* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download REVERSE GENETICS: USING RNAi TO MAKE PROTEIN KNOCK
Immunoprecipitation wikipedia , lookup
Gene regulatory network wikipedia , lookup
Index of biochemistry articles wikipedia , lookup
History of molecular evolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Silencer (genetics) wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Messenger RNA wikipedia , lookup
Molecular evolution wikipedia , lookup
List of types of proteins wikipedia , lookup
Magnesium transporter wikipedia , lookup
Epitranscriptome wikipedia , lookup
Expression vector wikipedia , lookup
Protein domain wikipedia , lookup
Protein design wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Protein folding wikipedia , lookup
Homology modeling wikipedia , lookup
Gene expression wikipedia , lookup
Protein structure prediction wikipedia , lookup
Protein moonlighting wikipedia , lookup
Interactome wikipedia , lookup
Protein (nutrient) wikipedia , lookup
Western blot wikipedia , lookup
RNA interference wikipedia , lookup
Protein purification wikipedia , lookup
Protein adsorption wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
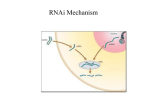
REVERSE GENETICS: USING RNAi TO MAKE PROTEIN KNOCK-DOWNS Developmental Biology Lab – Fall 2004 Using reverse genetics, one first identifies a gene of interest, and then determines what defect, if any, results when the corresponding protein is missing. This approach may be used to investigate whether a particular protein performs the same functions in one organism as a homologous protein (one with a similar sequence) does in another organism. Using a simple organism like C. elegans, one might investigate whether a C. elegans protein that is homologous to a mammalian protein has a related function. If so, C. elegans might then be used as a model system to study the normal function of the protein, the regulation of the protein, and molecular targets of its action. If the absence or mutation of the mammalian version of the protein leads to a disease, studying its C. elegans homolog might further our understanding of the molecular basis of the disease and could elucidate possible treatments. There are several different strategies for eliminating or severely depleting the expression of a particular protein, which are referred to as “knock-out” or “knock-down” strategies, respectively. C. elegans researchers, among others, employ a technique called RNA mediated interference, or RNAi. The central dogma of molecular biology is that DNA is transcribed into mRNA, and mRNA is translated into protein. Therefore, one would assume that more of a certain mRNA would result in more of that protein. This is true to a certain extent. What we now know is that if you introduce an extreme amount of a specific dsRNA into a worm, that dsRNA gets processed and leads to the degradation of the corresponding endogenous mRNA. This technique might not completely eliminate the corresponding protein, but it significantly diminishes the protein levels, thereby resulting in a phenotype that mimics a strong loss-of-function mutant. Remarkably, the mRNA can be introduced either by injection, soaking the animals in a solution of the mRNA, or even by feeding it to the animals on a plate! Each group will be responsible for performing the RNAi feeding protocol on one C. elegans gene. Due to our limited time together, the experiment will be spread out over the next four lab periods. Some of the preparatory tasks will be performed for you. SEE TABLE ON REVERSE SIDE. DATES: TASKS: Th 12/2; F 12/3 - Make M9-lactose plates for RNAi feeding protocol - Observe worms by DIC and learn basic worm anatomy - Bleach N2 hermaphrodites and allow embryos to hatch in M9 - Start overnight bacterial cultures from newly transformed plates of L4440 alone or L4440 with cDNA of interest - Seed M9-lactose plates with O.N. culture (L4440 alone or L4440 w/clone); allow to dry - Place L1 larvae on seeded plates - Observe N2 hermaphrodites with stereomicroscopes - Analyze data from RNAi feeding experiment - Characterize phenotypes using stereomicroscopes and DIC; document info - Use computer databases to find info regarding your gene/protein of interest (What type of protein? Any conserved domains?) - Use databases to determine whether humans have a homologous sequence. If so, is the function known? M 12/6 T 12/7; W 12/8 Th 12/9; F 12/10 T 12/14; W 12/15 ACCOMPLISHED BY: BIO 408 students Other BIO 408 students BIO 408 students BIO 408 students FINAL WRITTEN REPORT – 30 POINTS Developmental Biology Lab – Fall 2004 The following questions or requests should be addressed in a written report. One report per lab group of three should be submitted by 10 a.m. on Friday, December 17. The report should be typed, double-spaced, and have one-inch margins on all sides. The report grade will be based on the group’s ability to: - address all requested points - accurately and completely answer questions - demonstrate overall good writing skills; organization, topic sentences, proper grammar, correct spelling, and punctuation 1) What does “RNAi” stand for? 2) What is the purpose of RNAi? Why do researchers perform these kinds of experiments? 3) What is the cellular mechanism by which RNAi elicits its effects? 4) In your own words, briefly describe how the dsRNA gets generated and gets into the worm. 5) What protein did you deplete by RNAi and what served as the control in the experiment? 6) What phenotype(s) did you observe in comparison to the control? DOCUMENT WITH FIGURES. 7) Based on the observed phenotype, what organ system(s) might be affected? 8) What type of protein did you RNAi (i.e. a specific kinase, a cell-cycle protein, etc)? 9) Are there any conserved domains? If so, what are they and what functions are they associated with? 10) What kind(s) of cellular activities is this type of protein involved in? 11) Is there a human ortholog (a human version) of this protein? 12) If so, what is its name and what type(s) of cellular activities is it involved in? 13) Based on the results of the bioinformatics and any primary literature resources that you might have found, predict the molecular basis for the observed phenotype.