microtubule plus-ends depends on its
interaction with the EB1 homolog Bim1. Furthermore, Kar9 interacts with the type V myosin Myo2,
which walks along actin cables emanating from the bud cortex, carrying Kar9 and microtubule tips
towards the bud. Thereby, the Bim1-Kar9-Myo2 pathway drives both the orientation and alignment
of the spindle in mitotic yeast cells. Accordingly, symmetric localization of Kar9 to both asters causes
both SPBs to orient towards the bud and impairs spindle alignment with the mother-bud axis.
Therefore, the mechanisms ensuring Kar9 asymmetry are crucial for proper spindle positioning.
Here, we show that the protein phosphatase 1 (Glc7 in budding yeast), interacts with, and controls
Kar9 function, most likely by counteracting Clb4/Cdk1- and Dbf2-mediated phosphorylation of Kar9.
By using a series of Kar9 mutants (e.g. phospho-mimicking, phospho-ablating, phosphatase-docking
and chimeras), we aim to understand the mechanism by which Clb4/Cdc28, Dbf2 and Glc7 regulate
Kar9 distribution and function.
55
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 19
The phosphatase regulator NIPP1 is essential for the
proliferation and differentiation of male germ cells
Monica Ferreira1, 2, Shannah Boens1, Kathelijne Szeker1, Aleyde Van Eynde1,
Margarida Fardilha2, Mathieu Bollen1
1
Laboratory of Biosignaling & Therapeutics, Department of Cellular and Molecular Medicine, University of Leuven, Leuven,
Belgium.
2 Institute for Research in Biomedicine-iBiMED, Health Sciences Department, University of Aveiro, Aveiro, Portugal.
NIPP1, for Nuclear Inhibitor of Protein Phosphatase 1 (PP1), regulates transcription and pre-mRNA
splicing by targeting PP1 to specific nuclear substrates. The deletion of NIPP1 in mice is early
embryonic lethal. Studies on cultured cells suggest a role for NIPP1 in cell proliferation and
differentiation. To examine the role of NIPP1 in adult germline cell proliferation and differentiation,
we generated an inducible NIPP1 knockout (KO) mouse model. We observed that the Crerecombinase induced deletion of NIPP1 in four-weeks-old testis causes a progressive loss of germ
cells within 8 weeks, culminating in a Sertoli cell-only phenotype. At earlier time points, the mitotic
germ cells (spermatogonia) failed to progress beyond early spermatogenic events due to
proliferation defects and cell cycle dysregulation, resulting in an increased rate of apoptosis. Several
recombination-related proteins were downregulated after the deletion of NIPP1 and the incidence
of unrepaired DNA breaks was decreased, as detected by reduced γH2AX foci on the axes of
autosomal chromosomes in spermatocytes during meiotic prophase I. Consistent with this finding,
radiation-induced γH2AX foci formation was compromised in the NIPP1 KO testis. Altogether our
results show that NIPP1 is essential for both male germline stem-cell proliferation and the
differentiation of spermatocytes.
56
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 20
How budding yeast control extracellular matrix remodelling
during cytokinesis?
Magdalena Foltman1, 2, Iago Molist1, 2, Alberto Sanchez-Diaz1, 2
1
Instituto de Biomedicina y Biotecnología de Cantabria, Universidad de Cantabria, CSIC, Albert Einstein 22, 39011,
Santander, Spain
2 Departamento de Biología Molecular, Universidad de Cantabria, Facultad de Medicina, Cardenal Herrera Oria s/n, 39011,
Santander, Spain
During cytokinesis cells coordinate contraction of the actomyosin ring with the ingression of the
plasma membrane and remodelling of the extracellular matrix (ECM) but the underlying mechanisms
are still poorly understood. In eukaryotes, glycosyltransferases that synthesise ECM polysaccharides
are emerging as important players during cytokinesis. In budding yeast the chitin synthase Chs2
makes the primary septum, a special layer of ECM that is essential for cell division.
To try to understand how yeast cells coordinate actomyosin ring contraction, plasma membrane
ingression and remodelling of the extracellular matrix we have used budding yeast Chs2 and Inn1
proteins to isolate ‘ingression progression complexes’ (IPCs) that contain key actomyosin rings
components from yeast cells undergoing cytokinesis synchronously. We have identified, together
with Chs2 and Inn1, actomyosin ring components Cyk3, myosin type II, the IQGAP protein Iqg1 and
Hof1. We propose that IPCs are central to the mechanism by which cells coordinate cytokinesis. We
have found that the catalytic domain of Chs2 interacts directly with the C2 domain of Inn1 and the
transglutaminase-like domain of Cyk3. We have now data indicating that Inn1, Chs2 and Cyk3 form a
stable complex. We found that chitin synthase Chs2 is activated by C2 domain of Inn1, as well as the
transglutaminase-like domain of Cyk3. Finally we have exploited an experimental system that
allowed us to determine a previously unknown role for the C-terminus of Inn1 and to understand
how Inn1 and Cyk3 finely regulate Chs2 activity in vivo.
57
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 21
Aurora A mutant neuroblasts are defective for cyclin B
degradation in the absence of the Spindle Assembly
Checkpoint.
Régis GIET
Université of Rennes 1-CNRS
Control of the balance between stem cell renewal, differentiation and accurate chromosome
segregation are essential to ensure tissue homeostasis. Drosophila mutants for sas-4 and aurA
display brain tumors with extra Neuroblasts (NBs), triggered by loss of cell identity. These mutants
also show defective mitotic spindle assembly and consequent mitotic delay caused by the activation
of the Spindle Assembly Checkpoint (SAC).
In an attempt to determine if generating aneuploidy would compromise NB proliferation, we
inactivated the SAC by removing the SAC protein Mad2 from aurA and sas-4 mutants. Removal of
the SAC in the sas-4 mutant impaired NB proliferation and chromosome segregation. By contrast,
removing the SAC in the aurA mutant did not prevent brain overgrowth, tumor formation, and
chromosome segregation. Monitoring Mad2 and cyclin B dynamics in live aurA neuroblasts revealed
that chromosome alignment and SAC satisfaction were not coupled to timed cyclin B degradation.
We therefore show for the first time the existence of an Aurora A-dependent mechanism that
promotes timed and efficient cyclin B degradation.
As a consequence, aurA mutants NBs are delayed in mitosis, and perform accurate chromosome
segregation even in the absence of the spindle assembly checkpoint.
58
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 22
The kinetochore-to-cytoplasm ratio determines the strength of
the spindle assembly checkpoint during early embryonic
divisions of C. elegans
Matilde Galli, David Morgan
UCSF, Department of Physiology
The spindle assembly checkpoint (SAC) delays mitotic progression when chromosomes are not
properly attached to microtubules of the mitotic spindle. Interestingly, cells exhibit a large variation
in the extent to which they delay mitotic progression upon activation of the SAC. In C. elegans,
previous studies have shown that early embryonic divisions are only moderately delayed upon
disruption of their mitotic spindle. To determine whether this is due to an intrinsically weak
checkpoint in C. elegans, or whether the SAC becomes stronger during later developmental stages,
we systematically assayed the mitotic delay induced by microtubule disruption at different stages of
development. To disrupt microtubules during larval stages, when cells are impermeable to
microtubule drugs, we inducibly expressed DarpinD1, a designed ankyrin repeat protein selected to
bind tubulin and promote microtubule depolymerization. Interestingly, in contrast to the weak SAC
in early embryos, activation of the SAC during larval intestinal divisions delays mitosis for several
hours. To understand when this strong SAC arrest arises during development, we measured the SAC
at different stages of embryonic development. Strikingly, we found a gradual increase in the arrest
time, ranging from the reported 2-2.5-fold delay in 2-cell embryos to a 10-fold delay at the 50-cell
stage. We hypothesized that the gradual increase in strength of the SAC could be explained by a
decrease in cellular size in older embryos. Specifically, the strength of the SAC could depend on the
ratio between the amount of unattached kinetochores, where the checkpoint signal is generated,
and the amount of cytoplasm. To test whether cell size, and not a developmental timer, determines
the strength of the checkpoint, we depleted ani-2 to induce a spread in embryo sizes and then
assayed the mitotic delay in differently-sized two-cell embryos. Here again, smaller cells displayed a
longer mitotic delay, arguing against a developmental timer controlling the strength of the
checkpoint. Furthermore, we find that triploid embryos, which have an increased amount of
kinetochores, exhibit a stronger mitotic delay upon disruption of the mitotic spindle. Together, these
results demonstrate that the kinetochore to cytoplasm ratio determines the extent to which
embryonic cells delay mitosis upon activation of the SAC. Our work provides new insights into why
cells exhibit large variations in their ability to delay mitotic progression upon disruption of the
mitotic spindle.
59
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 23
PP2ACdc55 opposes Cdk substrate phosphorylation in budding
yeast interphase
Molly Godfrey, Frank Uhlmann
Chromosome Segregation Lab, Francis Crick Institute Lincoln's Inn Fields Laboratory
In the quantitative model for the cell cycle, rising levels of Cyclin-dependent-kinase (Cdk) activity
determine the timing of cell cycle events and progression through the different cell cycle stages.
Recently, the importance the importance of Cdk-opposing phosphatase activity in this quantitative
cell cycle model has come to light. This is particularly true when it comes to mitotic exit in S.
cerevisiae, where the ratio of Cdk/Cdc14 phosphatase activity determines the orderly progression
through mitotic exit in a quantitative manner.
Similarly, we postulate that Cdk-opposing phosphatase activity in interphase could be an important
element in this quantitative model for cell cycle progression. This activity could be required for
ordered phosphorylation of Cdk phosphoproteins during interphase and preventing too early cell
cycle transitions. Evidence from Xenopus as well as yeast suggests that PP2ACdc55 is a good
candidate for such a phosphatase. We have shown that in the absence of PP2ACdc55, global
interphase Cdk phosphorylation levels are elevated, independently of PP2A’s role in regulating Cdk
tyrosine phosphorylation. We have also identified specific Cdk targets whose phosphorylation is
advanced and increased in this case. This indicates that PP2ACdc55 may be directly opposing Cdk
phosphorylation events throughout interphase, from G1 until the onset of mitosis. PP2ACdc55 might
be part of a mechanism that sets Cdk thresholds for cell cycle progression.
60
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 24
Pausing the cell cycle at the mid-blastula transition in
Drosophila embryos
Jörg Großhans, Boyang Liu, Hung-wei Sung, Franziska Winkler
Institute for Developmental Biochemistry, Medical School, University of Göttingen
During early embryonic development in Drosophila, the mode of the cell cycle switches from fast
nuclear cycles to a mode with an extended G2 phase. This switch in cell cycle mode is an essential
part of a developmental switch, namely the mid-blastula transition. Concommitantly with the G2
pause, zygotic gene expression is activated, maternal mRNA are degraded and cellularisation starts.
To reveal the functional relationship between these processes and to determine the molecular
mechanism of the robust cell cycle pause, we isolated and characterised mutants with more or
fewer cell cycles prior to the switch. We previously reported that premature activation of zygotic
transcription is sufficient to trigger the complete developmental switch. By analysing a novel
mutation in the protein phosphatase V (PP-V) which leads to an additional nuclear division, we find
that PP-V required for timely pause of the cell cycle. Furthermore, a novel mutation in the the gene
encoding the enyzme serine hydroxymethyl transferase, leads to a precocious pause after 12 instead
of 13 nuclear divisions. We will present our unpublished analyses and findings how PP-V and SHMT
are involved in cell cycle control during MBT.
61
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 25
Identification of novel regulators of mitotic chromosome
architecture
Christoph Schiklenk, Jin Wang, Carlo Klein, Christian Haering
Cell Biology and Biophysics Unit, EMBL Heidelberg
Chromosome condensation is an essential prerequisite for successful chromosome segregation
during mitosis and meiosis. Even though some proteins that are essential for the formation of
mitotic chromosomes, including condensin complexes and topoisomerase II, have been recently
identified, the complete repertoire of molecular machines that organise mitotic chromosomes has
remained elusive.
Using a newly developed high-throughput assay that quantitatively measures chromosome
condensation dynamics in live fission yeast cells, we screened for yet undiscovered factors involved
in chromosome condensation. I will present novel candidate proteins that we identified in this
screen as regulators of mitotic chromatin architecture. One of these proteins encodes a large zincfinger protein named Zas1. Zas1 is essential for cell division and localizes to chromosomes during
interphase and during cell divisions, consistent with a direct role in regulating chromosome
architecture. I will discuss in detail the function of Zas1 and its role in mitotic chromosome
condensation.
62
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 26
Exploring the recruitment and regulation of Mad1 during
mitosis
Daniel Hayward, Ulrike Gruneberg
Sir William Dunn School of Pathology, University of Oxford
The spindle assembly checkpoint (SAC) ensures accurate chromosome segregation by delaying
mitotic progression until all kinetochores are attached to microtubules, therefore avoiding
aneuploidy which can lead to cell death or cancer. This is achieved by kinetochores unattached to
microtubules generating the mitotic checkpoint complex (MCC), which in turn inhibits the anaphasepromoting complex (APC/C) and maintains sister chromatid cohesion.
Mad1 (mitotic arrest deficient 1) is an essential SAC protein, recruiting Mad2 which is an
indispensable member of the MCC. Both Mad1 and Mad2 are the first SAC proteins lost at the
kinetochore upon microtubule-kinetochore attachment. Depletion of Mad1 in cells results in a
defective mitotic checkpoint whilst artificial tethering of Mad1 to the kinetochore results in mitotic
arrested cells with the SAC permanently on.
Although the importance of Mad1 has been extensively demonstrated, the nature of its regulation
and to what it binds at the kinetochore are poorly understood.
Here we explore how Mad1 binds to the kinetochore in Human cells and discover how mitotic
kinases and phosphatases effect its localisation. We expand upon previous work and show that both
MPS1 and Aurora B kinases are required for initial Mad1 localisation to kinetochores. Both kinases
are also required for maintenance of Mad1 at kinetochores in the presence of microtubules, but are
dispensable when microtubules are depolymerised. We further explore the action of these kinases
and their substrates. In addition to previous work suggesting the importance of Bub1 and the RZZ
complex in Mad1 localisation we also identify the Ndc80 complex as vital in establishing Mad1 at the
kinetochore. We can now present a clearer model of Mad1 regulation at the kinetochore.
63
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 27
Protein phosphatase 1 is required for inactivation of Greatwall
during mitotic exit in Xenopus embryos
Andreas Heim1, 2, Anja Konietzny1, Thomas U. Mayer1, 2
1
2
Department of Molecular Genetics, University of Konstanz, Universitätsstraße 10, 78457, Konstanz, Germany
Konstanz Research School Chemical Biology, University of Konstanz, Universitätsstraße 10, 78457, Konstanz, Germany
Entry into and exit from M-phase are triggered by changes in the phosphorylation state of multiple
cell cycle regulators. In Xenopus laevis oocytes and embryos, the M-phase promoting activity of
cyclin-dependent kinase 1 (Cdk1)/cyclin-B is actively counteracted by protein phosphatase 2A in
complex with a regulatory B55 subunit (PP2A-B55). Thus, to facilitate faithful and irreversible entry
into M-phase, Cdk1/cyclin-B and PP2A-B55 must be regulated in a manner such that their activities
are mutually exclusive. At mitotic entry, Cdk1/cyclin-B-activated Greatwall (Gwl) kinase
phosphorylates the small heat stable proteins Arpp19/Ensa which enables them to tightly bind to
and inhibit PP2A-B55. While it is well established that both Gwl and Arpp19/Ensa have to be
dephosphorylated during mitotic exit to facilitate PP2A-B55 re-activation, the precise underlying
mechanism remains elusive. Here, we demonstrate that protein phosphatase 1 (PP1) is essential for
Gwl inactivation at mitotic exit. PP1 dephosphorylates Gwl at autophosphorylation sites resulting in
Gwl inactivation. Subsequently, PP2A-B55 can completely dephosphorylate Gwl to reset it to the
interphasic state, which is important to prevent untimely Gwl reactivation. Thus, our work identifies
PP1 as the sought-after phosphatase that breaks the Gwl-Arpp19/Ensa module of PP2A-B55
inhibition and thereby provides a molecular explanation for the essential function of PP1 during exit
from mitosis in Xenopus early embryos.
64
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 28
Novel stage-specific fluorescent indicators for mapping
Ca2+ signals during mitosis in mammalian cells
Nordine Helassa1, 2, Lee Haynes2, Robert D Burgoyne2, Katalin Torok1
1
2
St George’s University of London, Institute of Cardiovascular and Cell Sciences, London, UK
University of Liverpool, Institute of Translational Medicine, Department of Cellular and Molecular Physiology, Liverpool, UK
The division of cells by mitosis is crucial for normal development, growth and tissue renewal and its
tight regulation is essential to prevent abnormal cell proliferation. Understanding the mechanistic
details and regulation of mitosis is important to mammalian cell biology since irregularities or failure
of this process can lead to aneuploidy, genetic instability, premature aging and malignant potential.
Ca2+ signalling is fundamental to multiple key physiological processes ranging from fertilisation to cell
death. In spite of the evidence indicating a role for Ca2+ signalling during progression of the cell cycle
and, in particular, during mitosis, there have been conflicting reports that have hindered progress in
this field. Our hypothesis is that Ca2+ signals are important for critical stages of cell division in
mammalian cells but that such events are spatio-temporally restricted to such an extent as to have
evaded detection in previous studies employing conventional cytosolic Ca2+ dyes. In order to address
these issues experimentally we are taking a novel approach of using targeted, genetically encoded,
calcium probes (GCaMPs) to monitor Ca2+ signalling during mitosis.
GCaMPs are based on a circularly permutated EGFP molecule (cpEGFP) flanked at the N and C
termini by the smooth muscle myosin light chain kinase derived RS20 peptide and calmodulin (CaM),
respectively. Upon Ca2+ binding, the formation of a tight complex between RS20 and CaM, stabilises
the deprotonated form of cpEGFP inducing a fluorescence enhancement. However, the high affinity
and slow Ca2+-response kinetics of the current GCaMPs does not make them suitable for measuring
localised, short lived mitotic Ca2+ transients.
To optimize GCaMP6 for mapping mitotic Ca2+ signals, we decreased the binding affinity of the
Ca2+.CaM.RS20 complex and accelerated the Ca2+-response kinetics by point mutations using
rationale design. Newly engineered GCaMP6 proteins were characterised in terms of dynamic range,
Ca2+ affinity and association and dissociation kinetics. Dissociation constants (Kd) for Ca2+ obtained
from the equilibrium Ca2+ binding experiments ranged from 0.1 to 2.6 microM with Fmax/Fmin of ~ 20,
making these variants suitable for tracking Ca2+ influx in multiple scenarios. In stopped-flow kinetics
experiments, we showed that the variant GCaMP6f RS1 EF4 had half times (t1/2) for Ca2+ rise and
decay 5-fold and 2-fold faster than GCaMP6f at 37°C, respectively (t1/2for rise of 3.3 ms and t1/2 for
decay 15.7 ms). The improved GCaMP6 variants were used to generate a library of mitosis stagespecific Ca2+ sensors by fusing to proteins relevant for mitosis that have specific localisations during
the process (e.g. RhoA, Arf6, pericentrin, actin, tubulin, lamp1). We demonstrated by spinning-disk
confocal microscopy that the targeted Ca2+ probes express and localize properly which provide
useful tools to accurately map Ca2+-signalling activity with high temporal and spatial resolution
throughout mitosis.
Funded by the Wellcome Trust (094385/Z/10/Z) to KT and Leverhulme Trust (RPG-2014-194) to LH
and RB.
65
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 29
Possible novel mechanism of degradation for the human new
cyclin I
Sara Hernández-Ortega, Samuel Bru, Natalia Ricco, Javier Jiménez, Josep Clotet
Grup de Noves Ciclines, Departament de Ciències Bàsiques, Universitat Internacional de Catalunya, Josep Trueta s/n, 08195,
Sant Cugat del Vallès.
Cell cycle is controlled by CDKs along with their correspondent partner cyclins in each phase of the
cycle. Some years ago, our group described in S. cerevisiae that Pcl1, a not essential cyclin, have
special relevance when the environmental conditions are not favorable. Furthermore, we also
described that this cyclin was degraded by Dma1 instead of the canonical E3 ligases for cell cycle
cyclins, Grr1 nor Cdc4, and that this E3 ligase acts recognizing a sequence that we named DDD
(Hernández-Ortega et al., JBC, 2013).
Our group is now interested in describing the possible homologous of this cyclin in human. We are
currently investigating if cyclin I, a small human new cyclin that appears to be acting with CDK5, the
Pho85 homolog, is being phosphorylated by this CDK. In addition, cyclin I has a peak of expression in
G1/S as Pcl1 (Nagano et al., Cell cycle, 2013). We made an alignment between these two cyclins and
we saw that cyclin I has conserved the DDD sequence so we are trying to find if the mechanism of
degradation is also conserved by the RING human E3 ligases, Chfr or Rnf8.
This possible conserved mechanism of regulation could help to elucidate the specific roles of the
redundant CDK/cyclins complexes along the cell cycle or their function in different stages of
development or tissues.
66
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 30
A Consensus Motif for PP2A-B56
Emil Peter Thrane Hertz1, Thomas Kruse1, Jón Otti Sigurdsson1, Norman Davey2,
Jesper Velgaard Olsen1, Jakob Nilsson1
1
The Novo Nordisk Foundation Center for Protein Research, Faculty of Health and Medical Sciences, University of
Copenhagen, Blegdamsvej 3b, 2200 Copenhagen, Denmark
2 UCD Conway Institute of Biomolecular and Biomedical Research, University College Dublin, Belfield, Dublin 4, Ireland
Reversible protein phosphorylation is central to biochemical processes in all living organisms. During
mitosis, kinases and phosphatases ensure fidelity in chromosome alignment and segregation. While
the kinases working during mitosis have been extensively studied the characterization of the
opposing phosphatases remains rudimentary. Protein Phosphatase 2A (PP2A) in complex with B56
regulatory subunits is emerging as an essential regulator of various mitotic and meiotic events. How
PP2A-B56 is regulated and recruited to binding partners remains unknown. We have now for the
first time discovered a short consensus motif in proteins binding to PP2A-B56 allowing us to predict
binding partners and precisely address the role of these interactions. Furthermore, we have
identified a fully conserved binding pocket on B56 subunits that mediate the interaction with the
consensus motif. We will present our progress in understanding the molecular determinants of the
consensus motif that dictates interaction with PP2A-B56 and the function of this important
phosphatase.
