* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Is this an inducible or repressible operon?
DNA vaccination wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Polyadenylation wikipedia , lookup
History of genetic engineering wikipedia , lookup
Non-coding DNA wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Histone acetyltransferase wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Microevolution wikipedia , lookup
Protein moonlighting wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Deoxyribozyme wikipedia , lookup
History of RNA biology wikipedia , lookup
Epigenomics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Helitron (biology) wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Messenger RNA wikipedia , lookup
Non-coding RNA wikipedia , lookup
Frameshift mutation wikipedia , lookup
Primary transcript wikipedia , lookup
Genetic code wikipedia , lookup
Expanded genetic code wikipedia , lookup
Point mutation wikipedia , lookup
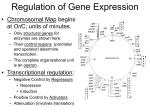
T R A N S L AT I O N , M U T AT I O N , R E G U L AT I O N , O H M Y ! AGENDA Translation 3 players Initiation, Elongation, Termination Mutation Point Mutations and their consequences Chromosomal Mutation Regulation of Gene Expression Prokaryotic Eukaryotic 3 PARTICIPANTS OF TRANSLATION 1.mRNA 2.Ribosome 3.tRNA MESSENGER RNA - What process creates mRNA? - What kind of mRNA modifications do prokaryotes have? - What kind of mRNA modifications do eukaryotes have? - From the below pictures, which mRNA is monocistronic and which mRNA is polycistronic? - Which would you find in a prokaryotic cell and which would you find in a eukaryotic cell? MESSENGER RNA - What process creates mRNA? transcription - What kind of mRNA modifications do prokaryotes have? none - What kind of mRNA modifications do eukaryotes have? Poly-A tail, 5’ Guanosine cap, and alternative splicing - From the below pictures, which mRNA is monocistronic and which mRNA is polycistronic? Bottom is monocistronic, top is polycistronic - Which would you find in a prokaryotic cell and which would you find in a eukaryotic cell? You can find monocistronic and polycistronic in a prokaryotic cell, but only monocistronic genes in eukaryotes. RIBOSOME - How many Svedburg units are the large and small subunits of the ribosome? - Ribosomes are made from ribosomal RNA (rRNA) and ribosomal proteins. - Where is the mRNA that codes for ribosomal proteins transcribed? - Where is rRNA transcribed? - Where are the ribosomal proteins and rRNA combined to make ribosomal subunits? - Where are the two subunits combined to make a ribosome? - When does rRNA get translated? - What does peptidyl transferase do? nucleus nucleolus chromosome - RIBOSOME How many Svedburg units are the large and small subunits of the ribosome? In the eukaryotic ribosome, the large subunit is 60s and the small is 40 s. The entire ribosome is 80s. - Ribosomes are made from ribosomal RNA (rRNA) and ribosomal proteins. - Where is the mRNA that codes for ribosomal proteins transcribed? in the nucleus, outside the nucleolus - Where is rRNA transcribed? in the nucleolus - Where are the ribosomal proteins and rRNA combined to make ribosomal subunits? In the nucleolus - Where are the two subunits combined to make a ribosome? cytoplasm - When does rRNA get translated? Never - What does peptidyl transferase do? Make peptide bonds chromosome nucleolus nucleus TRANSFER RNA - Where is tRNA transcribed? - Label the 5’ and 3’ ends on the tRNA below. - What amino acid should be in the box? - What is the name of the enzyme below? - How many different types of the enzyme below are there? - What anticodon sequence will the enzyme below bind with? - What would happen if the enzyme below was nonfunctional? - What is the difference between an uncharged tRNA and an aminoacyl tRNA? - TRANSFER RNA Where is tRNA transcribed? In the nucleus - Label the 5’ and 3’ ends on the tRNA below. - What amino acid should be in the box? Phe - What is the name of the enzyme below? Aminoacyl- tRNA synthetase - How many different types of the enzyme below are there? 