* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download NeuroAnatomic and Genetic Approaches to Memory Formation
Copy-number variation wikipedia , lookup
X-inactivation wikipedia , lookup
Minimal genome wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Oncogenomics wikipedia , lookup
Public health genomics wikipedia , lookup
Genomic imprinting wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Genome evolution wikipedia , lookup
Point mutation wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Genetic engineering wikipedia , lookup
Gene therapy wikipedia , lookup
Gene desert wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Genome editing wikipedia , lookup
The Selfish Gene wikipedia , lookup
Genome (book) wikipedia , lookup
Gene nomenclature wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Gene expression programming wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Gene expression profiling wikipedia , lookup
History of genetic engineering wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
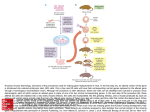
Neurobiology of Learning and Memory Prof. Stephan Anagnostaras Lecture 9: Molecular-Genetic Approaches to Learning & Memory CamKIIa and CREB are required for LTP QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Mechanisms of Learning and Memory Synaptic Kinase activation A NMDA D1 AMPA Cyclase cAMP CaMKII PDE Nuclear CREB activation Ca CREB Gene replacement and transgenic animals • Some genes are identified through mutant analysis Forward Genetics (mutant phenotype---> genotype) • To determine the function of these genes, it is possible to replace an organism’s wild type gene with an inactive gene to create a “gene knockout” Reverse Genetics (mutant genotype--->phenotype) • It is also possible to introduce additional genes (transgenes) to create a transgenic organism • Epigenetics Associate gene expression or polymorphisms with phenotype In mice, two main kinds of animals Transgenic A novel gene, the transgene, either man-made or borrowed from another animal is added to the genome. Works in many species - easy Knockout A gene is deleted; in practice the gene is replaced in the same location (homologous recombination) with a null mutation of the gene. Only possible in mice, difficult Related mutants: Dominant negative: a massively overexpressed transgene that interferes with the endogenous gene (easy) Knockin: Homologous recomination of a transgene (very difficult and rare) In vitro mutagenesis of a cloned gene Gene knockout and transgenic techniques usually involve mutagenesis of cloned genes prior to transfer into the organism Production of transgenic mice Confusing example is a dominant negative transgene (like Mayford & Kandel), not a knockout but a lot like a knockout Creation of mice embryonic stem (ES) cells carrying a knockout mutation Gene knockout in mice QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Qu ic kTime ™ an d a TIFF (U n co mpre ss ed ) d ec omp res so r a re ne ed ed to se e th is p ic tu re. Quick Time™ a nd a TIFF (U nc omp res se d) de co mpre ss or are n ee de d to s ee this picture . Qu ic kTime ™ an d a TIFF (U n co mpre ss ed ) d ec omp res so r a re ne ed ed to se e th is p ic tu re. Qu ic kTime ™ an d a TIFF (U n co mpre ss ed ) d ec omp res so r a re ne e de d to s ee this picture . Qu ic kTime ™ an d a TIFF (U n co mpre ss ed ) d ec o mpre ss or are n ee de d to s ee this picture . Qu ic kTime ™ an d a TIFF (U n co mpre ss ed ) d ec omp res so r a re ne ed ed to se e th is p ic tu re. T=Threonine (ACT, ACC, ACA) A=Alanine (GCT, GCC, GCA,GCG) Disruption of CamKIIa by T286A point mutation disrupts LTP and spatial memory •Disrupts autophosphorylation of Threonine at 286 Unable to switch to Ca-Calmodulin independent state Giese et al., Science,1998 Cell-type-specific gene knockouts in mice: Cre-Lox technique Spatial and/or temporal control of deletion Tsien et al., Cell, 1996: CA1 specific NMDAR1 knockout CA1 Tsien et al., Cell, 1996 DG McHugh et al., Cell, 1996: Place cells in CA1 NMDAR KO Abnormally large place fields Low coherent firing An Temporally controlled inducible CREB Repressor: The LBD Fusion Approach Kida, Josselyn et al., 2002 LBD Heat Shock Protein CREBr Tamoxifen LBD Cytoplasm CREBr LBD CREBr Nucleus The LBD-CREB-MT Vector (single Transgene) CaMKII promoter HA LBD CREB-MT intron polyA signal HA; influenza virus hemagglutinin (HA) tag LBD ; mutant estrogen receptor ligand binding domain (G521R) CREB-MT; dominant negative CREB (CREB S133A) Josselyn et al., 2002 - temporal control of the CREB repressor Fear conditioning Tetracycline-regulated expression of CamKIIaAsp286 (dominant negative, always phosphorylated T286D D= Aspartate, “constitutively active”) Double-transgenic • Cell-specific tTA • tetO + transgene to regulate Mayford & Kandel, TIG, 1999 D= GAT, GAC Tet-Off System Summary - Reverse Genetics •Molecular Biology Approaches Mutant Mice • Knockouts - global deletion • Transgenics - global addition Later generation: • Point mutants (PointLox method) • Region-specific deletion by Cre-Lox • Inducible deletions • Temporal and or region control of transgene - LBD fusion approach - Tet system Other Approaches - Quantitative Genetics and Functional Genomics Forward Genetic Approaches QTL (Quantitative Trait Loci) ENU (ethylnitrosurea - random mutagenesis) Dominant & recessive screens Mapping & Positional Cloning Epigenetic approaches DNA Microarrays Current mouse array = 11k genes/ 35k Correlate expression or polymorphisms with phenotype