67
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 31
De novo formation of centrosome-like structures and their
roles in the progression of the first mitosis and diploid nucleus
formation in parthenogenetic embryos of Drosophila
ananassae
Kazuyuki Hirai1, Haruka Suzuki2, Yohei Minakuchi3, Atsushi Toyoda3, Muneo Matsuda1
1
Kyorin University School of Medicine
International Christian University
3 National Institute of Genetics
2
Parthenogenesis, development of an unfertilized egg into a new individual, has been known to occur
naturally in a wide range of animal taxa. Genetic variants that allowed this unusual reproductive
strategy have arisen independently in evolutionary histories of sexual reproduction of each species.
Parthenogenetic Drosophila may serve as a useful experimental model for studying the adaptation
of basic nature in cytological as well as genetic respects for reproductive strategies. We have
recently established parthenogenetic strains of Drosophila ananassae and its closely related species.
Most of the unfertilized eggs laid by parthenogenetic mothers show developmental arrests and die
as haploid embryos, but others can develop into diploid females with a low but significant
probability (~8%). We now report the essential mechanisms by which the following problems are
solved: Mitotic spindle assembly around the female pronucleus without paternally-derived
centrosomes and restoration of diploidy. Our genetic and cytological experiments confirmed that
parthenogenetic eggs finished meiosis normally, giving rise to haploid gametes. We found that, in
unfertilized eggs of both sexual and parthenogenetic strains of the species, an acentrosomal
metaphase spindle is assembled around the chromosomes of the female pronucleus, after
completion of S phase. It is striking that eggs laid by parthenogenetic females assembled
centrosome-like structures de novo, which were scattered in the anterior part of the egg cytoplasm.
These microtubule organizing centers contain pericentriolar material recruitment factors Asterless
and Centrosomin. We observed some eggs with such centrosome-like structures located at one or
both of the poles of the bipolar spindle assembled around the female pronucleus. Interestingly,
anaphase or telophase figures were observed only in the presence of the centrosome-like structures
at spindle poles, while no mitotic stages beyond metaphase were observed in the absence of
centrosome-like structures at spindle poles in unfertilized eggs of sexual and parthenogenetics
strains. Our data suggest a role of the centrosome-like structures in stimulating the metaphaseanaphase transition by binding to the pole of the acentrosomal spindle around the female
pronucleus. The first mitosis produces two adjacent daughter nuclei, one associated with a
centrosome-like structure and the other without it. Two metaphase spindles are assembled around
the cleavage nuclei, but two poles from the separate spindles are often connected by a single
centrosome-like structure. We found that the first diploid nucleus is formed by fusion of two haploid
chromosome sets at telophase of the second nuclear cycle, resembling the gonomeric division in
sexual development. Among three nuclei produced by the second division, the diploid nucleus with a
centrosome-like structure might have the ability to proliferate preferentially. The function of
centrosomes essential for the initiation of sexual and parthenogenetic development will be
discussed.
68
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 32
Maintenance of genome integrity in Arabidopsis root meristem
is regulated by the RETINOBLASTOMA-RELATED protein
Beatrix Horvath1, 2, Hana Kourova3, Szilvia Nagy4, Zoltan Magyar5, Gabino SanchezPerez2, Csaba Papdi1, Edit Nemeth1, Tamas Meszaros4, Pavla Binarova3, Laszlo Bogre1,
Ben Scheres2, 6
1
School of Biological Sciences, Centre for Systems and Synthetic Biology, Royal Holloway, University of London, Egham Hill,
Egham, TW20 0EX, UK
2 Department of Molecular Genetics, Utrecht University, 3584 CH Utrecht, The Netherlands
3 Institute of Microbiology AS CS, Laboratory of Functional Cytology, v.v.i.,Víde?ská 1083 Prague 4 , 14 200, Czech Republic
4 Department of Medical Chemistry, Molecular Biology and Pathobiochemistry, H-1094, Budapest, Hungary
5 Institute of Plant Biology, Biological Research Centre, Temesvári krt. 62, POB 521, H-6701, Szeged, Hungary
6 Department of Plant Sciences, Wageningen University Research Centre, 6708 PB Wageningen, The Netherlands
Meristematic cells are prone to DNA damage, but are equipped with an array of response pathways
to protect genome integrity, including the activation of repair mechanisms, the delay or arrest of cell
cycle, the induction of differentiation or cell death. The RETINOBLASTOMA-RELATED is the major
regulator for cell proliferation, and its role during DNA damage response in plant is hitherto
unknown.
Using micro-array analysis, Chromatin –Immunoprecipitation and genetic studies
we show that RETINOBLASTOMA-RELATED, a central regulator for cell proliferation, in conjunction
with the E2FA and E2FB transcription factors, directly represses the expression of DNA damage
response genes; AtBRCA1, AtPARP2, and the newly characterised putative RING finger domain
containing E3 ligase, which we named LIFERING. AtBRCA1, a known DNA repair mediator, when
mutated becomes hypersensitive to genotoxin-induced cell death. LRN acts against the DNA
damage-induced cell differentiation, while AtPARP2 functions to restrain cell proliferation to impose
quiescence. RBR silencing leads on one hand to cell death, which relies on the function of the three
target genes, and on the other hand to overproliferation which depends on the opposing functions
of AtPARP2 and LIFERING. We conclude that RBR provides an input for mitogenic signals to regulate
distinct DNA damage responses. The balanced action of the RBR target genes, AtBRCA1, AtPARP2
and LIFERING collectively determines the outcome of DNA damage responses.
69
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 33
Cell length growth patterns in fission yeast reveal a novel size
control mechanism operating in late G2 phase
Anna Horváth, Ákos Sveiczer
Budapest University of Technology and Economics, Department of Applied Biotechnology and Food Science
Individual cell length growth patterns were analysed of several fission yeast strains, wild type and
some cell cycle mutants by fitting different functions to the data. During the growing period, most
cells’ growth patterns could be best described by a bilinear function (two linear segments separated
by a breakpoint, RCP2); however, linear patterns also occurred in several cases, but exponential ones
only very rarely. The bilinear patterns were separated into two growing parts by RCP2, and the
position of size control was examined by regression analyses of appropriate growth parameters in
both segments. The existence of known size controls in late G1, in mid G2, and in late G2 during the
fission yeast cell cycle was verified. However, in spite of the former general view, we now discovered
that late G2 size control is a general characteristic third event in the cycle, when cell size is
monitored. The level of the critical late G2 size to be reached in an individual fission yeast cell is
influenced by its growth rate, similarly to budding yeast, which suggests an evolutionary conserved
mechanism.
70
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 34
Modelling the role of systems-level feedbacks in the detection
of premature sister chromatid separation by the spindle
assembly checkpoint
Lukas Hutter1, Mihailo Mirkovic2, Raquel Oliveira2, Bela Novak1
1
2
Department of Biochemistry, University of Oxford
Instituto Gulbenkian de Ciencia, Lisbon
The Spindle Assembly Checkpoint (SAC) delays anaphase onset in response to chromosomes that are
erroneously attached to the mitotic spindle. Satisfaction of the SAC through biorientation of
chromosomes crucially depends on sister chromatid cohesion. However, recent findings suggest that
the SAC fails to robustly prevent mitotic progression in the absence of cohesin.
Here, we present a detailed theoretical study of the dynamics of the SAC in Drosophila Neuroblasts
that were challenged with precocious loss of cohesin. In this setup, full sister chromatid separation
does not elicit a robust checkpoint response and they abnormally exit mitosis with high segregation
errors after a short mitotic delay. By adopting a stochastic modelling approach to explain
quantitative live-cell imaging experiments performed on these cells, we are able to highlight the role
of multiple feedback loops in the mitotic signalling networks: our analysis shows how the coupling of
SAC-signalling and error correction mechanisms can promote the observed gradual accumulation of
stable end-on attachments as the signalling capacity of the SAC declines, and how transient
attachments in turn drive the decline in SAC signalling capacity. Thus, our results provide a
framework to understand how cohesion defects may escape SAC surveillance and therefore give rise
to aneuploid cells.
71
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 35
A dynamic description of progression through meiosis
Katarzyna Jonak, Wolfgang Zachariae
Laboratory of Chromosome Biology, Max Planck Institute of Biochemistry, 82152 Martinsried, Germany
Mitotic and meiotic divisions are controlled by the activities of Cdk1-cyclin B and APC/C-Cdc20.
However, these types of division differ in fundamental aspects. The events of the mitotic cell cycle
are controlled by a regulatory system that re-creates the starting-conditions for a new cycle. By
contrast, meiosis is a linear sequence consisting of a single S phase followed by two consecutive M
phases and a differentiation program dedicated to the generation of gametes or spores. It is unclear
how the mitotic cell cycle control mechanism is modified to regulate meiosis and, in particular, how
meiotic cells irreversibly exit from M phase after exactly two divisions. Thus, we combine
mathematical modelling with biological experiments in budding yeast to study the dynamics of key
meiotic regulators and to understand how these regulators form a network that generates two, and
only two, waves of Cdk1 and APC/C-Cdc20 activity. Recently, we have developed a model for the
irreversible transition from prophase I into metaphase I in response to the completion of meiotic
recombination (Okaz et al., 2012). To generate a model for the entire meiosis, we have
supplemented the prophase-to-metaphase I model with a Cdk1-APC/C-Cdc20 oscillator taken from
mitotic cell cycle models (Chen et al., 2004; Tyson and Novak, 2008) and a hypothetical terminator
mechanism that stops the oscillator at the exit from meiosis II. Currently, we are investigating the
mechanisms that govern exit from meiosis II in order to replace the hypothetical terminator with its
biological counterpart.
Chen, K., Calzone, L., Csikasz-Nagy, A., Cross, F.R., Novak, B., and Tyson, J.J. (2004). Mol. Biol. Cell
15(8), 3841-3862.
Okaz, E., Arguello-Miranda, O., Bogdanowa, A., Vinod, P.K., Lipp, J.J., Markova, Z., Zagoriy, I., Novak,
B., and Zachariae, W. (2012). Cell 151(3), 603-618.
Tyson, J.J., and Novak, B. (2008). Current Biology 18(17), R759-R768.
72
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 36
Assaying bistability and investigating its major determinants in
the G2/M switch system
Stephy Joseph
Genome damage and Stability centre
University of Sussex
Department of biochemistry
University of Oxford
Wellcome Trust Centre for Cell Biology
University of Edinburgh
Models of the Cdk1 activation feedback loops that ensure robust and irreversible transition between
G2 and M-phase suggest that this system behaves like a bistable switch. This model predicts that
cells can only be in a G2, or M-state but that intermediate states are not allowed implying a system
property termed hysteresis. If the G2/M transition displays hysteresis, Cdk1 activity thresholds
should differ between mitotic entry and exit. One would also expect a timelag in Cdk1 activation
inversely proportional to the input signal strength at levels close to the threshold. Both predictions
have been confirmed experimentally in cellfree Xenopus egg extracts. However, it remains to be
shown that the G2/M transition is bistable in intact mammalian cells. Moreover, we do not know
how the various components of the switch system contribute to bistability and how loss of this
property would affect mitotic progression. We have developed assays to investigate the dynamic
properties of the Cdk1 activation switch in cdk1 analogue sensitive DT40 and HeLa cells that allowed
us to verify bistability of the G2/M transition in these cellular model systems. We found that
modulating both the negative and positive feedback loops in the Cdk1 activation reaction shifts the
thresholds up and down, but does not abolish hysteresis. We are currently investigating how nuclear
translocation of cyclin B and cytoplasmic translocation of Greatwall kinase may affect the systems
behaviour and are hypothesising that these events may be the key determinants of hysteresis of the
Cdk1 activation switch. Overall, these data will be used to build an accurate and experimentally
verified mathematical model of the G2/M transition in mammalian cells.
73
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 37
Downregulation of PP2ACdc55 at anaphase onset by Zds1 and
separase
SORAYA JÁTIVA, et al.
BELLVITGE INSTITUTE OF BIOMEDICAL RESEARCH (IDIBELL)
Exit from mitosis and completion of cytokinesis require the inactivation of mitotic cyclin-dependent
kinase (Cdk) activity. In budding yeast, Cdc14 phosphatase is a key mitotic regulator that is activated
in anaphase to counteract Cdk activity. In metaphase, Cdc14 is kept inactive in the nucleolus
sequestered by its inhibitor Net1. At anaphase onset, downregulation of PP2ACdc55 phosphatase by
separase and Zds1 protein promotes Net1 phosphorylation and consequently, Cdc14 release from
the nucleolus. The mechanism by which Zds1 and separase impinge on PP2ACdc55 activity remains
to be elucidated. Previous results show that Zds1 exert its biological function as PP2ACdc55
regulator, by controlling the subcellular localisation of the PP2A regulatory subunit Cdc55. Our
previous results suggest that the activity of PP2ACdc55 cannot be modulated solely through
regulation of its localization, and that an additional regulatory step may be required to control
PP2ACdc55 activity during mitotic exit. Here we show that Cdc55 regulatory subunit is
phosphorylated during anaphase upon PP2ACdc55 downregulation. Our results suggest that
PP2ACdc55 activity is modulated throughout Cdc55 posttranslational modifications in a separase
and Zds1-dependent manner.
74
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 38
A comprehensive study of NRF2-controlled cell cycle arrest
during oxidative stress
Orsolya Kapuy1, Anita Kurucz1, Tamás Korcsmáros2, 3, Gábor Bánhegyi1
1
Department of Medical Chemistry, Molecular Biology and Pathobiochemistry, Semmelweis University, Budapest, Hungary
The Genome Analysis Centre, Norwich, UK
3 Institute of Food Research, Norwich, UK
2
Oxidative stress is one of the main environmental factors that can influence cellular homeostasis.
Treatment with oxidative agents (such as H2O2, tBHP) results in activation of several signal
transduction pathways of that stress response mechanism; meanwhile the ongoing cell division cycle
has to be blocked. NRF2 (nuclear factor erythroid 2-related factor 2) has a key role to enable cell
adaptation to oxidative stress via transcriptionally controlling more than 2000, mainly cytoprotective, genes. However, details about its effect on cell cycle regulation have not been explored
yet. Our goal is to reveal a crosstalk between NRF2 and the key elements of the cell cycle regulatory
network. We approach our scientific questions from a so called systems biological perspective by
using both experimental and theoretical methods.
It was shown recently that Cyclin D, the main inducer of G1/S transition, got immediately downregulated via PERK pathway in response to oxidative stress resulting in cell cycle arrest. We suggest
that PERK kinase inhibits the key cyclin molecule throughout NRF2 induction by using various
methods. We are planning to investigate whether this connection between NRF2 and Cyclin D is
direct and/or indirect. We assume that the indirect regulatory connection between NRF2 and Cyclin
D is generated by CDK inhibitors (for example p15, p16, p21, p27). We claim that a positive feedback
loop between NRF2 and CDK inhibitors assures a rapid cell cycle arrest even at low level of oxidative
stress. Performing a systematic analysis we explore the role of each above mentioned CDK inhibitor
in down-regulating Cyclin D with respect to oxidative agent. Our experimental results presumably
inter-correlated with our theoretically generated model. With computational simulations of the
underlying control network various mutant phenotypes might be also tested.
Our results suggest that the role of NRF2 during oxidative stress is even more complex as it was
known until now. NRF2 not only induces the survival process and arrests cell division cycle, but later
it is able to suppress the stress response by transcriptionally down-regulating the mechanism. We
claim that the generated feedback loops (both positive and negative ones) between NRF2 and its
targets guarantee the precise response mechanism with respect to oxidative agents. At low level of
stress NRF2 quickly induces the cyto-protective process, meanwhile mTOR (mammalian targets of
rapamycin), another key sensor of cellular homeostasis, gets inhibited. However at high level of
stress the cyto-protective process has only a transient induction followed by mTOR re-activation.
75
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 39
The role of Cyclin B3 in Mammalian Spermatogenesis
Mehmet Erman Karasu1, 2, Andrew C Koff1, 2, Scott Keeney1, 2, 3
1
Molecular Biology Program, Memorial Sloan Kettering Cancer Center
Louis V. Gerstner Jr. Graduate School of Biomedical Sciences, Memorial Sloan Kettering Cancer Center
3 Howard Hughes Medical Institute
2
Important components of the cell cycle machinery include cyclins and their catalytic kinase partners,
cyclin-dependent kinases (Cdks). It has been shown in budding yeast that temporal regulation of
cyclin-Cdk activity is critical for orchestrating key events during meiosis, as in mitosis. Such regulated
events include initiation of homologous recombination and segregation of homologous
chromosomes. Similar to budding yeast, in mouse male meiosis (spermatogenesis), cyclins show a
sequential expression pattern. In mouse, two cyclins – Cyclin A1 and Cyclin B3 – are meiosis-specific
and display interesting expression patterns. Cyclin A1 is expressed in spermatocytes from late
pachytene to diplotene, and disruption of Cyclin A1 causes arrest in meiotic prophase and infertility
in male mice. Cyclin B3 is expressed in both spermatocytes and oocytes from leptotene to zygotene,
when most DSB formation occurs, suggesting that Cyclin B3 may regulate events during early
meiosis, such as initiation of meiotic recombination or pairing of homologs. Moreover, in transgenic
mice in which expression of Cyclin B3 was prolonged until the end of meiosis, spermatogenesis was
disrupted and sperm count was reduced. Additionally, disruption of Cyclin B3 in flies resulted in
infertility in females. Based on these findings, Cyclin B3 has long been thought to be a prime
candidate for a key cell cycle regulator of early meiotic events. To investigate the role of Cyclin B3 in
meiosis, we generated Cyclin B3 knock-out (KO) animals by CRISPR-Cas9 genome editing. We
obtained in-frame deletions, out-of-frame deletions, substitutions and insertions at the Cyclin B3
locus. Analysis of Cyclin B3 KO animals indicate that Cyclin B3 protein is not present, and mRNA
transcript levels are reduced 6-10 fold. Surprisingly, however, Cyclin B3 KO mice are fertile and testis
size and weight are comparable to wild-type littermates. The detailed phenotype of Cyclin B3 KO
mice will be presented.
76
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 40
HOW DOES CDC14 ORDER EVENTS DURING MITOTIC EXIT IN
SACCHAROMYCES CEREVISIAE?
Meghna Kataria, Frank Uhlmann
The Francis Crick Institute - Lincoln's Inn Fields
Accurate progress through the cell division cycle necessitates robust order and timing of various
cytological events, the molecular basis for which is incompletely understood. The quantitative model
of cell cycle progression proposes that this order is dictated by increasing kinase activity as cells
traverse through the various stages. Lately, the historically overlooked contribution of the
concomitant phosphatases has come into focus – a process well studied in mitotic exit in budding
yeast. In this organism, Cdc14 is the major phosphatase which reverses cyclin-dependent kinase
(Cdk)-mediated phosphorylation events upon its activation at anaphase, and is required for cells to
exit mitosis. Ordered dephosphorylation of proteins dictates the sequence of events during mitotic
exit and cytokinesis. This order is largely imposed by differing catalytic efficiencies of Cdc14 towards
its substrates. This has spurred the idea that preferred substrates might possess high affinity for the
phosphatase.
In this study, we seek a molecular understanding of mitotic exit by probing the biochemical
characteristics of Cdc14 – sequence features within substrates that might underlie differential
affinities. We probe this by characterizing its interactions with substrates that are dephosphorylated
very early into anaphase, such as Fin1. Preliminary data indicates that Cdc14 binds to Cdk1
phosphorylation sites, in addition to other features that are now being validated. The eventual
objective is to alter the dephosphorylation timing of ‘early’ proteins not only in vitro, but also in vivo
to study its effects on cell cycle progression.
77
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 41
The condensin component NCAPG2 regulates chromosome
segregation
Jae Hyeong Kim1, Jaegal Shim1, Seoung Min Bong1, Young-Joo Jang2, Chang-Woo Lee3,
Byung Il Lee1,Kyungtae Kim1, et al.
1
Research Institute, National Cancer Center, Goyang, Gyeonggi 410-769, Republic of Korea
Laboratory of Cell Cycle and Signal Transduction, Department of Nanobiomedical Science & BK21 PLUS Research Center
for Regenerative Medicine, Dankook University, Cheonan, Chungnam 330-714, Republic of Korea
3 Department of Molecular Cell Biology, Samsung Biomedical Research Institute, Sungkyunkwan University School of
Medicine, Suwon, Gyeonggi 440-746, Republic of Korea
2
The early event of microtubule–kinetochore attachment is a critical stage for precise chromosome
segregation. Here we report that NCAPG2, which is a component of the condensin II complex,
mediates chromosome segregation through microtubule–kinetochore attachment by recruiting PLK1
to prometaphase kinetochores. NCAPG2 colocalises with PLK1 at prometaphase kinetochores and
directly interacts with the polo-box domain (PBD) of PLK1 via its highly conserved C-terminal
phospho-Thr (1007VLS-pT-L1011) containing region and conserved crystal structure with other PBD
binding phospho-peptide. In both humans and in vivo model system, when NCAPG2 is depleted, the
attachment of the spindle to the kinetochore is loosened and misoriented, caused by the disruption
of PLK1 localisation to the kinetochore and by the decreased phosphorylation of its kinetochore
substrate, BubR1. These findings suggest that NCAPG2 plays an important role in regulating proper
chromosome segregation through a functional interaction with PLK1 during mitosis.
78
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 42
Interaction of MAP kinase MPK6 with γ-tubulin and
microtubule plus-end protein EB1c and regulatory function of
MAP kinases in mitotic chromosome segregation in
acentrosomal plant cells
Lucie Kohoutová1, Hana Kourová1, Szilvia K. Nagy2, Jindřich Volc1, Petr Halada1, Tamás
Mészáros2, Irute Meskiene3, László Bögre4, Pavla Binarová1
1
Institute of Microbiology AS CR, v. v. i., Videnska 1083, 142 20 Prague 4, Czech Republic
Semmelweis University, Department of Medical Chemistry, Molecular Biology, Budapest, Hungary
3 Max F. Perutz Laboratories, University of Vienna, Vienna, Austria
4 Royal Holloway, University of London, School of Biological Sciences, Egham, United Kingdom
2
Mitogen-activated protein (MAP) kinases pathways play role in stress response and developmental
signalling in eukaryotes and modulate cell division through cytoskeleton. Knowledge of protein
targets of MAP kinase signalling among microtubular proteins is limited and uncovering of new
substrates and interactors of MAP kinases can help understanding of the cell cycle regulation.