20 - What anticodon sequence will the enzyme below bind with? 3’ UAA, UAG, UAU 5’ - What would happen if the enzyme below was nonfunctional? There would be no aminoacyl tRNA with Ile, which would probably stop translation in the cell - What is the difference between an uncharged tRNA and an aminoacyl tRNA? An aminoacyl tRNA has an amino acid 3’ 5’ INITIATION OF TRANSLATION - Which rRNA base pairs with the mRNA? - What proteins facilitate translation initiation? - What is the initiator tRNA for euks and for proks? - Draw a picture of the ribosome-mRNA complex at the end of initiation. What is in the E, P, and A sites? - Steps of initiation: - 1. small subunit binds to the 5’ UTR of mRNA - 2. initiator tRNA binds to small subunit - 3. small subunit scans for 5’AUG 3’ then stops - 4. large subunit joins INITIATION OF TRANSLATION - Which rRNA base pairs with the mRNA? 18s (the rRNA of the small subunit) - What proteins facilitate translation initiation? Translation initiation factors - What is the initiator tRNA for euks and for proks? In euks, it is a tRNA-met with different secondary structure. In proks, it is a tRNA- fmet with a formylated methionine. - Draw a picture of the ribosome-mRNA complex at the end of initiation. What is in the E, P, and A sites? The Exit site and Aminoacyl site are empty. The Peptidyl site has an initiator tRNA. - Steps of initiation: - 1. small subunit binds to the 5’ UTR of mRNA - 2. initiator tRNA binds to small subunit - 3. small subunit scans for 5’AUG 3’ then stops - 4. large subunit joins ELONGATION OF TRANSLATION What are the four steps of translation? Why do you think peptide bond formation happens before hydrolysis of the amino acid? What does the ribosome-mRNA complex look like after the first step of elongation? What does the ribosome-mRNA complex look like after translocation? ELONGATION OF TRANSLATION What are the four steps of translation? 1. Amino acylated tRNA enters A site 2. Peptide bond formation 3. Hydrolysis of bond between tRNA and amino acid in P-site 4. Translocation, movement of ribosome down one codon in 3’ direction in mRNA Why do you think peptide bond formation happens before hydrolysis of the amino acid? If the tRNA released the amino acid before the amino acid is bonded to the chain, the amino acid could possibly leave the ribosome before it is able to be incorporated into the protein. What does the ribosome-mRNA complex look like after the first step of elongation? There is an initiator tRNA in the P-site and an amino acylated tRNA in the A site. If it’s not the first round of elongation, there would be an uncharged tRNA in the Esite, and an aminoacyl tRNA in the P and A sites. What does the ribosome-mRNA complex look like after translocation? The E site has an uncharged tRNA, the P site has an aminoacyl tRNA, and the A site is empty. TERMINATION OF TRANSLATION What enters the A site in termination? Are there an anticodons for the stop codons? TERMINATION OF TRANSLATION What enters the A site in termination? Release factor protein Are there an anticodons for the stop codons? No, release factors don’t have RNA Release factors cause the release of the polypeptide, the release of the mRNA and the separation of the two subunits. WOBBLE HYPOTHESIS - The wobble hypothesis is the reason why there are 61 amino acid encoding codons but only about 45 different tRNAs. - There are exceptions to base pairing rules between the codon and the anticodon for the third nucleotide. - Guanine on the tRNA anticodon can base pair with either U or C on the mRNA codon. - Uracil on the tRNA anticodon can base pair with either A or G on the mRNA codon. - If the Anticodon is 3’GCA 5’, what codons can the tRNA base pair with? - If the anticodon is 3’UUU 5’, what codons can the tRNA base pair with? WOBBLE HYPOTHESIS - The wobble hypothesis is the reason why there are 61 amino acid encoding codons but only about 45 different tRNAs. - There are exceptions to base pairing rules between the codon and the anticodon for the third nucleotide. - Guanine on the tRNA anticodon can base pair with either U or C on the mRNA codon. - Uracil on the tRNA anticodon can base pair with either A or G on the mRNA codon. - If the Anticodon is 3’GCA 5’, what codons can the tRNA base pair with? 5’ CGU 3’ - If the anticodon is 3’UUU 5’, what codons can the tRNA base pair with? 5’ AAA 3’ and 5’ AAG 3’ POINT MUTATIONS - What are the four different point mutations? - List all the transition mutations. - List all the transversion mutations. POINT MUTATIONS - What are the four different point mutations? - 1. transition - 2. transversion - 3. insertion - 4. deletion - List all the transition mutations. - A G, GA, TC, CT - List all the transversion mutations. - AC, AT, GC, GT, TA, TG, CA, CG CONSEQUENCES OF POINT MUTATION ON A.A. A. Silent mutation- point mutation doesn’t change the amino acid encoded from the codon B. Missense mutation- causes change in encoded amino acid C. Nonsense mutation– a codon that originally encoded for an amino acid is changed so that it encodes a premature stop codon D. Frame shift mutation- causes reading frame to be shifted, often caused by point insertions or point deletions In what case are missense mutations neutral? In what cases are nonsense and frameshift mutations neutral? What consequences will happen for the following three point mutations? 