We found an interaction of MPK6 and γ-tubulin in cytoplasmic extract and with microtubules in
Arabidopsis. Signal of active MAP kinases detected by p-ERK antibody was present together with γtubulin on the specific subset of mitotic microtubules. We further found that microtubule plus-end
binding protein EB1c interacts with MPK6. We showed that EB1c is phosphorylated by MKK4activated MPK6 in vitro while phosphorylation was not shown for EB1a and γ-tubulin. Moreover, we
confirmed phosphorylation of EB1c in vivo in Arabidopsis and the phosphorylation was reduced after
treatment with specific MEK1/2 inhibitor U0126. Active MAP kinases and EB1c-GFP protein localized
with mitotic spindles with a slight enrichment in vicinity of the kinetochores and on shortening
kinetochore fibres.
Arabidopsis plants with reduced activity of MPK3, MPK4, and MPK6 kinases due to overexpression of
MAP kinase phosphatase AP2C3 showed mitotic and cytokinetic defects and misaligned spindles and
phragmoplasts. We found that congression and segregation of chromosomes was impaired in AP2C3
plants and similar defects of spindle assembly checkpoint were observed when activity of MAP
kinases was inhibited by U0126. KO mutants mpk6-2 showed misaligned spindles only in response to
stress.
Our data showed that γ-tubulin, a protein involved in microtubule nucleation and organization, and
microtubule plus-end protein EB1c with function in the cell division are two novel partners for MPK6
signalling. Furthermore, our data suggest synergistic and partially overlapping functions for MPK3,
MPK4, and MPK6 kinases in regulation of mitosis in plant cells.
Supported by Grant from Grant Agency of the Czech Republic P501-15-1657S
79
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 43
Functional analysis of the spindle checkpoint in plants - are we
really all MAD?
Shinichiro Komaki, Arp Schnittger
University of Hamburg
Cell division accuracy is required for genomic stability and proper organ development. The spindle
assembly checkpoint (SAC) plays a key role for the fidelity of cell division by preventing anaphase
onset until all chromosomes are correctly attached to the spindle. The core proteins of the SAC are
highly conserved from yeast to plants, including MAD1, MAD2, MAD3/BUBR1, BUB1, BUB3 and
MPS1. In yeast and mammalian cells, lack of the SAC activity causes missegregation of
chromosomes, resulting in aneuploidy. In plants, homologs of the core SAC components were
identified but surprisingly, all but one mutant in the core SAC genes are viable raising the question
whether the SAC or its regulators have a conserved function in the eukaryotic kingdom. To
investigate the requirement of the SAC activity in plants, mutants for the central SAC components of
Arabidopsis were also grown on media containing a microtubule-destabilizing drug oryzalin. Oryzalin
reduced root growth of mad1, mad2, bubr1, mad3.2 and mps1, suggesting that at least some of the
molecular functions of the SAC components are also conserved in Arabidopsis. Examination of the
expression patterns of the core SAC components during plant development using different genomic
fusions of the respective regulators to beta-glucuronidase (GUS) showed that most of the
Arabidopsis SAC genes have complex regulatory elements, most likely present in their intron and/or
3’ UTR region that may serve to integrate developmental with environmental signals. Based on the
GUS reporter lines, we have then constructed protein reporters fused to GFP. Live imaging revealed
that MAD1-GFP and MAD3.2-GFP localized to unattached kinetochores, which is the typical
localization pattern of the SAC reported previously. However, the other SAC components showed
diverse localization patterns. In particular, BUB3.1-GFP and BUB3.2-GFP concentrated at the
midzone of the plant-specific cytokinetic phragmoplast, suggesting a function of the SAC in
cytokinesis. Further analyses on the role of the SAC in mitosis and during cytokinesis are currently
under way.
80
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 44
CDK1 structures reveal conserved and unique features of the
essential cell cycle CDK.
Svitlana Korolchuk2, Nicholas Brown1, Mathew Martin2, Will Stanley2, Martin Noble2,
Jane Endicott2
1
2
MRC National Institute for Medical Research
Newcastle University
CDK1 is the only essential cell cycle CDK in human cells and is required for successful completion of
M-phase. It is the founding member of the CDK family and is conserved across all eukaryotes. We
have determined the crystal structures of complexes of CDK1–Cks1 and CDK1–cyclin B–Cks2. These
structures reveal a number of important aspects of CDK1 structure and function. They confirm the
conserved nature of the inactive monomeric CDK fold and its ability to be remodeled by cyclin
binding. In the context of previously reported CDK-cyclin structures, the structure of CDK1–cyclin B
increases the observed diversity with which the CDK responds to cyclin binding. Relative to CDK2–
cyclin A, CDK1–cyclin B is less thermally stable, has a smaller interfacial surface, is more susceptible
to activation segment dephosphorylation, and shows differences in the substrate sequence features
that determine activity. CDK1–cyclin B demonstrates a relatively relaxed specificity for residues
adjacent to the site of phosphorylation, a phenomenon that correlates with this CDK–cyclin pair
appearing to have a less rigidly constrained activation segment. The CDK1–cyclin B structures also
reveal potential novel protein interaction sites that might regulate CDK1 activity. In addition to a
conserved surface on the CDK1 C-terminal fold, there is a cleft between the two subunits not seen in
the structure of CDK2–cyclin A but apparent in CDK9–cyclin T, where it is exploited by the human
immunodeficiency virus 1 Tat. Both CDK1 and CDK2 are potential cancer targets for which selective
compounds are required. We have also determined the first structure of CDK1 bound to a potent
ATP-competitive inhibitor and identify aspects of CDK1 structure and plasticity that might be
exploited to develop CDK1-selective inhibitors. We are currently aiming to extend our observations
to characterise CDK1 activation by cyclin A and other activators.
81
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 45
Role of the DmUsp5 in coupling ubiquitin homeostasis to
development and apoptosis
Levente Kovács1, Olga Nagy1, Margit Pál1, Octavian Popescu2, Andor Udvardy1, Péter
Deák1
1
2
Institute of Biochemistry, Biological Research Centre, Hungarian Academy of Sciences, Szeged, Hungary
Interdisciplinary Institute of Bio-Nano-Sciences, Molecular Biology Center, Cluj-Napoca, Romania
Ubiquitin is a small regulatory protein which is used for posttranslational modification of proteins in
a reversible process called ubiquitylation. An enzyme cascade catalyzes the attachment of ubiquitins
to the target proteins. This covalent modification facilitates protein-protein interactions, changes
enzymatic activities, or targets proteins to proteasomal degradation. This dynamic process is
implicated in many basic cellular functions including cell cycle and cell death. For all these functions,
the ubiquitylation requires a free mono-ubiquitin pool which is maintained most prominently by
ubiquitin recycling. Members of the proteases family called deubiquitylating enzymes or DUBs are
implicated in ubiquitin recycling since they can remove ubiquitins from target proteins or process
polyubiquitins. Although DUBs seem to be indispensable for the maintenance of ubiquitin
homeostasis, their precise physiological importance is still poorly understood.
In the genetically well characterized Drosophila model, 46 DUBs have been identified, but the
function of most of them is still unknown. One of them is Usp5, an evolutionarily conserved DUB
enzyme involved in the disassembly of unanchored polyubiquitin chains. We identified and
genetically analyzed the Drosophila orthologue of the human Usp5 (DmUsp5) which turned out to
be essential for normal development. A heterologous complementation experiment confirmed
functional homology between the DmUsp5 gene and its yeast homologue, Ubp14. Loss of DmUsp5
function results in late lethality that is accompanied by the accumulation of unanchored
polyubiquitin chains. It also stabilizes p53 and induces a high incidence of apoptosis in larval brains
and imaginal discs. In addition to this, the expression of reaper and hid, but not the grim, proapoptotic genes becomes elevated in DmUsp5 mutants. Most importantly, the expression of
another, proteasome-associated DUB, DmUsp14 increased highly in DmUsp5 mutants. It was shown
in the unicellular budding yeast that Usp14 orthologue, Ubp6 is expressed and progressively
deubiquitylate proteasome-bound substrates at times of ubiquitin depletion. Elevated DmUsp14
expression together with dominant cycloheximide sensitivity indicates that loss of DmUsp5 cause
ubiquitin stress in these animals. These observations suggest that the DmUsp5 DUB enzyme plays a
critical role in regulating apoptosis and – together with DmUsp14 - moderating ubiquitin
homeostasis in Drosophila. This work is supported by HURO/1101/173/2.2., TAMOP-4.2.2.A11/1/KONV-2012-0035 and TAMOP- 4.1.1.C-13-1-KONV grants.
82
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 46
Axial contraction and short-range compaction of chromatin
synergistically promote the condensation of mitotic
chromosomes.
Tom Kruitwagen1, Annina Denoth-Lippuner1, Brian Wilkins2, Heinz Neumann2, Yves
Barral1
1
2
Institute of Biochemistry, ETH Zürich, Switzerland
Department of Applied Synthetic Biology, University of Göttingen, Germany
Prior to their symmetric segregation during mitosis, eukaryotic chromosomes must be extensively
folded into units that the spindle can act upon. Despite its importance, the molecular mechanisms
underlying chromosome condensation are still unclear. Serine 10 on histone 3 (H3 S10) has been
shown to be specifically phosphorylated during mitosis. In most eukaryotes, however, the role and
importance of this Aurora B dependent event has remained unclear, since the mutation of this
residue from serine to alanine failed to show any clear phenotype. However, we recently showed
that the yeast Sir2-related histone deacetylase Hst2 is recruited by H3 S10Ph and to deacetylate
histone 4 at lysine 16 (H4 K16), to promote mitosis-specific interactions between neighbouring
nucleosomes via H2A and H4.
A clearer role was assigned to the condensin complex, whose disruption lead to widespread defects
in chromosome condensation in frog egg extracts. However, in other model organisms condensin
seemed to be partially dispensable for chromosome condensation. Thus, although a number of
factors contribute to chromosome condensation, how they act together to shape mitotic
chromosomes remains unclear.
Here, we use two microscopic assays to distinguish short-range nucleosome-nucleosome
interactions from long-range, axial contraction of mitotic chromosomes. Using these methods, we
demonstrate that H3 S10 phosphorylation and the deacetylation of histone H4 tails in early
anaphase promote short-range compaction of chromatin through nucleosome-nucleosome
interactions, starting from the centromere. During late anaphase, condensin mediates the axial
contraction of chromosome arms. These processes are temporally and mechanistically distinct, since
mutants disrupting chromatin compaction, such as H3 S10A, have no observable effects on axial
contraction, and vice versa. When both pathways are inactivated, synergistic defects on
chromosome segregation and cell viability are observed. Interestingly, both pathways rely at least
partially on the deacetylase Hst2, the only factor to show both compaction and contraction defects
when deleted.
83
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 47
Rif1 is required for resolution of Ultrafine DNA Bridges in
anaphase to ensure genomic stability
Rutger Hengeveld1, Rudolf de Boer2, Pepijn Schoonen2, Elisabeth de Vries2, Marcel
van Vugt2, Susanne Lens1
1
Department of Molecular Cancer Research, University Medical Center Utrecht, Universiteitsweg 100, 3584 CG Utrecht, The
Netherlands.
2 Department of Medical Oncology, University Medical Center Groningen, University of Groningen, Hanzeplein 1, 9723 GZ
Groningen, The Netherlands.
Sister-chromatid disjunction in anaphase requires the resolution of DNA catenanes by
topoisomerase II together with PICH (Plk1-interacting checkpoint helicase) and BLM (Bloom’s
helicase). We here identify Rif1 as a novel factor involved in the resolution of DNA catenanes that
are visible as ultrafine DNA bridges (UFBs) in anaphase to which PICH and BLM localize. Rif1, which
during interphase functions downstream of 53BP1 in DNA repair, is recruited to UFBs in a PICHdependent fashion, but independently of 53BP1 or BLM. Similar to PICH and BLM, Rif1 promotes the
resolution of UFBs: Its depletion increases the frequency of nucleoplasmic bridges and RPA70positive UFBs in late anaphase. Moreover, in the absence of Rif1, PICH or BLM more nuclear bodies
with damaged DNA arise in ensuing G1 cells, when chromosome decatenation is impaired. Our data
reveal a thus far unrecognized function for Rif1 in the resolution of UFBs during anaphase to protect
genomic integrity.
84
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 48
Molecular ties between the cell cycle and differentiation in
embryonic stem cells
Victor Li, Marc Kirschner
Harvard Medical School
Attainment of the differentiated state during the final stages of somatic cell differentiation is closely
tied to cell cycle progression. Much less is known about the role of the cell cycle at very early stages
of embryonic development. Here, we show that molecular pathways involving the cell cycle can be
engineered to strongly affect embryonic stem cell differentiation at early stages in vitro. Strategies
based on perturbing these pathways can shorten the rate and simplify the lineage path of ES
differentiation. These results make it likely that pathways involving cell proliferation intersect at
various points with pathways that regulate cell lineages in embryos and demonstrate that this
knowledge can be used profitably to guide the path and effectiveness of cell differentiation of
pluripotent cells.
85
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 49
Feedback throughout G2 coordinates production of proteins
required for mitosis with mitotic entry
Karen Akopyan, Arne Lindqvist
Department of Cell and Molecular Biology, Karolinska Institutet, Stockholm, Sweden
During G2 phase a cell is considered to prepare for mitosis, but how such preparation is coordinated
with mitotic entry remains poorly understood. We have recently used quantitative
immunofluorescence to acquire temporally resolved data through the cell cycle and found that the
mitotic kinases Plk1 and Cdk1 are active throughout G2 phase (Akopyan et al, Mol Cell 2014). We
find that fitting the quantitative data to an ODE-based model of mitotic entry is most
straightforward if Cdk1 activity enhances Cyclin B protein levels. In accordance, we find that Cdk1
inhibition in G2 phase decreases gene-targeted Cyclin B1-YFP levels. Interestingly, during forced
stimulation of mitotic entry by Wee1 inhibition, the duration of mitosis inversely correlates to Cyclin
B1-YFP levels, suggesting that mid G2 cells lack factors required for mitotic progression. Similarly, a
pulse of Cdk1 inhibition in G2 phase prolongs the duration of a forced mitosis, despite full activation
of Cyclin B-Cdk1. Our working model is that Cdk1 through a feedback-loop increases mitotic entry
network components during G2 phase. Such a feedback could potentially synchronize production of
proteins required for a successful mitosis with timing of mitotic entry and may indicate why G2 takes
so long in human somatic cells.
86
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 50
Spatial-temporal model for silencing of mitotic spindle
assembly checkpoints
Jian Liu, et al.
National Institutes of Health, Bethesda, MD 20892, USA
The spindle assembly checkpoint arrests mitotic progression until each kinetochore secures a stable
attachment to the spindle. Despite fluctuating noise, this checkpoint remains robust and remarkably
sensitive to even a single unattached kinetochore among many attached kinetochores; moreover,
the checkpoint is silenced only after the final kinetochore-spindle attachment. Experimental
observations showed that checkpoint components stream from attached kinetochores along
microtubules toward spindle poles. Here, we incorporate this streaming behavior into a theoretical
model that accounts for the robustness of checkpoint silencing. Poleward streams are integrated at
spindle poles, but are diverted by any unattached kinetochore; consequently, accumulation of
checkpoint components at spindle poles increases markedly only when every kinetochore is properly
attached. This step-change robustly triggers checkpoint silencing after, and only after, the final
kinetochore-spindle attachment. Our model offers a unified framework that highlights systems-level
coupling between kinetochore-spindle attachment and spindle-pole formation in SAC silencing. It
not only explains the remarkable fidelity of chromosome segregation in normal somatic cell mitosis,
but also accounts for the frequent aneuploidy rate in cancer cell mitosis and mammalian oocyte
meiosis I.
87
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 51
Cell cycle control in MutiCellular Tumor Spheroids
Aurélie Gomes1, 2, Odile Mondésert1, 2, Céline Frongia1, 2, Bernard Ducommun1, 2,
3
, Valérie Lobjois1, 2
1
Université de Toulouse, ITAV-USR3505, Toulouse, France
CNRS, ITAV-USR3505, Toulouse, France
3 CHU de Toulouse,Toulouse, France
2
Major recent advances have recently been made in the understanding of the molecular pathways
regulating the cell cycle machinery and their deregulations in cancer. These results open new
avenues for the development of innovative antitumor pharmacological strategies targeting the cell
cycle and its checkpoints.
MultiCellular Tumor Spheroid (MCTS) are now considered as invaluable models to study cancer cell
biology and for the preclinical development of new antiproliferative drugs.
To fully exploit the features of MCTS, it is essential to preserve the 3D regionalization level of
analysis when investigating the effect of a drug. We will report here a breakthrough in the use of
MCTS with new technological developments that can be used to explore the regionalization and the
live dynamics of the effects of anticancer drugs in 3D.
Cell lines expressing fluorescent cell cycle reporters (i.e. Fucci) and biosensors were engineered and
used to study cell cycle dynamics and to characterize the cell cycle parameters regionalization within
growing spheroids. We examined the impact of anti-tumor drugs on cell cycle progression and DNA
damage Response (DDR) pathway activation induced by chemotherapeutic agents within spheroids
using innovative 3D imaging strategy based on 3D light sheet microscopy.
This study opens new perspectives for the understanding of cell cycle control and proliferation in 3D
multicellular models and for the investigation of the dynamics of the 3D response to novel
antiproliferative agents.
88
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 52
Greatwall dephosphorylation and inactivation upon mitotic exit
is triggered by PP1
Sheng Ma, Perle Robert, Suzanne Vigneron, Anna Castro, Thierry Lorca
CNRS-CRBM
Mitotic entry requires cyclin B-Cdk1 activation and PP2A-B55 inhibition. At G2/M the Greatwall
kinase (Gwl) is activated and promotes Arpp19/ENSA phosphorylation that once phosphorylated
bind and inhibit PP2A-B55. Mitotic exit requires the reactivation of this phosphatase by Gwl
inactivation and further dephosphorylation of its substrates. In this study we show that PP1
depletion in Xenopus egg extracts stabilises phosphorylation of Gwl on the activatory site S875
maintaining in this way the full activity of this kinase and blocking meiotic/mitotic exit. In contrast,
the inhibition of PP2A-B55 results in Thr194 stabilisation, partial Gwl inactivation and partial
meiotic/mitotic exit. Our data also show that if PP2A-B55 is fully reactivated by Arpp19 depletion,
this phosphatase is able to rapidly dephosphorylate Gwl in the two activatory sites even in the
absence of PP1 suggesting that PP1 would be essential to trigger the Gwl inactivation patwhay.
Finally, we show that both phosphatases PP1 and PP2A-B55 can dephosphorylate Gwl on S875 and
Thr194 in vitro. These data identify PP1 as the phosphatase first dephosphorylating Gwl and
triggering its inactivation upon meiotic/mitotic exit.
89
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 53
Developmental regulation of cell proliferation by E2FB in leaf
pavement cells and meristemoids
Tünde Leviczky1, Binish Mohammed2, Aladár Pettko-Szandtner1, Beatrix Horváth2,
Anita Kovács1, Márta Deli1, Csaba Papdi2, László Bögre2, Zoltán Magyar1
1
2
Institute of Plant Biology, Biological Research Centre, Hungarian Academy of Science, Szeged, Hungary
School of Biological Sciences, Royal Holloway University of London, Egham, United Kingdom
Leaf growth relies on the number of cells produced before pavement cells exit proliferation and the
number of daughter cells generated by asymmetric cell divisions of meristemoids. E2FB is an
activator of cell cycle genes in proliferating pavement cells in young developing leaves. In
accordance, pavement cells of e2fb mutant prematurely exit cell proliferation while elevated E2FB
level delays this exit. During cell cycle exit, RBR represses E2FB activity. We found that E2FB
regulates RBR both transcriptionally and on protein level but also controls its phosphorylation level.
In accordance CYCD3;1 transcript level was elevated in the ectopic E2FB lines. These feedbacks could
provide the underlying mechanism, which controls the switch from cell proliferation to cell cycle exit
and differentiation. Therefore elevated E2FB level can alter this switch in cells started to
differentiate and they reenter proliferation. In contrast, overexpression of a dominant negative E2FB
form, E2FB∆RBR, which is unable to activate cell cycle genes, results in early exit from proliferation
and therefore producing smaller leaves. In leaf meristemoids, change in E2F function has totally
different outcomes. On one hand, e2fb mutation causes extra divisions of the meristemoids resulted
in stomata clusters. On the other hand co-overexpression of the mutant E2FB∆RBR construct with
DPA expands the transit amplifying cells produced by meristemoid stem cells. RBR together with
E2FB regulates these processes as a repressor. Thus E2FB has dual functions regulated by RBR. (1)
RBR represses E2FB activity, dependent on its phosphorylation, regulating cell cycle exit in pavement
cells, (2) E2FB forms a repressor complex with RBR to regulate stem cell divisions in meristemoids.
90
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 54
Putting the brakes on Mps1 Kinase.
Davinderpreet Mangat, Ulrike Gruneberg
University of Oxford
Faithful execution of cell division is fundamental for the creation and survival of all organisms. The
spindle assembly checkpoint (SAC) is a cell cycle surveillance mechanism that delays mitotic exit in
the presence of unattached kinetochores. Both the timely activation and inactivation of the mitotic
checkpoint are important for accurate chromosome segregation. The Mps1 kinase, a highly
conserved serine/threonine kinase has emerged as a central regulator of the SAC. Mps1 kinase
triggers SAC activation by phosphorylating a number of substrates, of which phosphorylation of the
scaffold protein Knl1 is best understood. Mps1 mediated phosphorylation of Knl1 provides docking
sites for downstream checkpoint proteins including Bub1, Bub3 and BubR1. Inhibiting Mps1 with the
specific inhibitor, AZ3146, results in the loss of the Bub and Mad proteins from the kinetochore.
Thus, given the critical role Mps1 plays in SAC activation, we hypothesised that the regulation of
Mps1 activity may be important for SAC signalling.
It has previously been shown that autophosphorylation of the activation loop residue T676 is
necessary for efficient SAC signalling. Since the activation of Mps1 is phospho-dependent, we
investigated whether this event is negatively regulated by a mitotic phosphatase. Here, we have
taken an unbiased siRNA based approach to determine the protein phosphatase responsible for
controlling Mps1 activity. Our results suggest that a PP2A-B56 phosphatase complex opposes Mps1
activation by removing a phosphate group from Mps1’s activation loop. Knockdown of the catalytic
or regulatory subunits of this phosphatase complex results in the retention of the phospho-Mps1
signal. Moreover, the activation loop of Mps1 is a substrate for PP2A phosphatase complexes in
vitro. We suggest that PP2A-B56 promotes mitotic exit by inactivating Mps1 kinase as well as directly
inhibiting the Knl1-Bub1 interaction.
91
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 55
Phosphorylation on serine 283 of human CDC25A modulates its
stability and activity in G2 and contributes to accelerate entry
into mitosis
Laurent Mazzolini1, Christine Dozier1, Carine Froment2, Marlene Marcellin2, Odile
Schiltz2, Stéphane Manenti1
1
Cancer Research Center of Toulouse, INSERM Unité Mixte de Recherche 1037, CNRS Equipe de Recherche labellisée 5294,
Université de Toulouse, 2, avenue Hubert Curien 31037 Toulouse Cedex, France
2 Institute of Pharmacology and Structure Biology, CNRS Unité Mixte de Recherche 5089, Université de Toulouse, 205, route
de Narbonne 31077 Toulouse Cedex, France
In eukaryotic cells the progression through the cell cycle is tightly regulated in response to
intracellular as well as extracellular signals. Central actors of this regulation are constituted by cyclindependent kinases (CDK) which, together with their associated cyclin regulatory subunit, control all
cell cycle transitions. The activity of CDK-cyclin complexes themselves is subject to complex
regulatory networks involving mostly activating and inhibitory phosphorylation events. The removal
of key inhibitory phosphorylations on CDK is performed by the dual specificity phosphatases of the
CDC25A family. In mammalian cells three CDC25 isoforms have been identified: CDC25A, CDC25B
and CDC25C. While CDC25B and CDC25C activity appears restricted to the regulation of CDK-cyclin
complexes involved in mitotic onset, CDC25A regulates both G1/S and G2/M progression. The
intracellular function of CDC25A is strongly controlled by phosphorylation events which regulate its
stability and activity and are performed by numerous kinases including the CDK themselves. In order
to identify new putative regulatory phosphorylations that may impact CDC25A function in the cell,
phosphoproteomic studies of CDC25A were undertaken in our team. This led to the identification of
new phosphorylation sites among which one, phosphorylation of serine 283, was found to occur in
the G2 phase of the cell cycle. This phosphorylation appeared maintained in mitosis and is lost when
cells reenter into the next cell cycle. Transient tranfection studies and use of specific kinase
inhibitors indicate that this phosphorylation mainly involves CDK-cyclin complexes. Functional
studies using cell lines conditionally expressing non-phosphorylable CDC25A mutants showed that
this phosphorylation does not impact CDC25A intracellular localization but increases the stability of
CDC25A as well as its intracellular activity in G2, contributing to accelerate the G2 to M transition in
cells
92
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 56
Asymmetric segregation of a lysine deacetylase links
perinuclear organisation and the time of cell cycle entry in
budding yeast
Manuel Mendoza, et al.
Center for Genomic Regulation (CRG)
Nuclear organization is associated with cell type-specific patterns of gene expression. However, how
differences in nuclear architecture are established during development is not known. Here we report
that in asymmetrically dividing budding yeast, the lysine deacetylase Hos3 shapes perinuclear
organization in daughter cells to regulate the starting time of the next cell division cycle. Hos3
asymmetrically localizes to the periphery of anaphase daughter nuclei, where it associates with
nuclear pores. We find that the G1/S cyclin CLN2 locus is confined to the perinuclear region during
G1 in a Hos3-dependent manner, to favor its transcriptional repression. In hos3∆ mutants, the
perinuclear anchoring of the CLN2 locus is lost, resulting in premature transition into S phase.
Conversely, artificial anchoring of the CLN2 locus to the nuclear periphery delays onset of S phase,
even in the absence of Hos3. Thus Hos3 determines the time of cell cycle entry specifically in
daughter cells, possibly by regulating the nuclear localisation of the CLN2 locus. These data open the
possibility that asymmetric partitioning of lysine deacetylases establishes differences in nuclear
architecture to shape developmental decisions during asymmetric division of animal cells.
93
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 57
Identification of novel interaction partners of the Chromosomal
Passenger Complex
Amanda Meppelink, Sibel Bayrak, Susanne Lens
Department of Molecular Cancer Research, ?Center for Molecular Medicine, University Medical Center Utrecht, The
Netherlands
During mitosis the, in S-phase duplicated genomic contents of a cell needs to be equally distributed
over two new daughter cells. To accomplish this the kinetochores of the sister-chromatids have to
become attached to microtubules emanating from opposite spindle poles (bipolar attachments) In
anaphase the two sisters will then be pulled towards opposing sites of the cell. Chromosome biorientation is essential for genomic stability, and is supported by a correction system that
disconnects erroneous kinetochore-microtubule attachments, and that allows stabilization of
correct, bipolar attachments. The chromosomal passenger complex (CPC), consisting of INCENP,
Survivin, Borealin and Aurora B kinase, is a key component of the error correction machinery.
Through phosphorylation of kinetochore substrates that are involved in microtubule binding and in
the recruitment of mitotic checkpoint proteins, it destabilizes incorrect kinetochore-microtubule
attachments and prevents anaphase onset by keeping the mitotic checkpoint active [1]
To further elucidate the role of the CPC during mitosis, we performed large-scale
immunoprecipitation experiments in combination with mass spectrometry for the various CPC
subunits to identify potential novel interaction partners. Pull-downs were performed in HeLa Kyoto
cells expressing bacterial artificial chromosome (BAC) transgenes for Aurora B, INCENP and Borealin
[2]. BACs contain large stretches of DNA encoding the complete gene including their endogenous
regulatory sequences, resulting in near endogenous expression. All three proteins were C-terminally
tagged with a modified version of the localization and affinity purification (LAP) tag [2]. Cells were
blocked in mitosis with the Eg5 inhibitor STLC, to mimic a prometaphase-like state.
Upon immunoprecipitation (IP) of all three separate CPC components we could identify the
remaining three CPC subunits. Next to these we also identified CBX1, CBX3, CBX5, and KIF20A as
constant and significant interaction partners, often with a high intensity. We could thus efficiently
pull down the entire complex together with various interaction partners.
Interestingly, in the IPs of Borealin-LAP we identified SIRT1, a histone deacetylase protein that
belongs to the sirtuin family and is known to also deacetylate proteins other than histones [3]. Using
Western blot analysis we could validate SIRT1 as a true interactor of Borealin after IP from BorealinLAP cell lines as well as by co-IP of overexpressed SIRT1 and Borealin in HEK293T cells. Interestingly,
we never found SIRT1 in the Aurora B or INCENP IPs, both after analysis by mass spectrometry or by
Western blot, suggesting that SIRT1 might interact with a pool of Borealin that is not associated with
the other CPC subunits. Recently a role has been proposed for SIRT1 in chromosome condensation
during mitosis and ensuring faithful chromosome distribution [4,5]. We are currently investigating
whether the SIRT1-borealin association is involved in histone (de)acetylation and how this could
affect faithful chromosome segregation
[1] Lampson, M. A., & Cheeseman, I. M. (2011). Trends in Cell Biology
[2] Poser, I., et al. (2008). Nature Chemical Biology
[3] Blander, G., & Guarente, L. (2004). Annual Review of Biochemistry
[4] Fatoba, S. T., & Okorokov, A. L. (2011). Cell Cycle.
[5] Kim, J.-J., et al. (2015), Journal of Cellular Biochemistry
94
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 58
Investigating the factors regulating condensin association with
mitotic centromeres.
Catrina Miles, Miao Yu, Jon Baxter
Genome Damage and Stability Centre. Department of Life Sciences. University of Sussex, UK.
Condensin is a multi-subunit protein complex, essential for chromosome organization, resolution
and segregation during cell proliferation. Although condensin is enriched at centromeric regions
during mitosis, the full scope of regulation of condensin recruitment remains unclear. In the budding
yeast system we have analysed the potential intrinsic and extrinsic factors required for mitotic
recruitment of the condensin complex to centromeric regions. Using ChIP to ascertain condensin
enrichment levels, we find that the localisation of condensin to centromeric regions is regulated by
the extrinsic activities of mitotic kinases and phosphatases, and the intrinsic enzymatic activity of the
condensin complex itself. The importance of these data for models of condensin recruitment and
activity will be discussed.
95
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 59
Spindle checkpoint signalling from yeast and human
kinetochores
Jonathan Millar, Virginia Silio, Maria Del Mar Mora-Santos, Andrew McAinsh
University of Warwick
The spindle assembly checkpoint (SAC) is the major surveillance system that ensures sister
chromatids do not separate until all chromosomes are correctly bi-oriented. In early mitosis
components of the SAC, including Bub1, Bub3, Mad1, Mad2 and Mad3/BubR1 proteins, are recruited
to kinetochores. This induces a conformational change in Mad2 that promotes its association to
Mad3 and Cdc20 to form the mitotic checkpoint complex (MCC). The MCC inhibits the anaphasepromoting complex (APC/C) until the checkpoint is satisfied. Activation of Cdc20-APC/C activity
triggers the destruction of securin and cyclin, which allows the dissolution of sister chromatid
cohesion and mitotic progression. We recently showed that phosphorylation of multiple conserved
MELT motifs in Spc7 (KNL1) by Mph1 (Mps1) kinase recruits Bub1 and Bub3 to the kinetochore and
that this is required to maintain the SAC signal. Conversely, association of PP1 to the N-terminus of
Spc7 is necessary to silence the SAC. However the role of the KNL1-Bub3-Bub1 (KBB) pathway in
spindle checkpoint signalling in human cells remains unclear. We have examined this issue in
immortalised diploid retinal pigment epithelial (RPE1) cells. I will present evidence that, contrary to
the situation in yeast, there are at least two distinct receptors for the Mad1-Mad2 complex at
human kinetochores and, secondly, that these respond to distinct problems in microtubulekinetochore attachment.
96
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 60
Premature sister chromatid separation is poorly detected by
the spindle assembly checkpoint due to system-level feedbacks.
Mihailo Mirkovic1, Lukas Hutter2, Bela Novak2, Raquel Oliveira1
1
2
Instituto Gulbenkian de Ciência, Oeiras, Portugal
Oxford Center for Integrative Systems Biology, Department of Biochemistry, University of Oxford, Oxford
Mitotic errors are prevented by the Spindle Assembly Checkpoint (SAC), a surveillance mechanism
that inhibits anaphase onset until all chromosomes are properly aligned and bioriented.
Biorientation, in turn, depends on sister chromatid cohesion mediated by the cohesin complex. It
should therefore be expected that the SAC arrests mitosis when loss of sister chromatid cohesion
occurs prematurely. However, evidence suggests that the SAC is not robust enough to detect and
halt cell division in the absence of cohesin. The mechanism behind this poor response is not properly
understood. To address this issue we made use of a system to acutely induce sister chromatid
separation in Drosophila developing brains. We show that full sister chromatid separation does not
elicit a robust checkpoint response and cells abnormally exit mitosis with high segregation errors
after a short mitotic delay. Quantitative live-cell imaging analysis reveals that mitotic exit in the
presence of single sisters is caused by a gradual stabilization kinetochore-microtubules interactions
and consequently weak SAC signalling. Surprisingly, most single sisters attach to the spindle in an
end-on fashion, raising the question of how such interactions are escaping the tension-sensitive
error-correction mechanisms. To elucidate this, we adopted a mathematical modelling approach to
investigate the role of multiple feedback loops in the mitotic signalling network. Our results suggest
that frail SAC activation upon cohesion loss is not solely caused by an intrinsic weak signalling
capacity, but additionally potentiated by several feedback loops that gradually impair errorcorrection efficiency, which accelerate mitotic exit in response to premature cohesin loss. Our
results explain how cohesion defects may escape SAC surveillance and therefore give rise to
aneuploid cells.
97
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 61
CDK, PP2A And CDK-dependent PP2A-inhibitory pathway Can
Make A Switch-like Response of Mitotic Phosphorylation
Satoru Mochida1, Kaori Takaki1, Takeharu Nagai2
1
2
Kumamoto University
Osaka University
During cell cycle, M phase must be temporally separated from other phases to avoid chromosome
instability. This separation is achieved by the abrupt changes of phosphorylation level of CDK
substrates. We have shown that a subset of CDK substrates was dephosphorylated by a particular
form of PP2A complex including B55 regulatory subunit (PP2A-B55). Enzymatic activity of PP2A-B55
is suppressed in M phase (thereby opposite to CDK activity) by the Greatwall kinase (Gwl)ENSA/ARPP-19 (ENSA collectively) pathway, in which CDK-activated Gwl activates ENSA. Activated
ENSA then binds and inhibits PP2A-B55. In addition, recent reports by other groups suggested that
both ENSA and Gwl are substrates of PP2A-B55, suggesting double-negative feedback loops. In the
current study, we aimed to qualitatively and quantitatively analyse these proteins as a “system” to
know if they contribute to the abrupt change of phosphorylation of CDK substrates, especially
focusing on mitotic entry. For this purpose, we developed a bioluminescent energy-transfer (BRET)
probe, which enabled us to analyse kinetics of phosphorylation/dephopshorylation of a model CDK
substrate in real time. By using this probe in our reconstituted reaction, we observed a switch-like
behavior on the phosphorylation level of CDK substrate. Together with the established roles of
Cdc25 and Wee1, we suggest that PP2A and its regulators contribute to the robustness of the cell
cycle. The reconstituted system we built would be a very useful platform to test if and how your
protein of interest affects the cell cycle.
98
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 62
Defects in C/D snoRNPs assembly cause premature nucleolar
hyper-condensation and impair mitotic exit
Ana Isabel de los Santos-Velázquez, Fernando Monje-Casas
Anabel de los Santos-Velázquez and Fernando Monje-Casas
Dept. Genetics. University of Seville / Andalusian Center for Molecular Biology and Regenerative Medicine (CABIMER)
Avda. Américo Vespucio, s/n. 41092 Seville (SPAIN)
The faithful distribution of the chromosomes during mitosis requires the compaction of the DNA.
This is especially important for the segregation of the repetitive ribosomal DNA (rDNA) array, which
organizes the nucleolus. Chromosome compaction depends on a protein complex known as
condensin. In budding yeast, condensin becomes highly enriched in the nucleolus during anaphase
to facilitate chromosome segregation, and this process requires the activity of the Cdc14
phosphatase, a key determinant of mitotic exit. The activation of Cdc14 during the early stages of
anaphase promotes condensin enrichment at the rDNA locus through the inhibition of RNA
polymerase I transcription. Our data now suggest that not only PolI transcription, but also rRNA
maturation interferes with condensin accessibility to the rDNA locus. Specifically, we show that
defects in the assembly of C/D snoRNPs, which mediate site-specific modifications of the rRNA, lead
to premature nucleolar condensation. Interestingly, this precocious hyper-condensation of the rDNA
impairs Cdc14 release from the nucleolus, which is necessary to inactivate mitotic cyclin-CDK
complexes throughout the cell and therefore to promote mitotic exit. Unlike most other DNA
sequences, the rDNA fully condenses only during anaphase. Our results, together with the fact the
same regulatory mechanism is used to couple mitotic exit with the completion of chromosome
segregation in budding yeast, also allow us to explain why it is necessary to prevent full
condensation of the rDNA until anaphase.
99
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 63
Characterization of a new APC/C subunit in Drosophila
melanogaster
Ágota Nagy1, Margit Pál2, Olga Nagy2, Levente Kovács1, 2, Péter Deák1, 2
1
2
Department of Genetics, University of Szeged, Szeged, Hungary
Biological Research Centre, Hungarian Academy of Sciences, Szeged, Hungary
The Anaphase Promoting Complex (APC/C) plays a key role in controlling the protein level of cell
cycle regulators during mitosis and G1 phase. As an E3 enzyme, this large complex determines the
specificity of the ubiquitin-dependent destruction of target proteins. The APC/C contains at least 1315 subunits, many of which were identified and characterized by biochemical and genetic means
from several model organisms. We have shown recently, that one of these subunits, Apc11, interacts
with Mr/Apc2, and they together form a binding site for Vihar, the E2-C type ubiquitin conjugating
enzyme in Drosophila.
We have identified another two proteins in Drosophila with high degree of N-terminal sequence
similarity to yeast and human Cdc26, a small subunit of APC/C. Mutant alleles of the gene coding for
one of these proteins result in pupal lethality accompanied with chromosome overcondensation and
mitotic arrest in larval neuroblasts. This phenotype is comparable to that from knockdown of known
APC/C subunits. Due to its phenotype and higher similarity to known Cdc26 subunits, we
denominate this gene as DmCdc26. Interestingly, the other gene proved to be nonessential for
development and fertility, didn’t have any mitotic phenotype but nevertheless, it could complement
the loss of function phenotype of DmCdc26, therefore it was dubbed as DmCdc26-like. These
findings suggest that the APC/C machinery in Drosophila is somewhat different from those in yeast
and human cells and may warrant further analysis. We are investigating the genetic and biochemical
relationship between these proteins and the other subunits of the APC/C.
100
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 64
Nuclear dynamics and DNA replication in fission yeast
Patroula Nathanailidou1, Anna Maria Rapsomaniki1, Nickolaos Nikiforos
Giakoumakis1, Stella Maxouri1, Stavros Taraviras2, Zoi Lygerou1
1
2
Laboratory of General Biology, School of Medicine, University of Patras, Greece
Laboratory of Physiology, School of Medicine, University of Patras, Greece
DNA replication proceeds under a defined temporal and spatial pattern of origin activation, called
replication program. Experimental data support that cells are able to adapt their replication program
under different cellular conditions. There are various determinants of origin usage during S phase
and among them nuclear structure and the spatial organization of chromatin is considered to be of
great importance.
We are interested to understand how the replication program is affected with respect to origin
positioning in the 3-D space of the nucleus. Combining structural data from 3C studies in fission
yeast and data concerning the efficiency of each origin of replication along the fission yeast genome
we observed a correlation between positioning with respect to centromeres and origin efficiency. To
investigate experimentally how the position of centromeres affects replication, we constructed a
strain carrying GFP- tagged (Mis6-GFP), unclustered (Csi1Δ) centromeres and is able to incorporate
the thymidine analogue BrdU (hsv-TK,hENT1). We assesed the BrdU incorporation pattern both in
Csi1Δ and wild type -for this locus- cells, arrested in early S phase. In wild type cells BrdU foci are
located mostly around the centromeres. Upon centromere unclustering BrdU incorporates in a
significantly more dispersed manner than in the wild type situation. This result indicates that the
position of origins relative to centromeres may affect the efficiency and the timing of their firing.
In order to make a direct correlation between the distance from centromeres in the 3-D space and
the efficiency of origins at both the population and single cell level specific origins of replication are
being tagged using the lacO-lacI system.
101
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 65
Essential role of the atypical Cdk2 activator RingoA in meiotic
telomere tethering to the nuclear envelope
Angel Nebreda
Institute for Research in Biomedicine (IRB Barcelona) and Institució Catalana de Recerca i Estudis Avançats (ICREA),
Barcelona, Spain
CDKs are protein kinases normally activated by regulatory subunits named cyclins, which are
instrumental for cell cycle regulation. Genetic analysis in mice has revealed an essential role for Cdk2
in meiosis, which renders Cdk2 knockout mice infertile. Intriguingly, mice deficient for cyclins that
can potentially activate Cdk2 block meiosis earlier (cyclin B1), later (cyclins A1 and A2) or show
weaker phenotypes (cyclins E1 and E2), suggesting that Cdk2 might be activated independently of
cyclins in meiosis. Here we show that mice deficient in RingoA, an atypical activator of Cdk1 and
Cdk2 that has no amino acid sequence homology to cyclins, are infertile and display meiotic defects
virtually identical to those observed in Cdk2 knockout mice including non-homologous chromosome
pairing, unrepaired double strand breaks and undetectable sex body. Interestingly, RingoA is
required for Cdk2 targeting to telomeres and RingoA knockout spermatocytes display severely
affected telomere tethering as well as loss of telomeric Sun1, a protein essential for the attachment
of telomeres to the nuclear envelope. Our results identify RingoA as an essential activator of Cdk2 at
meiotic telomeres, which regulates telomere tethering to the nuclear envelope and proper synapsis
of homologous chromosomes. This is the first genetic evidence of a physiological function for
mammalian Cdk2 that depends on a non-cyclin regulatory subunit.
102
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 66
The Contribution of Cell Size to Functional Decline during Aging
Gabriel Neurohr, Angelika Amon
David H. Koch Institute, Massachusetts Institute of Technology
Functional decline with increasing age has been observed in most known organisms. Genetic and
environmental modulators of aging have been identified, yet the mechanisms leading to functional
decline remain poorly understood. Conditions that reduce cell growth delay aging from yeast to
mammals. Interestingly aging in yeast and mammalian cells correlates with a marked increase in cell
size. Whether and how this large cell size itself contributes to functional decline is not clear. I used
cell division cycle mutants to generate oversized yeast cells. The resulting decreased DNA:cytoplasm
impaired cell proliferation in big cells. Increasing cell size was sufficient to reduce lifespan and
slowed cell cycle progression. These delays correlated with impaired induction of cell cycle regulated
genes and other inducible transcription systems. Importantly old yeast cells showed the same cell
cycle progression and transcription defects as big young cells, indicating that age associated cell
cycle defects are indeed a consequence of big cell size. The finding that increasing cell size impairs
cell proliferation and contributes to age associated functional decline challenges currently accepted
models of aging in budding yeast.
103
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 67
Re-visiting the regulation of centrosome duplication
Zsofia Novak, Jordan Raff
Sir William Dunn School of Pathology, University of Oxford
Centrosomes are the main microtubule organizing centres in animal cells and thereby participate in
many key cellular processes, such as cell polarity, motility or bipolar spindle assembly. In most cell
types centrosome numbers are carefully controlled, which is critical for the accurate regulation of
centrosome-dependent cellular events. Accordingly, numerical centrosome aberrations have been
linked to a number of human diseases. A key regulation point of centrosome formation is the
assembly of centrioles - the structures that act as both the physical core and the precursor of
centrosomes. Centrosomes contain a pair of centrioles, and during a cell division cycle these
undergo a single round of semiconservative replication in S-phase to ensure that centrosome
duplication occurs strictly once per cell cycle. We have recently shown in Drosophila that the
conversion of newly assembled daughter centrioles to duplication-competent mother centrioles
requires the centriolar incorporation of Asl/Cep152. The centriolar presence of Asl/Cep152 acts as a
primary license for centriole replication, and its recruitment is initiated as new centrioles pass
through their first mitosis, prior to their first round of duplication in the subsequent S-phase. The
initial incorporation of Asl at new centrioles is dependent on the core centriolar component Sas-4,
which itself is present in centrioles from the earliest stages of their assembly at S-phase onset, but
appears unable to recruit Asl until late mitosis. Here we investigate the cell cycle-dependent
regulation of the Sas-4 –Asl interaction at centrioles in vivo as the key regulation mechanism that
permanently converts newly assembled centrioles into a replication-competent form.