5’ AUG-UCA –AAC- UAU-UAA 3’ U U G CONSEQUENCES OF POINT MUTATION ON A.A. In what case are missense mutations neutral? If the amino acid that was changed wasn’t very important for the function of the protein. Also, if the amino acid was changed to a similar amino acid. In what cases are nonsense and frameshift mutations neutral? If a nonsense mutation happens near the 3’ end of the mRNA, a premature stop won’t be too bad because the code was about to stop anyway. If a frameshift mutation happens near the end of the mRNA, it may still result in a viable protein. What consequences will happen for the following three point mutations? 5’ AUG-UCA –AAC- UAU-UAA 3’ U U G For the blue, it would be a missense mutation, from Ser to Leu. For the Green, it would be a silent mutation. Both encode for Asn. For the red, it would be a nonsense mutation, from Tyr to STOP. CHROMOSOMAL MUTATION 1. Deletion 2. Duplication 3. Inversion a. pericentric- centromere included b. Paracentric- does not include centromere 4. Translocation- a change in position of chromosome segments to a different location in the genome a. Nonreciprical intrachromosomal b. Nonreciprical interchromosomal c. Reciprocal interchromosomal Deletions and duplications can result from unequal crossing over. The consequences of chromosomal mutation varies widely and are NOT missense, silent, frameshift, nonsense. Intrachromosomal means the segment is moving to the same chromosome, Interchromosomal means the segment is moving to a different chromosome MY C GENE - This gene encodes for the Myc protein which causes the cell to move from G phase to S phase (cell division). - There is a mutation that is found in some cancer cells where the promoter of the myc gene is deleted. As a result, the gene uses an alternative promoter without a repressor binding site causing the gene to be expressed constitutively. GENE REGULATION - House keeping genes: constitutively expressed - Regulated genes: regulated expression Inducible Operon Repressible Operon When an effector (inducer) binds to a repressor protein, it increases expression. When an effector (corepressor)binds to a repressor protein, it decreases expression. Negative Regulation Positive Regulation Protein binds to DNA and turns off expression Protein binds to DNA and turns on expression PROKARYOTIC REGULATION: LAC OPERON - Draw a picture of the lac operon - What do lac Z and lac Y encode? - What does lac I encode? - Is lac I regulated? - Where does the repressor bind? - What happens when lactose binds to the repressor? Is this inducible or repressible operon? - If there is no expression of the lac operon, how does lactose get in the cell? - Does the repressor protein do negative or positive regulation? - In what conditions are cAMP levels high? - What happens when cAMP binds to CAP? - Is CAP protein positive or negative regulation? - Where does CAP bind? - In terms of lactose and glucose levels, when is the lac operon expressed the most? PROKARYOTIC REGULATION: LAC OPERON - What do lac Z and lac Y encode? Lac Z encodes B-galactosidase, and lac Y encodes galactoside permease which lets lactose into the cell - What does lac I encode? The repressor protein for the lac operon - Is lac I regulated? No, lac I is a housekeeping gene and is always expressed - Where does the repressor bind? The operator - What happens when lactose binds to the repressor? Is this inducible or repressible operon? When lactose binds to the repressor, the repressor falls off the DNA and expression is increased. This is an inducible operon. - If there is no expression of the lac operon, how does lactose get in the cell? There is always a minimal expression of the lac operon so that there is always some galactoside permease to let lactose in. - Does the repressor protein do negative or positive regulation? Negative regulation - In what conditions are cAMP levels high? cAMP is high when glucose levels are low - What happens when cAMP binds to CAP? CAP binds to DNA and increases expression - Is CAP protein positive or negative regulation? positive - Where does CAP bind? The lac promoter - In terms of lactose and glucose levels, when is the lac operon expressed the most? The lac operon is expressed the most when lactose is