104
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 68
Coordinating Cdk1 and Plk1 activation for mitotic entry in early
C. elegans embryos and human cells
Yann THOMAS1, Luca CIRILLO2, Costanza PANBIANCO2, Lisa MARTINO1, Nicolas
TAVERNIER1, Lucie VAN HOVE1, SCHWAGER Françoise2, Anna Santamaria4, Monica
GOTTA2, 3, Lionel PINTARD1
1
Jacques Monod Institute, UMR7592, Paris-Diderot University - CNRS, Paris, France
Department of Cellular Physiology and Metabolism, Faculty of Medicine, University of Geneva, 1 rue Michel Servet,
Geneva 4, Switzerland
3 Swiss National Centre for Competence in Research Program Chemical Biology, Geneva 1211, Switzerland
4 Biomedical Research Unit in Gynecology, Collserola building, Vall Hebron Research Institute (VHIR) 08035 Barcelona, Spain
2
Mitosis is orchestrated by several protein kinases including cyclin-dependent kinases (Cdks), Pololike kinases (Plks) and Aurora kinases. Despite considerable progress toward understanding the
individual function of these protein kinases, how their activity is coordinated in space and time
during mitosis is less well understood. We have recently shown that CDK-1 regulates PLK-1 activity
during mitosis in C. elegans embryos through multisite phosphorylation of the PLK-1 activator SPAT1 (Tavernier et al. 2015). Early embryos expressing SPAT-1 variants mutated on CDK-1
phosphorylation sites presented severe delays in mitotic entry, mimicking embryos lacking spat-1 or
plk-1 function. We further showed that SPAT-1 phosphorylation by CDK-1 promotes its binding to
PLK-1 and stimulates PLK-1 phosphorylation on its activator T-loop by Aurora A kinase.
In human cells, Bora, the ortholog of SPAT-1, regulates several aspects of mitosis (Bruinsma et al.
2014, Chan et al 2008, Macůrek et al. 2008, Seki et al. 2008. Although phosphorylation of Bora is not
strictly required to activate Plk1 (Seki et al. 2008), we found that Cdk1 phosphorylation of Bora
greatly increases Plk1 activation in vitro (Tavernier et al. 2015). We have now mapped the Cdk1
phosphorylation sites of Bora. Based on the results obtained in C. elegans, we have mutated a
number of N-terminal sites and found that Bora ability to activate Plk1 is greatly reduced. We are
currently testing the function of these sites in human cells. Our data will help to shed light on the
activation mechanisms of Plk1 by Bora. Since Plk1 is overexpressed in many cancers, understanding
the precise mechanisms of activation of this kinase will help the design of anti-cancer drugs.
References:
• Tavernier et al. JCB 2015
• Bruinsma W. et al. J Cell Sci. 2014
• Chan EH. et al. Chromosoma. 2008
• Macůrek L. et al. Nature. 2008
• Seki A. et al. Science. 2008
105
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 69
Cycling to change destiny: dream or reality?
Fabienne PITUELLO, Frédéric BONNET, Angie MOLINA-DELGADO, Eric AGIUS
Centre de Biologie du Développement UMR5547 CNRS/UPS
Our aim is to understand how cell cycle features of neural stem/progenitor cells (NS/PC) time the
switch from proliferating progenitors towards differentiating neurons. We previously showed that a
positive regulator of the G2/M transition, the phosphatase CDC25B, is promoting neuronal
differentiation. CDC25B expression correlates remarkably well with areas where neurogenesis occurs
and downregulating the CDC25B phosphatase, results in G2-phase lengthening and defective
conversion of NSC to differentiating neurons. We thus identified a new function of the CDC25B
phosphatase in promoting neuronal differentiation associated with a G2 phase shortening (Agius et
al., 2015; Peco et al., 2012). More recently we began to elucidate how the CDC25B phosphatase
switches a proliferating neural progenitor into a differentiating neuron (unpublished data). To help
us accomplish this goal we set up a novel high resolution time-lapse imaging technique that allows
measuring the duration of each phase of the cell cycle in single neural progenitors in the chicken
developing neural tube and to track the fate of daughter cells after mitosis. Neural progenitors can
perform three mode of division: self-renewing (one progenitor gives two progenitors, P to PP),
asymmetric neurogenic (one progenitor gives a progenitor and a neuron, P to PN), symmetric
terminal neurogenic (one progenitor gives two neurons, P to NN). Using specific markers of each
division mode, we show that CDC25B promotes neurogenic divisions. Interestingly, asymmetric or
symmetric terminal neurogenic divisions will be favored depending on the context. Moreover part of
this CDC25B function is independent of its role in the cell cycle suggestive of new targets for the
phosphatase linked to the molecular network orchestrating neurogenesis. Our recent unpublished
data will be presented.
Agius, E., Bel-Vialar, S., Bonnet, F. and Pituello, F. (2015). Cell cycle and cell fate in the developing
nervous system: the role of CDC25B phosphatase. Cell Tissue Res 359, 201-13.
Peco, E., Escude, T., Agius, E., Sabado, V., Medevielle, F., Ducommun, B. and Pituello, F. (2012). The
CDC25B phosphatase shortens the G2 phase of neural progenitors and promotes efficient neuron
production. Development 139, 1095-104.
This work is supported by the "Fédération pour la Recherche sur le cerveau (FRC)" and the
"Fondation ARC pour la Rcecherche sur le Cancer".
106
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 70
Competition between microtubules and MPS1 kinase for
kinetochore binding directly regulates spindle checkpoint
signaling
Yoshitaka Hiruma1, 2, 3, Carlos Sacristan2, 3, Spyridon Pachis2, 3, Athanassios
Adamopoulos1, Timo Kuijt2, 3, Marcellus Ubbink4, Eleonore van Castelmur1, Anastassis
Perrakis1, Geert Kops2, 3
1
Division of Biochemistry, Netherlands Cancer Institute, 1066 CX, Amsterdam, The
Netherlands.
2 Molecular Cancer Research, University Medical Center
Utrecht, 3584 CG, Utrecht, The Netherlands.
3 Cancer Genomics Netherlands, University Medical Center
Utrecht, 3584 CG, Utrecht, The Netherlands.
4 4 Leiden Institute of Chemistry, Leiden University, Post Office Box 9502, 2300 RA Leiden. The
Netherlands
Cell division progresses to anaphase only after all mitotic chromosomes are connected to spindle
microtubules via their kinetochores and the spindle assembly checkpoint (SAC) is satisfied. The
mechanism by which the SAC senses kinetochore-microtubule interactions remains unknown. Here
we show that microtubules directly compete with the essential SAC kinase MPS1 for
binding to the outer kinetochore NDC80 complex. The N-terminal localization module of MPS1
directly interacts with the NDC80 complex in vitro, in a manner dependent on its phosphorylation.
That interaction is mediated by residues in the calponin-homology domain of HEC1 and can be
disrupted by addition of microtubule polymers. In cells, MPS1 binding to kinetochores or to ectopic
NDC80 complexes is prevented by end-on microtubule attachment, independent of phosphatase
activity or stripping by dynein. Competition for kinetochore binding between SAC proteins and
microtubules may thus represent a direct and evolutionary conserved
way to silence the kinetochore-derived SAC signal.
107
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 71
Biochemical Characterization of CENP-N/L complex during
Kinetochore assembly
Satyakrishna Pentakota, et al.
MaxPlanck Institue of Molecular Physiology
ABSTRACT: Accurate segregation of chromosomes during mitosis is crucial for the maintenance of
genome integrity. A detailed understanding of the mechanisms of chromosomal segregation is very
difficult owing to the molecular complexity of this process. Key to proper segregation of the
replicated genetic material is the physical attachment of the microtubule-based spindle to each
sister chromatid. This attachment is directly mediated by macromolecular structures known as
kinetochores. Kinetochores are built on the centromeres, which are the specific centers for
Kinetochore assembly. Kinetochores can be conceptualized as having an outer and inner
kinetochore. The outer kinetochore consists of the KMN network (Knl1, Mis12 and Ndc80 complex),
a 10 subunit complex known to bind to microtubules directly. The inner kinetochore comprises of 16
subunit of the constitutive centromere associated network (CCAN). The CCAN, which is organized in
sub complexes, acts as a bridge between the centromeric chromatin and the outer kinetochore
complex. Centromeric chromatin contains CENP-A, a histone H3 variant that is heavily enriched at
the centromeres that functions as an epigenetic mark for centromere identity. To date, only two
subunits of CCAN, namely CENP-C and CENP-N are known to bind to CENP-A. However, the
molecular detail of how CENP-C, CENP-N and CENP-A coordinate to bring about faithful chromosome
segregation is an ongoing study.
In this study, we demonstrate that CENP-N forms a heterodimer with CENP-L, consistent with
orthologous proteins in budding yeast. We then confirm that CENP-N/L has a preferential selectivity
for the CENP-A nucleosomes when compared with the canonical H3 nucleosomes by employing both
GST-pull down and size exclusion chromatography experiments. We produce a fragment of CENP-N
lacking the dimerization interface, and show that it binds specifically but weakly to CENP-A
nucleosomes when compared with CENP-N/L complex. However, CENP-L does not bind to either
CENP-A nucleosomes or canonical H3 nucleosomes indicating that CENP-N could be the prime
candidate binding to CENP-A nucleosomes. Our Analytical Ultra Centrifugation experiments (AUC) on
CENP-A nucleosomes in complex with CENP-N/L indicate that there are two copies of the CENP-N/L
complex that bound to each CENP-A histone within an octameric CENP-A nucleosome. Further
biochemical and structural studies with recombinant CENP-C will address the interplay between
these two proteins and the CENP-A nucleosome.
108
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 72
CHARACTERISATION OF THE PHOSPHATASE CONTROL SYSTEM
THAT PREVENTS PREMATURE MITOTIC ENTRY IN MAMMALIAN
CELLS
Nisha Peter, Priya Amin, Helfrid Hochegger
Genome Damage and Stability Centre, University of Sussex
Accurate chromosome segregation during mitosis prevents aneuploidy and cancer. The Wee1Cdc25-CDK1 feedback loop ensures cells enter mitosis in a timely and orderly manner. This signalling
system has been proposed to work as a bistable switch that is maintained by the counterbalancing
action of kinases and phosphatases. However, the phosphatases critical for this transition in
mammalian cells are yet to be identified. Cdc14 is the major phosphatase antagonising CDK in yeast.
But its eukaryotic homologue does not appear to play an essential role in mitotic control. Studies in
Drosophila and Xenopus have shown PP2A/B55 inhibition is crucial for CDK activation, and that this
is brought about by Greatwall kinase and its substrates ENSA/Arpp19. However, it remains unclear, if
PP2A/B55 inhibition by phosphorylated Ensa/ARPP is a critical event in regulating mitotic entry in
somatic cells.
To address this question, we have analysed the effects of ARPP19/ENSA depletion. We found that
ENSA depletion causes very similar phenotypes to the depletion of its upstream kinase Greatwall
causing mitotic delays and cytokinesis failure. Surprisingly, siRNA depletion of ARPP, which is
expressed at 10fold lower levels than Ensa, causes changes in cell shape and spontaneous cell death
in interphase cells, but does not affect mitosis. To better understand the dynamics of the G2/M
switch we have established a Cdk1 analogue sensitive mutation in HeLa cells that allows a wellcontrolled analysis of mitotic entry in a block/release experiment. Treatment of these cdk1as HeLa
cells that are arrested in G2 with the phosphatase inhibitor okadaic acid (OA) causes a rapid trigger
of the G2/M switch suggesting that an OA sensitive phosphatase plays a crucial role in preventing
mitotic entry. Interestingly depletion of PP2A/B55 and/or inactivation of this phosphatase with
constitutively phosphorylated ARPP19 do not overcome this G2 arrest. This suggests that another as
yet unknown phosphatase participates in this critical control step. We are currently deploying siRNA
screening in our cdk1as HeLa system to identify this additional player in the G2/M switch.
Ultimately these experiments will help understand how the balanced activity of the Cdk1 activation
switch and its counteracting phosphatases contribute to the regulation of mitotic entry.
109
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 73
Analysis of Drosophila protein interaction networks reveals
new centromeric and kinetochore components
Marcin Przewloka1, Max Convay2, Pietro Lio2, David Glover1
1
2
Department of Genetics, University of Cambridge
Computer Laboratory, University of Cambridge
We employed affinity purification – mass spectrometry (AP-MS) method to purify proteins from
Drosophila melanogaster cultured cells and syncytial embryos. Baits were chosen from the list of
proteins, which are important for the proper cell cycle progression. A very large volume of data was
accumulated, so we created a searchable database in order to efficiently analyse the results of our
AP-MS experiments. The database was subsequently equipped with custom designed modules for
the statistical analysis and visualization of protein-protein interaction networks. Preliminary analyses
using these new bioinformatics tools revealed that some novel proteins specifically co-purified with
the centromeric or kinetochore baits. We chose few candidates and analysed their localization in
cultured cells using GFP fusions or specific antibodies. Interestingly, we found that Drosophila vesicle
trafficking protein SNAP29 (or Ubisnap), an adaptor protein Homer and the uncharacterised protein
CG15107 indeed localize do centromeres, in some cases in the cell cycle-dependent manner.
Centromeric/kinetochore localization of these proteins suggests that previously unknown players
and regulatory pathways take part in the kinetochore assembly and function. This finding validates
the usefulness of our bioinformatics utility and underscores the importance of the systems biology
approach to analyse protein interaction networks. Currently we are selecting new candidates for
further studies and in parallel are trying to gain mechanistic insight into the function of newly
discovered kinetochore proteins and their role in the cell cycle regulation.
110
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 74
MEN-independent role of Cdc5 polo kinase in the Cdc14
activation during mitosis
Jose Antonio Rodriguez-Rodriguez, Ethel Queralt
Instituto de Investigaciones Biomédicas de Bellvitge (IDIBELL)
During mitosis dividing cells equally segregate their replicated chromosomes to the daughter cells.
To complete mitosis, Saccharomyces cerevisiae needs to activate the mitotic phosphatase Cdc14.
Before anaphase, Cdc14 is kept inactive in the nucleolus bound to the nucleolar protein Net1. Net1
is kept dephosphorylated by the activity of the PP2ACdc55 phosphatase during metaphase.
Separase-dependent downregulation of PP2ACdc55 at anaphase onset allows Cdk-dependent Net1
phosphorylation and Cdc14 release. A number of additional proteins contribute to Cdc14 regulation
commonly included in two pathways, FEAR (Cdcfourteen early anaphase release) and MEN (mitotic
exit network). Cdc5 polo kinase is a mitotic exit component, which best-characterized role is the
inactivation by phosphorylation of the MEN inhibitor Bfa1 in anaphase. However, the role of Cdc5
regulating the Cdc14 release it still not well defined. Here we present new data providing new
insight into the mechanism how Cdc5 contributes to the timely Cdc14 release from the nucleolus.
We showed that Cdk-dependent Net1 phosphorylation is not required for Cdc5-induced Cdc14
release from the nucleolus and that Cdc5 phosphorylate different residues to Cdk1 within Net1.
Apart of regulating MEN activity through Bfa1 phosphorylation, Cdc5 has a parallel role to MEN
during anaphase. This MEN-independent Cdc5 function in the Cdc14 activation requires activation by
Cdk1-dependent phosphorylation mostly at T242. In addition, Cdc5 requires not only previous Cdk1
activation but also active separase to promote Cdc14 release from the nucleolus.
111
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 75
Self-organization of a Rho GTPase module during polarity
establishment
Péter Rapali1, 2, Romain Mitteau1, 2, Aurelie Massoni-Laporte1, 2, Julien Meca1, 2, Sylvain
Tollis1, 2, Derek McCusker1, 2
1
2
Dynamics of Cell Growth and Division Laboratory, European Institute of Chemistry and Biology, Bordeaux, France
Institut de Biochimie et Génétique Cellulaires (IBGC), UMR 5095, Bordeaux, France
The most basic function of cells is to grow and divide, thus propagating life via the cell division cycle.
Our understanding of the regulatory mechanisms linking cell growth and the cell cycle are
surprisingly limited. In budding yeast, Cyclin dependent kinase 1 (Cdk1) triggers polarization of the
actin cytoskeleton and growth of a new cell via activation of the Rho GTPase Cdc42 in late G1 of cell
cycle. Cdc42 is a key regulator of polarized growth in diverse eukaryotes. The activity of Cdc42 relies
on its GDP and GTP nucleotide binding status where the GTP bound state is the active form. The
rapid formation of a stable pole of Cdc42 at the cell cortex may be established via feedback
mechanisms, wherein the Guanine nucleotide Exchange Factor (GEF) Cdc24 activates Cdc42. It has
been proposed that Bem1, a multi-domain scaffold protein is involved in feedback mechanisms
underlying Cdc42 regulation, but its exact role has been enigmatic. Here, using a FRET-based
spectroscopy assay of Cdc42 activity, we show that Bem1 increases the GEF-catalyzed GDP to GTP
exchange rate of Cdc42. In addition, Cla4, a Cdc42 effector kinase, phosphorylates Cdc24 GEF more
extensively in the presence of Bem1; moreover, Bem1 accelerates this otherwise slow Cla4mediated phosphorylation of Cdc24 GEF. Since phosphorylation of Cdc24 by Cla4 has been proposed
to down-regulate Cdc24 GEF activity, Bem1 may thus also be involved in a delayed negative
feedback loop that regulates Cdc42 activity. In conclusion, our results suggest that Bem1 is an active
scaffold protein that is directly involved in Cdc42 activation; however, it may also play a role in the
late attenuation of Cdc42 activation. This dual function of Bem1 may ensure the rapid establishment
and development of an active and stable Cdc42 pole during the cell cycle, while the capacity to
down-regulate GEF activity in response to Cdc42 activation suggests that the GTPase module has the
propensity to self-organize.
112
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 76
A kinetic model for an irreversible anaphase switch in human
cells
Scott Rata1, Susanne Hellmuth2, Olaf Stemmann2, Bela Novak1
1
2
Department of Biochemistry, University of Oxford, South Parks Road, Oxford, OX1 3QU, UK
Chair of Genetics, University of Bayreuth, Universitatsstraße 30, 95440, Bayreuth, Germany
The cleavage of centromeric cohesin by separase, a giant cysteine endopeptidase, is the universal
trigger of anaphase onset in eukaryotes. Separase activation is under multiple layers of control,
consisting of securin binding, auto-cleavage, and isomerisation. Based on recent findings in human
cells [1, 2], a mathematical model is presented demonstrating the irreversible activation of separase
at anaphase by the interplay of these mechanisms. During interphase, separase binds to securin and
protein-phosphatase 2A (PP2A). Securin inhibits the endopeptidase activity of separase, while PP2A
keeps four sites of the bound securin dephosphorylated. At the beginning of metaphase the
Anaphase Promoting Complex/Cyclosome (APC/C) becomes activated, which preferentially
promotes the degradation of free, phosphorylated securin. Once separase is liberated of securin
inhibition it auto-cleaves close to its PP2A binding site, thereby disrupting PP2A binding. With
intermolecular cleavage, securin-liberated separase promotes the phosphorylation of other
separase-bound securin molecules and makes them a better substrate of the APC/C. Furthermore,
securin-liberated separase could be isomerised by the prolyl-directed isomerase Pin1 into a securinresistant configuration, which can be inhibited by cyclin-B binding only. However the mutual
inhibition between separase and cyclin-B takes place at late stages of mitosis. This network shows an
intriguing similarity to the positive-feedback-promoted separase activation in budding yeast [3].
References
1) Hellmuth, S., Bottger, F., Pan, C., Mann, M., and Stemmann, O. (2014). PP2A delays APC/Cdependent degradation of separase-associated but not free securin. EMBO J. 33, 1134–1147.
2) Hellmuth et al., (2015). Human Chromosome Segregation Involves Multi-Layered Regulation of
Separase by the Peptidyl-Prolyl-Isomerase Pin1. Mol. Cell 58: 495 – 506
3) Holt LJ, Krutchinsky AN, Morgan DO. (2008). Positive feedback sharpens the anaphase switch.
Nature 454: 353 – 357
113
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 77
ROLE OF THE APC/C COMPLEX-MEDIATED DEGRADATION OF
GRK2 IN MITOSIS
Clara Reglero1, 2, Verónica Rivas1, 2, Federico Mayor1, Petronila Penela1, 2
1
2
Centro de Biología Molecular Severo Ochoa (CBMSO) - Universidad Autónoma de Madrid (UAM). Madrid. Spain
Instituto de Investigación Sanitaria La Princesa. 28006, Madrid, Spain
Proteasome-dependent degradation of key kinases represents a major regulatory mechanism in the
control of cell cycle. We have recently reported that timely degradation of GRK2, a serine/threonine
G protein-coupled receptor kinase, is part of an intrinsic pathway that ensures cell cycle progression.
Besides being a key player in the desensitization of manifold GPCR receptors, GRK2 emerges as a
signaling node due to its ability to regulate a variety of signaling proteins and cellular functions. We
have previously found that GRK2 protein levels progressively decay during G2/M as a result of the
sequential cooperation of the E3 ligases Mdm2 and APC/C. Thus, during G2 the functional
interaction of Pin1 with the CDK2/cyclinA-dependent phosphorylated GRK2 triggers its
ubiquitination by Mdm2. However, at mitosis onset GRK2 decay is promoted by the APC/C-Cdc20
complex that recognizes a D-Box in GRK2, in a spindle checkpoint-independent manner.
Impairment of this timely down-regulation of GRK2 by mutation of the D-box motif results in
significant alterations in cellular proliferation and mitosis progression. Our data have unveiled the
functional interaction of GRK2 with relevant regulatory factors involved in cytokinesis. Thus, we have
demonstrated that GRK2 is a key modulator of microtubule dynamics through the phosphorylation
of the tubulin deacetylase HDAC6. In addition, we have found that GRK2 influences the extent of
PI3K activation by a direct interaction with the regulatory subunit p85, which is involved at the
cleavage furrow. Our data reveal the presence of GRK2 in the midbody, a structure wherein HDAC6
and p85 are also found. Dynamic changes in the levels and functionality of GRK2 might help to timely
activate HDAC6 or/and PI3K in order to orchestrate microtubule dynamics and to recruit/activate
factors involved in organizing the contractile ring and setting the symmetry of the division plane. Our
results suggest that defective or sustained degradation of GRK2 during mitosis could impair
cytokinesis and contribute to polyploidy and chromosomal instability.
114
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 78
Characterization of the spindle assembly checkpoint in meiosis
Julie Rojas, Wolfgang Zachariae
Laboratory of Chromosome Biology, Max Planck Institute of Biochemistry, 82152 Martinsried, Germany
Faithfull chromosome segregation in both mitosis and meiosis requires coordinating APC/C-Cdc20
activation with correct microtubule-kinetochores attachment. In mitosis, kinetochore-microtubule
attachments are monitored by the spindle assembly checkpoint (SAC), which destabilizes incorrect
attachments and delays the cell cycle at metaphase until every chromosome is properly bi-oriented
(Musacchio and Salmon, 2007). Several studies performed in different model organisms suggest that
the SAC is also present in meiosis (Sun and Kim, 2012). However, SAC function and regulation in
meiosis are poorly understood. Thus, we study SAC regulation during meiosis in budding yeast. We
develop a technique to dynamically assess SAC activity in living cells using time-lapse microscopy.
Our results show that the SAC is transiently activated in both metaphase I and metaphase II. We also
observed that the SAC remains active longer in several mutants having tension or attachment
defects. Current work aims to identify aspects of SAC regulation, which are specific to either the first
or the second division of meiosis.
Musacchio, A., and Salmon, E.D. (2007). Nat Rev Mol Cell Biol 8, 379-393.
Sun, S.C., and Kim, N.H. (2012). Hum Reprod Update 18, 60-72.
115
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 79
Opening the cohesin ring:
Asymmetric ATP hydrolysis triggers DNA release
Ahmed Elbatsh, Judith Haarhuis, Benjamin Rowland
The Netherlands Cancer Institute
Sister chromatid cohesion mediated by the cohesin complex is essential for faithful chromosome
segregation in mitosis. By resisting the pulling forces of microtubules up to the moment that all
chromosomes are correctly attached, cohesin ensures that the sister chromatids separate to the
opposite poles of the cell, and that each of the daughter cells receives an equal karyotype.
Cohesin stably holds the sister chromatids together by co-entrapping them inside its ring-shaped
structure from the moment of cohesion establishment in S phase until mitosis. Cohesin can only
stably associate with DNA when it is protected against its cellular antagonist Wapl. This factor is
thought to remove cohesin from DNA through the opening up of a DNA exit gate in cohesin rings.
How Wapl achieves this ring opening is unknown. In S phase, Eco1 acetylates cohesin’s Smc3 subunit
and hereby renders cohesin resistant to Wapl.
We used yeast genetics to dissect how Wapl drives cohesin from chromatin. Hereby we identified
mutants of cohesin that can entrap, but can’t release DNA. We pinpointed an unexpected role of the
highly conserved ABC-like ATPase domain of cohesin’s Smc1 subunit. Using a combination of
genetics, microscopy, and in vitro assays, we found that hydrolysis of specifically one of cohesin’s
associated ATPs is the trigger for DNA release from cohesin rings. We propose that Eco1 locks
cohesin rings around the sister chromatids by counteracting cohesin-mediated ATPase activity.
116
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 80
Bub3-associated BubR1 sequesters Fizzy/Cdc20 at DNA breaks
and promotes correct segregation of broken chromosomes
Anne Royou
CNRS, Université de Bordeaux, IECB/IBGC, UMR 5095, 2 rue Robert Escarpit, 33607 Pessac, France
The presence of DNA double-strand breaks during mitosis is particularly challenging for the cell as it
produces broken chromosomes lacking a centromere. This situation can cause genomic instability
due to improper segregation of the broken fragments into daughter cells. We have recently
uncovered a process by which broken chromosomes are faithfully transmitted, via the BubR1dependent tethering of the two broken chromosome ends. However, the mechanisms underlying
BubR1 recruitment and function on broken chromosomes were not determined. Here, we show that
BubR1 requires interaction with Bub3 to localize on the broken chromosome fragments and to
mediate their proper segregation. We also find that Cdc20, a co-factor of the E3 ubiquitin ligase
Anaphase-Promoting-Complex/Cyclosome (APC/C), accumulates on DNA breaks in a BubR1 KEN boxdependent manner. A biosensor for APC/C activity demonstrates a BubR1-dependent local inhibition
of APC/C around the segregating broken chromosome. Therefore, we propose that the Bub3/BubR1
complex on DNA breaks functions to inhibit the APC/C locally via the sequestration of Cdc20, thus
promoting proper transmission of broken chromosomes.
117
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 81
Hi-C analysis of condensin action in budding yeast
Stephanie Schalbetter1, Jon-Matthew Belton2, Job Dekker2, Jon Baxter1
1
2
GDSC, University of Sussex, UK
University of Massachusetts Medical School, United States
Mitotic chromosome condensation is thought to occur through hierarchical folding of the chromatin
polymer in mitosis. The activity of condensin and topoisomerase II are required for full compaction
of mitotic chromosomes. Here we assess the function of condensin and topoisomerase II in mitotic
chromosome compaction by Hi-C analysis in the yeast Saccharomyces cerevisiae. We find that both
condensin and topoisomerase II act throughout the yeast genome to resolve yeast mitotic
chromosomes from one another and prevent long-range intra-chromosomal interactions during
mitosis. However, condensin and topoisomerase II achieve this in distinct manners. Condensin
promotes short-range interactions within chromosomes that prevent long-range interactions, whilst
topoisomerase II is required to resolve long-range interactions. The impact of these findings on the
current models of condensin and topoisomerase II action in mitotic chromosomes will be discussed.
118
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 82
A MEIOTIC CHECKPOINT REGULATES TRANSLATION OF THE
SIGNALING FACTOR GURKEN IN DROSOPHILA OOGENESIS.
Trudi Schupbach1, Wei Li1, Attilio Pane2
1
2
Princeton University
University of Rio de Janeiro, Brazil
The spatial patterning of the egg and embryo of Drosophila depends on signaling events that occur
during oogenesis. In particular the gene gurken (grk) which encodes a secreted molecule with
homologies to TGF-alpha like growth factors, plays an important role in transmitting pattern
information from the oocyte to the overlying follicle cells. The levels of Grk are particularly crucial in
dorso-ventral patterning, and the production of Grk protein is tightly controlled both at the level of
RNA localization, as well as at the level of protein accumulation. We have shown that mutations in
genes required for DSB repair in meiosis affect dorso-ventral patterning through activation of a
checkpoint that couples progression through meiosis to the translation of Gurken. We have also
shown that mutations in genes necessary for the production of piRNAs activate the same checkpoint
and, among other effects, lead to a decrease in Gurken protein accumulation. The molecular level at
which this meiotic checkpoint affects the translational machinery in oogenesis is under investigation.
119
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 83
The dynamics of Cyclin A2 in the cell cycle: A time and a place
for Cyclin A2
Helena Silva Cascales, Erik Müllers, Arne Lindqvist
Karolinska Institutet
Progression through the cell cycle is a highly regulated process driven by the coordinated action of
many enzymes, in particular Cyclin-Cdk complexes. Using a novel method based on accurate
quantification of immunofluorescence, we have recently shown that Cdk1 activity starts to
accumulate at completion of S-phase. Interestingly, in parallel with Cdk1 activation we observe a
change in the localisation of Cyclin A2 from being only nuclear to being both nuclear and
cytoplasmic, suggesting a role for Cyclin A2 in the cytoplasm during G2 phase. We find that all cells
express cytoplasmic Cyclin A2 in the last hours preceding mitosis. Preliminary results show that
perturbations of Cdk activity affect the dynamics of Cyclin A2 localisation to the cytoplasm; however
how Cdk activity modulates this process is still an open question. Interestingly, inflicting DNA
damage induces a relocalisation of cytoplasmic Cyclin A2 to the nucleus suggesting that the activities
needed for a DNA damage response could prevent cytoplasmic localisation of Cyclin A2.
We are currently assessing the functional importance of cytoplasmic Cyclin A2 for progression
through G2 and mitosis. However our current hypothesis is that the G2 biochemical state brought
about by the increase in Cdk activity at the S/G2 transition triggers an unknown mechanism that
induces cytoplasmic localisation of Cyclin A2 to allow the phosphorylation of Cdk targets in the
cytoplasm needed for correct mitotic entry.
120
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 84
Regulatory mechanism and physiological meaning of anaphase
delay during mouse pronuclear formation
Shou Soeda1, 2, Miho Ohsugi1, 3
1
Department of Biological Sciences, Graduate School of Sciences, The University of Tokyo
Japan Society for the Promotion of Science, Research Fellowship for young Scientists
3 Department of Life Sciences, Graduate School of Arts and Sciences, The University of Tokyo
2
Mammalian matured oocytes are arrested at metaphaseⅡ. After the fusion with sperm, oocytes
resume meiosis, leading to female chromosome segregation, polar body emission and pronuclear
formation. During this process, it takes more than one hour for female chromosomes to be
enveloped after chromosome segregation ends. This is in contrast to somatic cells in which nuclear
envelope reassembly begins just after chromosome segregation has completed. Here, we investigate
the regulatory mechanisms and the physiological importance of this anaphase delay during mouse
pronuclear formation.
We previously reported that p90RSK, which is a downstream kinase for MOS-MAPK pathway in
oocytes, has an activity to delay anaphase during pronuclear formation. However our previous
results also demonstrated that pronuclear formation preceded the inactivation of MOS-MAPKp90RSK pathway, indicating that the other factors than p90RSK determine the timing of pronuclear
formation. We speculated that the factors that counteract MOS-MAPK-p90RSK pathway are the
potent candidates for pronuclear formation accelerators and thus focused on mitotic phosphatases.
PP1 and PP2A are reported to facilitate mammalian mitotic exit by counteracting the activities of
CDK or other mitotic kinases. We tested the involvement of these phosphatases in temporal
regulation of pronuclear formation. Pronuclear formation timing was delayed by okadaic acid in a
dose dependent manner, suggesting that pronuclear formation timing is determined by the degree
of these phosphatase activities. In agreement with this hypothesis, PP1 over-expression accelerated
the timing of pronuclear formation. To address the question of how phosphatase activation delay is
achieved, we investigated the CDK-Mastl-Ensa/Arpp19 pathway that inhibits PP2A activity during
metaphase in somatic cells. By immunorfluorescent studies we found that phopsho-Ensa was
maintained for more than one hour after the onset of anaphase II, but drastically decreased before
pronuclear formation. Next, we tested the cross talk between p90RSK pathway and Ensa/Arpp19
pathway. The p90RSK inhibitor BI-D1870 treatment accelerated the decrease of phospho-Ensa as
well as pronuclear formation. Importantly, even in this condition, a decrease of phospho-Ensa
preceded pronuclear formation. These results suggest that phospho-Ensa decrease is a prerequisite
for pronuclear formation and that p90RSK is involved in upstream of Ensa/Arpp19 pathway to delay
PP2A activation.
We next investigated the physiological importance of delayed pronuclear formation by accelerating
the pronuclear formation timing by two ways; PP1 over expression and intra-cytoplasmic sperm
injection one hour after parthenogenetic activation (PA-ICSI). We found that PP1 over expressed
zygotes formed aberrant small male pronucleus and caused severe chromosome bridges in first
cleavage division stage while PP1 over expression didn’t affect pronuclear size in parthenogenetic
embryos with two pronuclei and had much less effect on first cleavage division. The similar
phenotype was observed in the zygotes obtained by PA-ICSI. Thus we propose that anaphase delay
in pronuclear formation is required for sperm chromatin to be reorganized into proper zygotic
chromatin.
Sperm chromatin remodeling and female chromosome segregation are fulfilled in the same
cytoplasm. Our findings revealed that oocytes accomplish these complicated processes by adjusting
the cell cycle.
121
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 85
Nek7-interactor RGS2 is required for mitotic spindle
organization
Edmarcia Elisa de Souza1, Heidi Hehnly3, Arina Marina Perez1, Gabriela Vaz Meirelles1,
Juliana Smetana1, Stephen Doxsey3, Jörg Kobarg2
1
Brazilian Laboratory of Biosciences/National Center For Research in Energy and Materials, Campinas, SP, Brazil
Institute of Biology, State University of Campinas, Campinas, SP, Brazil.
3 University of Massachusetts Medical School, Program in Molecular Medicine, MA, USA.
2
The mitotic spindle apparatus is composed of microtubule (MT) networks attached to kinetochores
organized from two centrosomes (a.k.a. spindle poles). In addition to this central spindle apparatus,
astral MTs assemble at the mitotic spindle pole and attach to the cell cortex to ensure appropriate
spindle orientation. We propose that cell cycle-related kinase, Nek7, and its novel interacting protein
RGS2, are involved in mitosis regulation and spindle formation. We found that RGS2 localizes to the
mitotic spindle in a Nek7-dependent manner, and along with Nek7 contributes to spindle
morphology and mitotic spindle pole integrity. RGS2-depletion leads to a mitotic-delay and severe
defects in the chromosomes alignment and congression. Importantly, RGS2 or Nek7 depletion or
even overexpression of wild-type or kinase-dead Nek7, reduced γ-tubulin from the mitotic spindle
poles. In addition to causing a mitotic delay, RGS2 depletion induced mitotic spindle misorientation
coinciding with astral MT-reduction. We propose that these phenotypes directly contribute to a
failure in mitotic spindle alignment to the substratum. In conclusion, we suggest a molecular
mechanism whereupon Nek7 and RGS2 may act cooperatively to ensure proper mitotic spindle
organization.
122
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 86
Dual regulation of Drosophila APC/C activity by Rca1
Frank Sprenger, Matthias Kies
University of Regensburg
APC/C activity has to be tightly controlled to ensure accumulation of mitotic cyclins in G2. Negative
regulation of interphase APC/C activity is achieved by multiple mechanisms: 1) by phosphorylation
(and thereby inhibition) of its activator Fzr/Cdh1 by cyclin-dependent kinase activity, and 2) by
pseudosubstrate inhibition by Rca1/Emi1. We have analyzed the functional domains in Rca1 in
Drosophila Schneider cells and could confirm that the mechanisms proposed for APC/C inhibition by
the vertebrate homologue Emi1 are also valid for Rca1 in Drosophila. Further analyses revealed that
the F-box domain of Rca1 is also required for efficient APC/C inhibition. Several lines of evidence
indicate that Rca1 is part of an SCF complex that can target other proteins for ubiquitination and
subsequent degradation and that this function is indespensible for full APC/C inhibition. Dap, a
specific CKI inhibitor of CycE/Cdk2, could be identified as a potential target of SCF-Rca1. This
suggests that Rca1 contributes to APC/C inhibition in a dual way: as a pseudosubstrate inhibitor, and
as part of an SCF-complex, degrading a CKI that would otherwise restrain kinase activity necessary
for APC/C inhibition.
123
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 87
Ubiquitin receptor protein UBASH3B mediates a switch-like
mechanism of Aurora B localization to microtubules.
Ksenia Krupina1, Charlotte Kleiss1, Stephane Schmucker1, Kay Hofmann2, Olivier
Poch3, Laurent Brino1, Izabela Sumara1
1
Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC), Illkirch, France.
Institute for Genetics, University of Cologne, Cologne, Germany.
3 Computer Science Department, ICube, UMR 7357, University of Strasbourg, CNRS, Fédération de médecine
translationnelle, Strasbourg, France.
2
Mitosis ensures equal segregation of the genome and is controlled by variety of ubiquitylation
signals on substrate proteins. However, it remains unexplored how the versatile ubiquitin code can
be read during mitotic progression. Here, we identify the ubiquitin receptor protein UBASH3B as an
important regulator for mitosis. UBASH3B interacts with ubiquitylated Aurora B, one of the main
kinases regulating chromosome segregation, and controls its subcellular localization but not protein
levels. Importantly, UBASH3B is a limiting factor in this pathway, and is sufficient to target Aurora B
to microtubules prior to anaphase, as revealed by super resolution imaging. Moreover, targeting
Aurora B to microtubules by UBASH3B is necessary for the timing and fidelity of chromosome
segregation in human cells. Our findings uncover an important mechanism how ubiquitin
attachment to a substrate protein is decoded during mitosis.
124
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 88
The inner centromere-shugoshin network prevents
chromosomal instability
Yuji Tanno, Hiroaki Susumu, Miyuki Kawamura, Yoshinori Watanabe
Laboratory of Chromosome Dynamics, Institute of Molecular and Cellular Biosciences, University of Tokyo
Chromosomal instability (CIN) is a major trait of cancer cells and a potent driver of tumor
progression. However, the molecular mechanisms underlying CIN still remain elusive. Here, we show
that a number of CIN+ cell lines exhibits impaired centromeric localization of SGO1 and Aurora B,
which are the core component of the inner centromere-shugoshin (ICS) network. These defects are
linked with the hyperstabilization of kinetochore-microtubule attachment, which is a major cause of
chromosome segregation error. The defects in ICS network are caused mostly by the loss of histone
H3 lysine-9 tri-methylation at centromeres, and sometimes by a reduction in chromatin-associated
cohesin; both pathways separately sustain centromeric SGO1 stability. Remarkably, artificial
restoration of the ICS network suppresses chromosome segregation errors in a wide range of CIN+
cells. Thus, dysfunction of the ICS network might be a key mechanism underlying CIN in human
tumorigenesis.
125
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 89
Polo-like kinase activation through SPAT-1/Bora
phosphorylation in human cells and in C. elegans embryos
Yann Thomas1, Luca Cirillo2, Costanza Panbianco2, Lisa Martino1, Nicolas Tavernier1,
Lucie Van Hove1, Francoise Schwager2, Thibault Leger3, Anna Santamaria5, Monica
Gotta2, 4, Lionel Pintard1
1
Jacques Monod Institute, UMR7592, Paris-Diderot University - CNRS, Paris, France
Department of Cellular Physiology and Metabolism, Faculty of Medicine, University of Geneva, 1 rue Michel Servet,
Geneva 4, Switzerland.
3 Mass Spectrometry Facility, Jacques Monod Institute, UMR7592, Paris-Diderot University - CNRS, Paris, France
4 Swiss National Centre for Competence in Research Program Chemical Biology, Geneva 1211, Switzerland
5 Biomedical Research Unit in Gynecology, Collserola building, Vall Hebron Research Institute (VHIR) 08035 Barcelona, Spain
2
Mitosis is orchestrated by several protein kinases including cyclin-dependent kinases (Cdks), Pololike kinases (Plks) and Aurora kinases. Despite considerable progress toward understanding the
individual function of these protein kinases, how their activity is coordinated in space and time
during mitosis is less well understood. We have recently shown that CDK-1 regulates PLK-1 activity
during mitosis in C. elegans embryos through multisite phosphorylation of the PLK-1 activator SPAT1 (Tavernier et al. 2015). Early embryos expressing SPAT-1 variants mutated on CDK-1
phosphorylation sites presented severe delays in mitotic entry, mimicking embryos lacking spat-1 or
plk-1 function. We further showed that SPAT-1 phosphorylation by CDK-1 promotes its binding to
PLK-1 and stimulates PLK-1 phosphorylation on its activator T-loop by Aurora A kinase.
In human cells, Bora, the ortholog of SPAT-1, regulates several aspects of mitosis (Bruinsma et al.
2014, Chan et al 2008, Macůrek et al. 2008, Seki et al. 2008. Although phosphorylation of Bora is not
strictly required to activate Plk1 (Seki et al. 2008, Tavernier et al. 2015), we found that Cdk1
phosphorylation of Bora greatly increases Plk1 activation in vitro. We have now mapped the Cdk1
phosphorylation sites of Bora. Based on the results obtained in C. elegans, we have mutated a
number of N-terminal sites and found that Bora ability to activate Plk1 is greatly reduced. We are
currently testing the function of these sites in human cells. Our data will help to shed light on the
activation mechanisms of Plk1 by Bora. Since Plk1 is overexpressed in many cancers, understanding
the precise mechanisms of activation of this kinase will help the design of anti-cancer drugs.
References:
• Tavernier et al. JCB 2015
• Bruinsma W. et al. J Cell Sci. 2014
• Chan EH. et al. Chromosoma. 2008
• Macůrek L. et al. Nature. 2008
• Seki A. et al. Science. 2008
126
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 90
The budding yeast cell cycle without Cdc14 phosphatase
Sandra Touati, Frank Uhlmann
The Francis Crick Institute
Cell cycle progression is driven by a succession of phosphorylation and dephosphorylation events,
mediated by the interplay of kinases and phosphatases. Correct balance of kinase/phosphatase
activity is essential to order the cell cycle events. In G1/S-phase, low CDK (cyclin dependent kinases)
activity permits accurate DNA replication, whereas high Cdk activity in G2/M is crucial for mitotic
entry. The metaphase-to-anaphase transition is a key event of the cell cycle where
kinase/phosphatase ratio is gradually reversed. Indeed, mitotic cyclins are degraded, CDK activity is
inhibited and phosphatases dephosphorylate CDK substrates.
In budding yeast, the main mitotic phosphatase is Cdc14. In anaphase, Cdc14 is released from the
nucleolus into the nucleus and cytoplasm and counteracts CDK activity. Substrate dephosphorylation
allows anaphase regulation, cytokinesis and mitotic exit. Accordingly, cdc14-1 cells are arrested in
anaphase. Glc7 (PP1) and PP2A are also indispensable for cell cycle progression. Indeed, glc7-ts
mutants lead to lethality and the cell cycle is highly perturbed upon deletion of either catalytic
subunit of PP2A or deletion of the regulatory subunits Cdc55 (B55) and Rts1 (B56).
In mammalian cells, Cdc14 is dispensable for mitotic exit and PP2A-B55 seems to be the main
phosphatase driving CDK substrate dephosphorylation. In budding yeast, the predominance of Cdc14
could hide fundamental roles for Glc7 and PP2A. Accordingly, we propose to study their roles and
substrates after metaphase in a context where Cdc14 is absent. We also intend to investigate if the
mitotic arrest observed in Cdc14 depleted cells may be overcome by the modulation of the Glc7,
PP2A and expression of its regulatory subunits.