high and glucose is low TRP OPERON - Draw the trp operon - Is the effector an inducer or corepressor? - What happens when trp binds to the repressor? - Is this an inducible or repressible operon? - Is this negative or positive regulation? - Why would the trp operon shut down when there are high levels of trp? TRP OPERON - Draw the trp operon - Is the effector an inducer or corepressor? It is a corepressor - What happens when trp binds to the repressor? When trp binds to the repressor, the repressor is able to bind to the operator and stop expression. - Is this an inducible or repressible operon? repressible - Is this negative or positive regulation? negative - Why would the trp operon shut down when there are high levels of trp? The trp operon makes proteins that make trp. If there are already high levels of trp in the cell, there is no need to make trp. REGULATION BENEFITS What are the advantages of regulating transcription? What are the advantages of regulating translation? CHROMATIN - Chromatin is the combination of DNA and proteins - Histone proteins help compact the DNA and are highly conserved across species - What kind of side chains do histones have and what are their charge? - The nucleosome: - 2 H2A, 2 H2B, 2 H3, 2 H4, and 200 bp of DNA 3 Levels of DNA packaging: 1. nucleosome 2. 30 nm fiber (facilitated by H1) 3. looped domains If DNA is in a looped domain, can it be expressed? CHROMATIN - Chromatin is the combination of DNA and proteins - Histone proteins help compact the DNA and are highly conserved across species - What kind of side chains do histones have and what are their charge? Histones have basic side chains with a positive charge that is attracted to the negative charge of DNA - The nucleosome: - 2 H2A, 2 H2B, 2 H3, 2 H4, and 200 bp of DNA 3 Levels of DNA packaging: 1. nucleosome 2. 30 nm fiber (facilitated by H1) 3. looped domains If DNA is in a looped domain, can it be expressed? Yes, looped domains are still considered euchromatin and is still relatively loosely packed. If it was packed tighter than a looped domain, it would be heterochromatin and not easily expressed. CHROMATIN CHANGES - Chromatin can be remodelled or chemically modified - Chemical modification: acetylation of histones to lysine side chains which neutralizes positive charge - Remodeling: histones are moved upstream or down stream - What kind of proteins acetylate histones? - What if a cell had inactive HDAC proteins? CHROMATIN CHANGES - Chromatin can be remodelled or chemically modified - Chemical modification: acetylation of histones to lysine side chains which neutralizes positive charge - Remodeling: histones are moved upstream or down stream - What kind of proteins acetylate histones? Histone acetyl transferase (HAT) acetylate histones and make DNA more loose and easier to translate. - What if a cell had inactive HDAC proteins? HDAC or histone deacetylaces take off the acetyl group and make the histones positively charged again to make DNA tightly packaged. Without them, DNA would remain loosely packaged and regulation would be much more difficult if not impossible. EUKARYOTIC GENE REGULATION - Eukaryotes have promoter proximal elements and promoter distal elements. - What’s the difference? - What are the elements made from, and what is their job? - Activators recruit HAT proteins or chromatin remodeling complexes. - Silencers recruit HDAC or chromatin remodeling complexes DNA Enhancers DNA Silencers Activators (positive regulation) Repressors (negative regulation) HAT Proteins HDAC proteins EUKARYOTIC GENE REGULATION - Eukaryotes have promoter proximal elements and promoter distal elements. - What’s the difference? Proximal elements are at least 500 bp upstream of the promoter, whereas distal elements are more than 500 bp upstream of the promoter or somewhere downstream of the gene. - What are the elements made of, and what is their job? The elements are made of DNA and are in the genome, they provide binding sites for either activator proteins or repressor proteins. - Activators recruit HAT proteins or chromatin remodeling complexes. - Silencers recruit HDAC or chromatin remodeling complexes DNA Enhancers DNA Silencers Activators (positive regulation) Repressors (negative regulation) HAT proteins HDAC proteins CORTISOL - Cortisol is a stress hormone that binds to a receptor in the cell’s cytoplasm. - What happens when cortisol binds to its receptor? - Cortisol increases the expression of what type of protein? Dsx protein Transferrin RNAiTranslation initiation Maternal genes Ferritin ubiquitination