127
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 91
Cytokinetic abscission is an acute G1 event
Amit Tzur1, Ofir Gershony2, Tal Pe'er1, Meirav Noach-Hirsh1, Natalie Elia2
1
The Mina and Everard Goodman Faculty of Life Sciences and the Institute of Nanotechnology and Advanced Materials,
Bar-Ilan University, Ramat-Gan 5290002, Israel
2 Department of Life Sciences and the National Institute for Biotechnology in the Negev, Ben-Gurion University, Beer-Sheva,
8410501, Israel
Animal cell division ends with the cutting of the microtubule and membrane intercellular bridge
connecting the two daughter cells. This process, known as cytokinetic abscission (abscission), is
widely regarded as the last step of cytokinesis, i.e., the last step of the cell cycle. Major
breakthroughs have been recently achieved, illuminating mechanistic aspects of abscission;
however, the timing of abscission with respect to the mammalian cell cycle remains unclear. In this
study, we carefully measured the onset and progression of abscission in dividing cells expressing a
G1 reporter. We conclude that abscission commences long after cells enter the G1 phase. Affiliating
abscission with G1 is beyond semantics since it essentially postulates that the last step of the cell
cycle is regulated in, and probably by, the following cycle.
128
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 92
CDK-1 inactivation and PLK-1 gradient cooperate for the
sequential assembly of the Nuclear Envelope during mitotic
exit.
Paola Vagnarelli, Maria Laura Di Giacinto, Ines Castro, Lorena Ligammari
Brunel University London
Nuclear Envelope (NE) reformation around the segregating chromatin after mitosis is a very
controlled process regulated in both space and time.
As with many mitotic exit events it is triggered by inactivation of CDK1/cyclinB and by the activation
or re-localisation of several protein phosphatases. Thus far PP1 and PP2A have been both implicated
in the de-phosphorylation events necessary for NE reorganisation.
Moreover, spatial clues contribute to the coordination of the reassembly process. This spatial
information is provided by both the chromatin and a gradient of kinases. Recent work has identified
Aurora B kinase as a negative regulator of NE reassembly but little is known about the role of other
mitotic kinases that demonstrate a gradient distribution within the anaphase cells.
Lagging chromosomes incapable of re-joining the main nucleus after mitosis lead to the formation of
micronuclei (MN). These chromatin structures present defects in Nuclear Envelope assembly and
provide a great tool to study the role of mitotic kinases enriched at the spindle midzone in the
formation of NE. Furthermore, MN undergo rupture during the cell cycle and represent a major
source of genomic instability.
Using human cell lines and following the destiny and the Nuclear Lamina composition of missegregating chromosomes, we have identified PLK-1 as the kinase that negatively regulates the
assembly of Nuclear Pores (but not the Lamina). Moreover we show that LaminA reassembly can be
triggered solely by the inactivation of CDK-1 independently of any other spatial clue.
We therefore propose that both a clock (CyclinB degradation) and a gradient (PLK, AuroraB)
contribute to the timely assembly of the Nuclear Envelope after mitosis.
129
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 93
The phosphatase Cdc14 is essential for anaphase spindle
elongation
Rosella Visintin, Michela Roccuzzo, Clara Visintin
European Institute of Oncology
Sister chromatid separation requires the dissolution of the cohesin linkages that hold them together,
whilst their segregation relies on the activity of the mitotic spindle and spindle-associated proteins.
Cleavage of cohesins and CDK inhibition are thought to be sufficient for triggering chromosome
segregation. Here we identified a novel essential requirement for anaphase chromosome movement
following cohesin cleavage. We showed that, at anaphase onset, the phosphatase Cdc14 and the
polo-like kinase Cdc5 are redundantly required to drive spindle elongation. This essential role of
Cdc14 is mediated by the FEAR network, a group of proteins that activates Cdc14 at anaphase onset
and we suggest that Cdc5 facilitates both Cdc14 activation and CDK inhibition. We further identify
the kinesin-5 motor protein Cin8 as a key target of Cdc14. Indeed, of all the known spindleassociated Cdc14 substrates tested, only a non-phosphorylatable allele of Cin8 is capable of
bypassing the cdc14cdc5 double mutant arrest. This result suggests that although stabilization of
microtubule dynamics and building a stable midzone are an important requirement for anaphase
spindle elongation the limiting step for chromosome segregation is the activation of motor proteins
of which Cin8 is an example.
130
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 94
Regulation of origin selection is critical for genome integrity
Blanca Gomez-Escoda, Pei-Yun Wu
Institute of Genetics and Development of Rennes - CNRS
The accurate duplication and transmission of genetic information is critical for cell growth and
proliferation as well as for development and differentiation. This is ensured in part by multiple layers
of regulation of DNA synthesis and checkpoint pathways. One of the key aspects of this process is
the selection and activation of the sites of replication initiation, or origins, across the genome.
Interestingly, changes in origin usage have been observed during development and in different
pathologies, but the physiological consequences of undergoing S phase with specific replication
programs remain largely unexplored. To address this question, we have investigated the effect of
alterations in replication initiation induced when checkpoint-defective fission yeast cells encounter
replication stress, a situation analogous to that found in cancer. In these conditions, we observe
major increases in replication initiation clustered in discrete genomic domains. Strikingly, this origin
deregulation leads to the generation of single-stranded DNA and ultimately DNA damage specifically
at these loci, constituting a major source of genome instability in these cells. Our findings provide
evidence that the organization of DNA replication along the chromosomes plays a critical role in
genome maintenance and integrity.
131
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 95
Analysis of prematurely condensed chromosomes in chicken
DT40 cells
Tao Zhang1, Paul Kalitsis1, 2, Damien Hudson1, 2
1
2
Murdoch Childrens Research Institute, Royal Children’s Hospital, Parkville, Melbourne, Victoria 3052, Australia.
Department of Paediatrics, University of Melbourne, Parkville, Melbourne, Victoria 3052, Australia.
There are two types of condensins in high eukaryotic cells, which play important roles in
chromosome condensation. Condensin I is excluded from the nucleus until prometaphase when
nuclear membrane is broken down (NEBD); while condensin II remains in the nucleus throughout the
whole cell cycle. There is little information on what role, if any, condensins play at the end of
anaphase, during which chromosomes need to be decondensed and condensins to be redistributed
into their cellular compartments. In our study in chicken DT40 cells, we focused on the transition
between anaphase and interphase and in particular what factors are involved in premature
chromosome condensation (PCC). Using quantitative live cell imaging in conjunction with 3D
reconstruction, we spatially and temporally dissected the role of condensins in G1 synchronized and
PCC cells. We found that shortly upon the completion of anaphase, chromatin bound CAP-H
(condensin I) signal dramatically decreased from the nucleus. By contrast, CAP-D3 (condensin II)
signal remained and gradually increases in interphase. Surprisingly, PCC led to recruitment of both
condensin I and condensin II into G1 premature condensed chromatins without NEBD. However, the
characteristic axial staining of the scaffold proteins is lost in G1 PCC chromosomes, and structurally
they exhibit heightened fragility. PCC replicated chromosomes show mislocalization of the
chromosome scaffold protein KIF4A but not TOPOII and SMC2. Our results show that G1 premature
condensed chromosomes require both condensin I and condensin II, but not KIF4A.
132
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Poster presentations
Poster 96
Fly-FUCCI - a versatile tool for studying cell proliferation in
complex tissues
Norman Zielke, et al.
Deutsches Krebsforschungszentrum (DKFZ) - Zentrum für Molekulare Biologie der Universität Heidelberg (ZMBH) Allianz, Im
Neuenheimer Feld 282, 69120 Heidelberg, Germany
The development of organs and tissues often involves strictly orchestrated lineages, in which only
certain cell-types proliferate at a time. The decision to proliferate is highly dependent on signals
from surrounding cells and hence one of the future challenges is to study cell proliferation within its
microenvironment. The recently introduced FUCCI system (Fluorescent Ubiquitination-based Cell
Cycle Indicator) allows the monitoring of cell cycle phasing in living cells. To enable the specific
labeling of small subpopulations of cells we have generated a fly-specific FUCCI system (Fly-FUCCI),
whose expression can be spatially and temporally controlled. The Fly-FUCCI system is based on E2F1
and Cyclin B, which are sequentially degraded by the E3-ligases CRL4-Cdt2 and APC/C. Simultaneous
expression of both Fly-FUCCI probes allows a distinction of all categories of interphase cells: G1 cells
are marked by GFP/CFP; cells in S phase are labeled by RFP/YFP and cells in G2 express both
markers. To support a broad range of experimental settings we have generated a toolkit of fly lines
expressing the Fly-FUCCI probes under control of UASt, UASp and QUAS promoters. We demonstrate
that the Fly-FUCCI system is capable of recapitulating the developmentally programmed cell cycle
patterns in eye and wing discs. Furthermore, we have applied the Fly-FUCCI method to the stem cell
lineage of the adult midgut, which revealed that the terminally differentiated enterocytes re-enter
the cycle during regeneration. Altogether, our work demonstrates that the Fly-FUCCI system is a
valuable tool for visualizing cell cycle activity during development and tissue homeostasis.
133
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
List of contributors
ALPHABETICAL LIST OF CONTRIBUTORS
Adamopoulos, Athanassios,
107
Adriaans, Ingrid, 38
AGIUS, Eric, 106
Akopyan, Karen, 86
Aldea, Marti, 4
Alvarez-Fernández, Mónica, 31
Alves-Rodrigues, Isabel, 41
Amin, Priya, 109
Amon, Angelika, 29, 103
Anger, Martin, 12
Araujo, Ana Rita, 39
Argüello-Miranda, Orlando, 40
Ayte, Jose, 41
Bade, Debora, 42
Bakal, Chris, 43
Bánhegyi, Gábor, 75
Barr, Alexis, 43
Barr, Francis, 34
Barral, Yves, 35, 36, 55, 83
Baxter, Jon, 95, 118
Bayer, Mathias, 55
Bayrak, Sibel, 94
Beaven, Robin, 44
Belton, Jon-Matthew, 118
Benaud, Christelle, 45
Biedermann, Sascha, 19
Binarová, Pavla, 79
Blaukopf, Claudia, 28
Blayney, Martyn, 10
Blythe, Shelby, 1
Boens, Shannah, 56
Bogre, Laszlo, 5, 69
Bohlen, Jonathan, 20
Bollen, Mathieu, 46, 56
Bolognesi, Alessio, 36
Bonaiuti, Paolo, 49
Bong, Seoung Min, 78
Bonner, Amanda, 47
BONNET, Frédéric, 106
Borsos, Mate, 13
Brandeis, Michael, 48
Brino, Laurent, 124
Brown, Nicholas, 81
Bru, Samuel, 66
Buellens, Monique, 46
Buffin, Eulalie, 11
Burgoyne, Robert D, 65
Burkhardt, Sabrina, 13
Cadart, Clotilde, 2
Castro, Anna, 33, 89
Castro, Ines, 129
Chiroli, Elena, 49
Ciliberto, Andrea, 49
CIRILLO, Luca, 105
Cladière, Damien, 11
Clotet, Josep, 66
Cohen, Paula, 13
Collart, Clara, 17
Convay, Max, 110
Coudreuse, Damien, 50
Csikasz-Nagy, Attila, 4
Cundell, Michael, 34
Cuylen, Sara, 28
Davey, Norman, 67
de Boer, Rudolf, 84
de los Santos-Velázquez, Ana
Isabel, 99
De Munter, Sofie, 46
de Souza, Edmarcia Elisa, 122
de Vries, Elisabeth, 84
Deák, Péter, 82, 100
de-Carvalho, Jorge, 25
Dekker, Job, 118
Delgado-Barea, María, 51
Deli, Márta, 90
Deneke, Victoria, 22
Denoth-Lippuner, Annina, 83
Deshpande, Ojas, 25
Desvoyes, Bénédicte, 51
Di Giacinto, Maria Laura, 129
Di Talia, Stefano, 1, 22
Dick, Amalie, 52
Ditte, Peter, 53
Dörner, Peter, 19
Doxsey, Stephen, 122
Dozier, Christine, 92
Ducommun, Bernard, 88
Echard, Arnaud, 54
Edgar, Bruce, 20
Eichele, Gregor, 53
El Yakoubi, Warif, 11
Elbatsh, Ahmed, 116
Elder, Kay, 10
Elia, Natalie, 128
Ellenberg, Jan, 28
Endicott, Jane, 81
Farcas, Ana-Maria, 55
Fardilha, Margarida, 56
Ferreira, Monica, 56
Foltman, Magdalena, 57
Françoise, SCHWAGER, 105
Froment, Carine, 92
Frongia, Céline, 88
134
Gallego, Carme, 4
Galli, Matilde, 59
Gérard, Claude, 50
Gerlich, Daniel W., 28, 52
Gershony, Ofir, 128
Giakoumakis, Nickolaos
Nikiforos, 101
GIET, Régis, 58
Girard, Juliet, 32
Glover, David, 110
Godfrey, Molly, 60
Godwin, Jonathan, 13
Gomes, Aurélie, 88
Gomez-Escoda, Blanca, 131
Gotta, Monica, 126
Groenewold, Vincent, 42
Gross, Fridolin, 49
Großhans, Jörg, 61
Gruneberg, Ulrike, 30, 63, 91
Guerra-Moreno, Angel, 41
Gueydon, Elisabeth, 14
Gutierrez, Crisanto, 18, 51
Haarhuis, Judith, 116
Haering, Christian, 62
Halada, Petr, 79
Harashima, Hirofumi, 19
Hawley, R. Scott, 47
Haynes, Lee, 65
Hayward, Daniel, 63
Hehnly, Heidi, 122
Heim, Andreas, 64
Helassa, Nordine, 65
Hellmuth, Susanne, 113
Hengeveld, Rutger, 84
Henriques, Rossana, 5
Hernández-Ortega, Sara, 66
Hertz, Emil Peter Thrane, 67
Hidalgo, Elena, 41
Hirai, Kazuyuki, 68
Hiraoka, Daisaku, 8
Hirota, Takayuki, 13
Hiruma, Yoshitaka, 107
Hochegger, Helfrid, 109
Hofmann, Kay, 124
Holubcova, Zuzana, 10
Horváth, Anna, 70
Horváth, Beatrix, 69, 90
Hotz, Manuel, 35
Huang, Jin, 46
Hudson, Damien, 132
Hutter, Lukas, 52, 71, 97
Hyman, Anthony A., 28
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Jang, Young-Joo, 78
JÁTIVA, SORAYA, 74
Jiménez, Javier, 66
Jonak, Katarzyna, 72
Joseph, Stephy, 73
Juanes, Maria Angeles, 36
Kalitsis, Paul, 132
Kapuy, Orsolya, 75
Karasu, Mehmet Erman, 76
Kataria, Meghna, 77
Kawamura, Miyuki, 125
Keeney, Scott, 76
Kermi, Chames, 16
Kies, Matthias, 123
Kim, Jae Hyeong, 78
Kim, Kyungtae, 78
Kirschner, Marc, 85
Kishimoto, Takeo, 8
Klein, Carlo, 62
Kleiss, Charlotte, 124
Klinkert, Kerstin, 54
Kobarg, Jörg, 122
Koff, Andrew C, 76
Kohoutová, Lucie, 79
Kõivomägi, Mardo, 3
Komaki, Shinichiro, 80
Konietzny, Anja, 64
Kops, Geert, 42, 107
Korcsmáros, Tamás, 75
Korolchuk, Svitlana, 81
Kourová, Hana, 69, 79
Kovács, Anita, 90
Kovács, Levente, 82, 100
Kruitwagen, Tom, 83
Krupina, Ksenia, 124
Kruse, Thomas, 67
Kuijt, Timo, 107
Kurucz, Anita, 75
Le Dez, Gaelle, 45
Lee, Byung Il, 78
Lee, Chang-Woo, 78
Leger, Thibault, 126
Lengefeld, Jette, 35
Lens, Susanne, 38, 84, 94
Leontiou, Ioanna, 11
Lesage, Bart, 46
Leviczky, Tünde, 90
Li, Victor, 85
Li, Wei, 119
Ligammari, Lorena, 129
Lindqvist, Arne, 86, 120
Lio, Pietro, 110
Liu, Boyang, 61
Liu, Jian, 87
Lobjois, Valérie, 88
Lopez, María Isabel, 51
Lorca, Thierry, 89
Losada, Ana, 27
Lu, Dan, 32
Lygerou, Zoi, 101
Ma, Sheng, 89
Magiera, Magda, 14
Magyar, Zoltán, 5, 90
Maiorano, Domenico, 16
Makarova, Maria, 6
Malumbres, Marcos, 31
Manenti, Stéphane, 92
Mangat, Davinderpreet, 91
Marcellin, Marlene, 92
Marquina, Maribel, 41
Martin, Mathew, 81
Martino, Lisa, 126
Massoni-Laporte, Aurelie, 112
Matsuda, Muneo, 68
Maxouri, Stella, 101
Mayer, Thomas U., 64
Mayor, Federico, 114
Mazzolini, Laurent, 92
McAinsh, Andrew, 96
McCusker, Derek, 112
Meca, Julien, 112
Melbinger, Anna, 22
Mendoza, Manuel, 93
Mengoli, Valentina, 7, 40
Meppelink, Amanda, 94
Merlini, Laura, 36
Meskiene, Irute, 79
Mészáros, Tamás, 69, 79
Miles, Catrina, 95
Millar, Jonathan, 96
Minakuchi, Yohei, 68
Mirkovic, Mihailo, 71, 97
Mironov, Svetlana, 45
Mitteau, Romain, 112
Miyawaki, Atsushi, 15
Mizrak, Arda, 32
Mochida, Satoru, 98
Mohammed, Binish, 90
Mohammed, Shabaz, 34
MOLINA-DELGADO, Angie, 106
Molist, Iago, 57
Momen Roknabadi, Amir, 1
Mondésert, Odile, 88
Monje-Casas, Fernando, 99
Monnier, Sylvain, 2
Mora-Santos, Maria Del Mar,
96
Moreno, David, 4
Morgan, David, 32, 59
Müller-Reichert, Thomas, 28
Müllers, Erik, 120
Nagai, Takeharu, 98
135
List of contributors
Nagy, Ágota, 100
Nagy, Olga, 82, 100
Nagy, Szilvia, 69
Nathanailidou, Patroula, 101
Nebreda, Angel, 102
Nemeth, Edit, 69
Neumann, Beate, 28
Neumann, Heinz, 83
Neurohr, Gabriel, 103
Nilsson, Jakob, 67
Nitzan, Mor, 48
Noach-Hirsh, Meirav, 128
Noble, Martin, 81
Novák, Béla, 43, 50, 52, 71, 97,
113
Novak, Zsofia, 104
Nunes Bastos, Ricardo, 34
O'Farrell, Patrick H, 21
Ohkura, Hiro, 44
Ohsugi, Miho, 121
Oliferenko, Snezhana, 6
Oliveira, Raquel, 71, 97
Olsen, Jesper Velgaard, 67
Otero, Sofía, 51
Pachis, Spyridon, 107
Pál, Margit, 82, 100
Panbianco, Costanza, 126
Pane, Attilio, 119
Papdi, Csaba, 5, 69, 90
Parisi, Eva, 4
Pe'er, Tal, 128
Penela, Petronila, 114
Pentakota, Satyakrishna, 108
Perez, Arina Marina, 122
Perrakis, Anastassis, 107
Peter, Nisha, 109
Peters, Jan-Michael, 26
Petit, Julie, 14
Pettkó-Szandtner, Aladár, 5
Piatti, Simonetta, 36
Piel, Matthieu, 2
Pintard, Lionel, 126
PITUELLO, Fabienne, 106
Poch, Olivier, 124
Ponsioen, Bas, 38
Popescu, Octavian, 82
Poser, Elena, 34
Poser, Ina, 28
Prieto, Susana, 16
Prigent, Claude, 45
Przewloka, Marcin, 110
Qian, Junbin, 46
Queralt, Ethel, 111
Raff, Jordan, 104
Rapali, Péter, 112
Rapsomaniki, Anna Maria, 101
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
Rata, Scott, 113
Reboutier, David, 45
Recolin, Benedicte, 16
Reglero, Clara, 114
Ricco, Natalia, 66
Rivas, Verónica, 114
Robert, Perle, 89
Roccuzzo, Michela, 130
Rodriguez-Rodriguez, Jose
Antonio, 111
Rojas, Julie, 115
Rowland, Benjamin, 116
Royou, Anne, 117
Ruiz-Torres, Miguel, 27
Sacristan, Carlos, 107
Saitou, Mitinori, 13
Sajman, Julia, 48
Sanchez-Diaz, Alberto, 57
Sanchez-Perez, Gabino, 69
Santamaria, Anna, 105, 126
Santos, Silvia, 23, 39
Sanz-Castillo, Belén, 31
Sanz-Flores, María, 31
Schalbetter, Stephanie, 118
Scheres, Ben, 69
Schiklenk, Christoph, 62
Schiltz, Odile, 92
Schmoller, Kurt, 3
Schmucker, Stephane, 124
Schnittger, Arp, 19, 80
Schoonen, Pepijn, 84
Schuh, Melina, 10
Schupbach, Trudi, 119
Schwager, Francoise, 126
Schwob, Etienne, 14
Sheriff, Rahuman, 39
Shermoen, Antony W, 21
Shim, Jaegal, 78
Sigurdsson, Jón Otti, 67
Silio, Virginia, 96
Silva Cascales, Helena, 120
Skotheim, Jan, 3
Smetana, Juliana, 122
Smith, Jim, 17
Soeda, Shou, 121
Sprenger, Frank, 123
Stanley, Will, 81
Stemmann, Olaf, 113
Sumara, Izabela, 124
Sung, Hung-wei, 61
Susumu, Hiroaki, 125
Suzuki, Haruka, 68
Sveiczer, Ákos, 70
Szeker, Kathelijne, 56
Tachibana-Konwalski, Kikue,
13
Takahashi, Naoki, 19
Takaki, Kaori, 98
Talarek, Nicolas, 14
Tanno, Yuji, 125
Tapon, Nicolas, 24
Taraviras, Stavros, 101
Tavernier, Nicolas, 126
Teixeira, Antoinette, 42
Telley, Ivo, 25
Thomas, Yann, 126
Tollis, Sylvain, 112
Torok, Katalin, 65
Touati, Sandra, 127
Toyoda, Atsushi, 68
Tsanov, Nikolay, 16
Turner, Jonathan, 3
Tyson, John, 50
Tzur, Amit, 128
Ubbink, Marcellus, 107
136
List of contributors
Udvardy, Andor, 82
Uhlmann, Frank, 60, 77, 127
Umeda, Masaaki, 19
Vagnarelli, Paola, 129
Vallot, Antoine, 11
van Castelmur, Eleonore, 107
van der Laan, Siem, 16
Van Eynde, Aleyde, 56
Van Hove, Lucie, 126
van Vugt, Marcel, 84
Vaz Meirelles, Gabriela, 122
Vergassola, Massimo, 22
Vietri, Marina, 37
Vigneron, Suzanne, 89
Visintin, Clara, 130
Visintin, Rosella, 130
Volc, Jindřich, 79
Wang, Jin, 62
Wassmann, Katja, 11
Watanabe, Yoshinori, 9, 125
Weimer, Annika, 19
Wieschaus, Eric, 1
Wilkins, Brian, 83
Winkler, Franziska, 61
Wu, Pei-Yun, 131
Xiang, Jinyi, 20
Yahya, Galal, 4
Yu, Miao, 95
Yu, Xueyang, 20
Yuan, Kai, 21
Zachariae, Wolfgang, 7, 40, 72,
115
Zagoriy, Ievgeniia, 7, 40
Zegerman, Philip, 17
Zenvirth, Drora, 48
Zhang, Tao, 132
Zhang, Tongli, 43
Zielke, Norman, 20, 133
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
List of participants
LIST OF PARTICIPANTS
ADRIAANS, Ingrid
UMC Utrecht
Universiteitsweg 100
3584 CG, Utrecht, Netherlands
[email protected]
BARRAL, Yves
ETH Zuerich Institute of Biochemistry
IBC HPM D12.2
8093, Zuerich, Switzerland
[email protected]
ALDEA, Marti
Molecular Biology Institute of
Barcelona, CSIC
Baldiri Reixac 15 3-A16
08028, Barcelona, Spain
[email protected]
BEAVEN, Robin
University of Edinburgh
Michael Swann Building
EH9 3JR, Edinburgh, United Kingdom
[email protected]
ALVAREZ-FERNÁNDEZ, Mónica
Spanish National Cancer Research
Centre (CNIO)
C/ Melchor Fernandez Almagro, 3
28029, Madrid, Spain
[email protected]
AMON, Angelika
Koch Institute/HHMI/MIT
77 Mass Ave
02139, Cambridge, United States
[email protected]
ANGER, Martin
Masaryk University
Department of histology and
embryology
Kamenice 3
Brno, Czech Republic
[email protected]
ARAUJO, Ana Rita
MRC-Clinical Sciences Centre,
Imperial College London
B409, 80 Wood Lane
W12 0BZ, London, United Kingdom
[email protected]
ARGUELLO-MIRANDA, Orlando
Max Planck Institute
Roentgen Str 1a
82152, Planegg, Germany
[email protected]
AYTE, Jose
Universitat Pompeu Fabra
Doctor Aiguader 88
08003, Barcelona, Spain
[email protected]
BADE, Debora
UMC Utrecht
Universiteitsweg 100
3584 CG, Utrecht, Netherlands
[email protected]
BARR, Alexis
Institute of Cancer Research
Chester Beatty Laboratories
SW3 6JB, London, United Kingdom
[email protected]
BENAUD, Christelle
IGDR-UMR6290
2 av Leon Bernard
35043, Rennes, France
[email protected]
BOGRE, Laszlo
Royal Holloway University of London
Egham High Hill
TW20 0EX, Egham, United Kingdom
[email protected]
BOLLEN, Mathieu
KU Leuven
Campus Gasthuisberg 0&N1
3000, Leuven, Belgium
[email protected]
BONNER, Amanda
Stowers Institute for Medical Research
1000 East 50th Street
64111, Kansas City, United States
[email protected]
BRANDEIS, Michael
The Hebrew University of Jerusalem
Safra Campus
9190401, Jerusalem, Israel
[email protected]
CADART, Clotilde
Institut Curie
12 rue Lhomond
75007, Paris, France
[email protected]
CASTRO, Anna
CRBM-CNRS
1919 route de Mende
34293, Montpellier, France
[email protected]
CILIBERTO, Andrea
IFOM, Milan
Via Adamello 16
Milan, Italy
[email protected]
COUDREUSE, Damien
Institute of Genetics and Development
of Rennes
2, avenue du Pr. Léon Bernard
35043, Rennes, France
[email protected]
137
CUNDELL, Michael
University of Oxford, Department of
Biochemistry
South Parks Road
OX13QU, Oxford, United Kingdom
[email protected]
CUYLEN, Sara
Institute of Molecular Biotechnology of
the Austrian Academy of Sciences
(IMBA)
Dr.-Bohr Gasse 3
1030, Vienna, Austria
[email protected]
DE SOUZA, Edmarcia Elisa
National Center For Research in Energy
and Materials
Giuseppe Máximo Scolfaro 10000
13083-970, Campinas, Brazil
[email protected]
DESHPANDE, Ojas
Instituto Gulbenkian de Ciencia
Rua Quinta Grande 6
Oeiras, Portugal
[email protected]
DESVOYES, Bénédicte
Centro de Biología Moleculal Severo
Ochoa, CSIC
calle Nicolas Cabrera,1
28049, Madrid, Spain
[email protected]
DI TALIA, Stefano
Duke University Medical Center
Nanaline Duke Building
27710, Durham, United States
[email protected]
DICK, Amalie
IMBA - Institute of Molecular
Biotechnology GmbH
Dr. Bohr-Gasse 3
1020, Wien, Austria
[email protected]
DITTE, Peter
Max Planck Institute for Biophysical
Chemistry
Am Faßberg 11
Göttingen, Germany
[email protected]
ECHARD, ARNAUD
INSTITUT PASTEUR
Membrane Traffic and Cell Division Lab
28 RUE DU DOCTEUR ROUX
75724, PARIS CEDEX 15, France
[email protected]
EDGAR, Bruce
University of Heidelberg
Im Neuenheimer Feld 282
Heidelberg, Germany
[email protected]
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
List of participants
FARCAS, Ana-Maria
Institute of Biochemistry, ETH Zurich
Otto-Stern Weg 3
8093, Zurich, Switzerland
[email protected]
HELASSA, Nordine
University of Liverpool
Crown Street
L69 3BX, Liverpool, United Kingdom
[email protected]
JOSEPH, Stephy
University of Sussex
Genome damage and stability centre
BN1 9RH, Brighton, United Kingdom
[email protected]
FERREIRA, Monica
KU Leuven
Campus Gasthuisberg
3000, Leuven, Belgium
[email protected]
HERNÁNDEZ-ORTEGA, Sara
Universitat Internacional de Catalunya
Josep Trueta s/n
08195, Sant Cugat del Vallès, Spain
[email protected]
KAPUY, Orsolya
Semmelweis University
Tűzoltó utca 37-47
1094, Budapest, Hungary
[email protected]
GALLI, Matilde
UCSF
600 16th Street
94158, San Francisco, United States
[email protected]
HERTZ, Emil Peter Thrane
Novo Nordisk Foundation Center for
Protein Research, Faculty of Health and
Medical Sciences, University of
Copenhagen
Blegdamsvej 3b
2200, Copenhagen, Denmark
[email protected]
GIET, Régis
Université of Rennes 1-CNRS
2, Avenue du Pr Leon bernard
35043, RENNES, France
[email protected]
GODFREY, Molly
The Francis Crick Institute
44 Lincoln's Inn Fields
WC2A 3LY, London, United Kingdom
[email protected]
GROHANS, Jörg
University of Göttingen
Justus-von-Liebig Weg 11
37077, Göttingen, Germany
[email protected]
GRUNEBERG, Ulrike
University of Oxford
South Parks Road
OX1 3RE, Oxford, United Kingdom
[email protected]
GUTIERREZ, Crisanto
Centro de Biologia Molecular Severo
Ochoa, CSIC-UAM
Nicolas Cabrera 1
Madrid 28049, Spain
[email protected]
HAERING, Christian
EMBL
Meyerhofstr. 1
69117, Heidelberg, Germany
[email protected]
HIRAI, Kazuyuki
Kyorin University School of Medicine
6-20-2
181-8611, Mitaka, Japan
[email protected]
HORVATH, Beatrix
Royal Holloway, University of London
Egham
TW20 OEX, Egham, United Kingdom
[email protected]
HORVÁTH, Anna
Budapest University of Technology and
Economics, Department of Applied
Biotechnology and Food Science
Szent Gellért tér 4.
H-1111, Budapest, Hungary
[email protected]
HUNT, Tim
Cancer Research UK
Rose Cottage
EN6 3LH, Potters Bar, United Kingdom
[email protected]
HUTTER, Lukas
Department of Biochemistry/ University
of Oxford
Lincoln College
OX1 3DX, Oxford, United Kingdom
[email protected]
HAYWARD, Daniel
University of Oxford
Sir William Dunn School of Pathology
OX1 3RF, Oxford, United Kingdom
[email protected]
JÁTIVA, SORAYA
BELLVITGE INSTITUTE OF BIOMEDICAL
RESEARCH (IDIBELL)
Av. Gran Via de L'Hospitalet 199-203
08908, L'Hospitalet de Llobregat, Spain
[email protected]
HEIM, Andreas
University of Konstanz
Universitätsstraße 10
78457, Konstanz, Germany
[email protected]
JONAK, Katarzyna
Max Planck Institute for Biochemistry
Am Klopferspitz 18
82152, Martinsried/Munich, Germany
[email protected]
138
KARASU, Mehmet Erman
MSKCC- Gerstner Sloan Kettering
Graduate School
1233 York Ave Apt 16 B
10065, New York, United States
[email protected]
KATARIA, Meghna
The Francis Crick Institute
London WC2A 3LY, United Kingdom
[email protected]
KIM, Kyungtae
National Cancer Center, Research
Institute
323 Ilsan-ro, Ilsandong-gu,
410-769, Goyang-si, Korea, Republic of
(South)
[email protected]
KISHIMOTO, Takeo
Ochanomizu University
Ootsuka 2-1-1
112-8610, Bunkyo-ku, Japan
[email protected]
KOHOUTOVÁ, Lucie
Institute of Microbiology AS CR, v. v. i.
Videnska 1083
142 20, Prague, Czech Republic
[email protected]
KOMAKI, Shinichiro
University of Hamburg
Ohnhorststr. 18
22609, Hamburg, Germany
[email protected]
KOROLCHUK, Svitlana
Newcastle University
NICR
NE24HH, Newcastle upon Tyne, United
Kingdom
[email protected]
KOVÁCS, Levente
Institute of Biochemistry, Biological
Research Centre, Hungarian Academy
of Sciences
Temesvári krt
6726, Szeged, Hungary
[email protected]
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
List of participants
KRUITWAGEN, Tom
Institute of Biochemistry, ETH Zurich
Otto-Stern-Weg 3
8093, Zurich, Switzerland
[email protected]
MANGAT, Davinderpreet
University of Oxford
16 vicarage farm road
TW3 4NW, London, United Kingdom
[email protected]
NAGY, Ágota
University of Szeged
Közép Fasor
6726, Szeged, Hungary
[email protected]
LEGATE, Kyle
Nature Communications
4 Crinan Street
London, United Kingdom
[email protected]
MAZZOLINI, Laurent
CNRS
2, avenue Hubert Curien
31037, Toulouse, France
[email protected]
NATHANAILIDOU, Patroula
University of Patras
Telou Agra 6A
26442, Patra, Greece
[email protected]
LENS, Susanne
UMC Utrecht
Universiteitsweg 100
3584 CG, Utrecht, Netherlands
[email protected]
MENDOZA, Manuel
Center for Genomic Regulation (CRG)
C/ Dr Aiguader 88
08003, Barcelona, Spain
[email protected]
NEBREDA, Angel
IRB Barcelona
C/ Baldiri Reixac 10
08028, Barcelona, Spain
[email protected]
LI, Victor
Harvard Medical School
200 longwood Avenue
02115, Boston, United States
[email protected]
MEPPELINK, Amanda
University Medical Center Utrecht
Universiteitsweg 100
3584 CG, Utrecht, Netherlands
[email protected]
NEUROHR, Gabriel
Massachusetts Institute of Technology
68 Wells Rd
Lincoln, United States
[email protected]
LINDQVIST, Arne
Karolinska Institutet
von eulers v 3
171 77, Stockholm, Sweden
[email protected]
MILES, Catrina
Genome damage and Stability center
(Sussex)
Science Park road
BN1 9QG, Falmer, United Kingdom
[email protected]
NOVAK, Bela
Department of Biochemistry/University
of Oxford
South Parks Road
OX1 3QU, Oxford, United Kingdom
[email protected]
MILLAR, Jonathan
University of Warick
Gibbet Hill Road
Coventry, United Kingdom
[email protected]
NOVAK, Zsofia
Dunn School of Pathology, University of
Oxford
South Parks Road
OX1 3RE, Oxford, United Kingdom
[email protected]
LIU, Jian
NIH
50 South Drive, Room 3306
Bethesda, United States
[email protected]
LOBJOIS, Valerie
CNRS-ITAV USR3505
1 place Pierre POTIER
31106, TOULOUSE, France
[email protected]
LORCA, Thierry
CNRS
1919 route de mende
34293, Montpellier, France
[email protected]
LOSADA, Ana
Spanish National Cancer Research
Centre
Melchor Fernandez Almagro 3
E28029, Madrid, Spain
[email protected]
MAGYAR, ZOLTÁN
Institute of Plant Biology, Biological
Research Centre, Szeged
TEMESVÁRI KRT 62
6726, SZEGED, Hungary
[email protected]
MAIORANO, Domenico
Institute of Human Genetics. CNRSUPR1142.
rue de la cardonille
34396 Cedex 5, Montpellier, France
[email protected]
MIRKOVIC, Mihailo
Instituto Gulbenkian de Ciencia
Rua Quinta das Palmeiras 45,5f
2780-156, Oeiras, Portugal
[email protected]
MIYAWAKI, Atsushi
RIKEN
2-1 Hirosawa, Wako
Saitama, Japan
[email protected]
MOCHIDA, Satoru
Kumamoto university
Kyoyotou-Honjo 1
860-0811, Kumamoto, Japan
[email protected]
MONJE-CASAS, Fernando
University of Seville
CABIMER. Avda. Americo Vespucio, s/n
Seville, Spain
[email protected]
MORGAN, David
University of California, San Francisco
UCSF Genentech Hall, Room N312B
94158, San Francisco, United States
[email protected]
139
O'FARRELL, Patrick
UCSF
600-16th Street
94153, San Francisco, United States
[email protected]
OLIFERENKO, Snezhana
King's College London
New Hunt's House
SE1 1UL, London, United Kingdom
[email protected]
PACHIS, Spyridon
UMC Utrecht
Vredenburg 28C
3511 BC, Utrecht, Netherlands
[email protected]
PENTAKOTA, satya krishna
Max planck institute of Molecular
physiology
AMRODE
44149, DORTMUND, Germany
[email protected]
PETER, Nisha
Genome Damage and Stability Centre,
University of Sussex
Science Park Road
BN2 4HY, Brighton, United Kingdom
[email protected]
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
PETERS, Jan-Michael
Research Institute of Molecular
Pathology
Dr. Bohr-Gasse 7
1030, Vienna, Austria
[email protected]
PIATTI, Simonetta
CNRS
1919 Route de Mende
34293, Montpellier, France
[email protected]
PINTARD, Lionel
CNRS
15 rue hélène Brion
75013, PARIS, France
[email protected]
PITUELLO-BERNIERE, Fabienne
Centre de Biologie du Développement
UMR5547 CNRS/UPS
118 rte de Narbonne Bât 4R3
31062 cedex 09, Toulouse, France
[email protected]
PRZEWLOKA, Marcin
University of Cambridge
Department of Genetics
CB2 3EH, Cambridge, United Kingdom
[email protected]
QUERALT, Ethel
Instituto de Investigaciones Biomédicas
de Bellvitge (IDIBELL)
Avenida Gran via de L'Hospitalet 199203
08908, Barcelona, Spain
[email protected]
RAPALI, Peter
Institut de Biochimie et Génétique
Cellulaires, France
14 Rue du Haut Carre
33400, Talence, France
[email protected]
RATA, Scott
Department of Biochemistry, University
of Oxford
Keble College
OX1 3PG, Oxford, United Kingdom
[email protected]
REGLERO, Clara
Centro de Biología Molecular Severo
Ochoa (CBMSO) - Universidad
Autónoma de Madrid (UAM)
C/ Nicolás Cabrera, 1
28049, Madrid, Spain
[email protected]
ROJAS, Julie
Max Planck Institute of Biochemistry
Am Klopferspitz 18
82152, Martinsried, Germany
[email protected]
List of participants
ROWLAND, Benjamin
The Netherlands Cancer Institute
Plesmanlaan 121
1066 CX, Amsterdam, Netherlands
[email protected]
SOEDA, Shou
The University of Tokyo
3-8-1 Komaba
153 8902, Meguro, Japan
[email protected]
ROYOU, Anne
IECB/CNRS
2 rue Robert Escarpit
33607, PESSAC, France
[email protected]
SPRENGER, Frank
University of Regensburg
Universitaetsstrasse 31
93053, Regensburg, Germany
[email protected]
SANCHEZ-DIAZ, ALBERTO
UNIVERSITY OF CANTABRIA
INSTITUTO DE BIOMEDICINA Y
BIOTECNOLOGIA DE CANTABRIA
39012, SANTANDER, Spain
[email protected]
SUMARA, Izabela
IGBMC
1, rue Laurent Fries
67400, Illkirch, France
[email protected]
SANTOS, Silvia
MRC-Imperial College London
Clinical Sciences Centre
W12 0NN, London, United Kingdom
[email protected]
SCHALBETTER, Stephanie
GDSC, University of Sussex
Science Park Road
Brighton, United Kingdom
[email protected]
SCHMOLLER, Kurt
Stanford University
337 Campus Drive
Stanford, United States
[email protected]
SCHNITTGER, Arp
University of Hamburg
Ohnhorststr. 18
22609, Hamburg, Germany
[email protected]
SCHUH, Melina
MRC Laboratory of Molecular Biology
Francis Crick Avenue
CB2 0QH, Cambridge, United Kingdom
[email protected]
SCHUPBACH, Trudi
Princeton University
Washington Road
08544, Princeton, United States
[email protected]
SCHWOB, Etienne
CNRS Institute of Molecular Genetics
IGMM CNRS
34293, Montpellier, France
[email protected]
SILVA CASCALES, Helena
Karolinska Institutet
Ölandsgatan 52
116 63, Stoclholm, Sweden
[email protected]
140
TACHIBANA-KONWALSKI, Kikue
IMBA, Institute of Molecular
Biotechnology of the Austrian Academy
of Sciences
Dr. Bohr Gasse 3
1030, Vienna, Austria
[email protected]
TANNO, Yuji
IMCB, University of Tokyo
Yayoi
113-0032, Bunkyo-ku, Tokyo, Japan
[email protected]
TAPON, Nicolas
Cancer Research UK London Research
Institute
5 Hartswood Road
W12 9NQ, London, United Kingdom
[email protected]
THOMAS, Yann
Institut Jacques Monod/ Universite
Paris Diderot
15 rue Helene Brion
75205, Paris, France
[email protected]
TOUATI, Sandra
CRICK
44 Lincoln's Inn Fields
London, United Kingdom
[email protected]
TZUR, Amit
Bar-Ilan
Keren Hayesod 15/a
Givat Shmuel, Israel
[email protected]
UHLMANN, Frank
The Francis Crick Institute
44 Lincoln's Inn Fields
WC2A 3LY, London, United Kingdom
[email protected]
VAGNARELLI, Paola
Brunel University London
Heinz Wolff Building
UB8 3PH, Uxbridge, London, United
Kingdom
[email protected]
EMBO Cell Cycle Workshop, Budapest, Hungary (2015)
VIETRI, Marina
2Department of Molecular Cell Biology,
Institute for Cancer Research, Oslo
University Hospital
Montebello
0379, Oslo, Norway
[email protected]
VISINTIN, Rosella
European Institute of Oncology
Via Adamello, 16
Milan, Italy
[email protected]
WASSMANN, Katja
IBPS- Institut de Biologie Paris Seine
9 quai St. Bernard
75005, Paris, France
[email protected]
WATANABE, Yoshinori
University of Tokyo
Yayoi 1-1-1, Bunkyo-ku
113-0032, Tokyo, Japan
[email protected]
WIESCHAUS, Eric
Princeton University
435 Moffett Laboratory
08544, Princeton, United States
[email protected]
WU, Pei-Yun Jenny
Institute of Genetics and Development
of Rennes - CNRS
2 av. du Pr. Léon Bernard
35043, Rennes, France
[email protected]
ZACHARIAE, Wolfgang
Max Planck Institute of Biochemistry
Am Klopferspitz 18
D-82152, Martinsried, Germany
[email protected]
ZEGERMAN, Philip
Gurdon Institute, University of
Cambridge
Tennis Court Road
Cambridge, United Kingdom
[email protected]
ZHANG, Tao
Murdoch Childrens Research Institute
Royal Childen Hospital
3052, Melbourne, Australia
[email protected]
ZIELKE, Norman
DKFZ
Im Neuenheimer Feld 282
69120, Heidelberg, Germany
[email protected]
141
List of participants
PREVIOUS EUROPEAN CELL CYCLE CONFERENCES
Year
Location
Organizer
1st
1969
Copenhagen (Denmark)
Erik Zeuthen
2nd
1972
Innsbruck (Austria)
Wilhelm Sachensenmaier
3rd
1974
Zürich (Switzerland)
Gerhard Braun
4th
1976
Edinburgh (Scotland)
Murdoch Mitchison
5th
1978
Madrid (Spain)
Jorge Lopez-Saez
6th
1981
Prague (Czechoslovakia)
Eva Streiblova
7th
1984
Heidelberg (Germany)
Christian Petzelt
8th
1987
Bath (England)
Allan Wheals
9th
1990
Stockholm (Sweden)
Anders Zetterberg
10th 1993
La Rochelle (France)
Christian Petzelt
11th 1997
Gardone (Italy)
Lilia Alberghina
12th 2001
Mayerhofen (Austria)
Wilhelm Sachsenmaier
13th 2004
Salamanca (Spain)
Sergio Moreno
14th 2007
Spetses (Greece)
Stathis Gonos
15th 2011
Montpellier (France)
Etienne Schwob
16th 2015
Budapest (Hungary)
Béla Novák & Frank Uhlmann
registration
desk
MAP OF THE HOTEL GROUND FLOOR
Stairs to
Auditorium
Jázmin corridor
Platán