* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Plasmid Sex Introduction .....In most bacteria there are several
Oncogenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Minimal genome wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Genome evolution wikipedia , lookup
Genome (book) wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Primary transcript wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Epigenomics wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Non-coding DNA wikipedia , lookup
Point mutation wikipedia , lookup
DNA supercoil wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Designer baby wikipedia , lookup
DNA vaccination wikipedia , lookup
Genome editing wikipedia , lookup
Genetic engineering wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genomic library wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Helitron (biology) wikipedia , lookup
Molecular cloning wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Microevolution wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
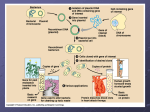
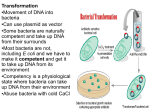
Plasmid Sex Introduction .....In most bacteria there are several pieces of DNA. .. One is the somatic genome - a huge circle of double-stranded DNA that actually measures about 2 mm in length, and is all crammed into the little cell. ..This large piece of DNA is what defines the type of bacterium it is. .. The cell cannot live without this circle of DNA. In addition, there are various optional smaller circles of DNA, which are usually called plasmids... To repeat: ..these are 'optional' and the cell can get along without them unless the genes on those plasmids allow it to survive under unusual conditions such as when a particular antibiotic is in the neighborhood, and the plasmid contains a gene that protects the cell from that antibiotic by any one of several different mechanisms. .....A cell can duplicate a plasmid and then send one copy over to another cell via a thin tube called a pilus. ..Seemingly, each type of plasmid codes for its own special type of pilus. .....A widely used plasmid of E. coli is one called "F" (for fertility). .. Cells that possess "F" are called male (F-donors or "F+"), these cells usually possess two F-pili for the transport of the F into cells lacking it. .. Those without F are called females (potential recipients, F-). .. In this exercise you will mix a very few F+ with a much larger number of F-, and show that the F- cells are converted to F+ at a rather rapid pace. ..The rapidity is because because it is a chain-reaction as the newly form F+ cells are added to the pool of donors. ..(You can change this exercise into an experiment if you can design an experiment from which you can determine the time it takes to convert an F- into an F+.) .....First, you must have a strain of donor that possesses a somatic genome that is different from that of the recipient. ..That way you can "select against" the original donor cells by using a mating medium in which they cannot grow, but in which the recipients can. ..For this we will use donors that are normal with respect to sensitivity to the antibiotic streptomycin, while the recipients will be streptomycin resistant (strR). ..We will add streptomycin to the medium, and none of the original donors will be able to grow up to form colonies. ..(Always keep in the back of your mind a very important word: "controls"!) .....Second, you must have a way to tell whether or not the recipients have acquired the F. ..We will do a little trick here, and use an F that has the lactose operon inserted into it. ..This is called "F-lac." ..(Whenever other genes are incorporated into F, the F is now called a type of F-prime (F'). ..Thus F-lac is a type of F'. ..We will therefore start with F- cells that do not have a functional lac-operon (lac-), and so when they acquire the F-lac, they will be able to use lactose sugar to grow. .. (This is not a genetically engineered product, but one that can be made naturally, and therefore doesn't not need to be approved by an institutional recombinant DNA committee. ..Indeed, no part of this exercise is considered genetic engineering. ..It is all genetic recombination using natural means. ..That is one of the important things about this exercise - this happens naturally around you all the time in the genetically mobile and fluid microbial world.) .....Back in the first decade of the 1900's, Dr. MacConkey concocted a medium bearing his name. ..On MacConkey agar (page 61), E. coli that possess the ability to ferment lactose (lac+) grow up as red colonies, while lac- mutants grow up as pale colonies. ..Thus what do we need? ..Yes! ..MacConkey agar into which we have added some streptomycin. (Controls?) THE EXERCISE 1. Make two separate overnight cultures of the donors and recipients in 0.7% tryptone (a la page 2). .. (Please don't follow the directions on the bottles! ..Use TAP-water as these little critters need minerals to grow.) .. Do NOT add any sugars to the medium! 2. Into a new sterile bottle of 20 ml of tryptone add 20 drops of the recipient F-, and 1 drop of the donor F-lac. ..(Here strict aseptic technique is not required because the product of this reaction will be plated on antibiotic plates.) 3. At timed intervals make plate counts (page 112) of the culture using the MacConkey agar PLUS streptomycin.* ..(The next day, your results should show many pale colonies on the plates, but as time proceeds there will be more and more red ones, which are clones of the recombinants.) * Making antibiotic medium is a little tricky as most antibiotics are destroyed in heat, and therefore cannot be autoclaved. ..Of course you are wondering why worry about sterilizing an antibiotic in the first place. .. Because they are not free of bacteria! ..They are either made by a type of bacteria or fungi, which will then contaminate the powder. ..Thus the antibiotic powder must be dissolved in water at the desired concentration, and then sterilized. ..The most common way to do so is by "ultra-filtration" through sterile filters that have such small holes that bacteria cannot get through while the antibiotic molecules easily fit. .. The filtrate is collected in a cool pre-sterilized container. ..The most common devices are little filters that fit onto the ends of large hypodermic syringes. .. The syringe (without the filter) is filled with the antibiotic solution, and then the sterile filter device is screwed onto the bottom of the syringe... The liquid is forced through the filter by pushing down on the syringe piston. ..The stream or drops of sterilized antibiotic solution is then allowed to flow into a sterilized bottle that can the be screw capped tight. .. A popular brand of such filter assemblies are 0.2nm AcroDisks®, and are sold through most laboratory supply houses. .. (Virus suspensions and heat labile vitamins are also sterilized in this same way.) -----------------------------------------------------------------------------------------------I WANT TO ESCAPE THIS PAGE! Mechanisms of Evolution Evolutionary stages of resistance There are several stages in the development of resistance; full antibiotic resistance is not necessarily conferred by an immediate change in the bacterial genome. Tolerance is the ability of bacteria to survive in the presence of antibiotics, but not to continue cell division. Resistance, on the other hand, is when bacteria can both survive and duplicate when antibiotics are present (Novak, 1999). For example, there is concern that vancomycin tolerant strains are now appearing, with documented cases in Streptococcus pneumoniae (Novak, 1999). Because tolerance is often a direct precursor to resistance, this observation is of serious concern. Resistance to vancomycin might leave no alternative for patients. It is hypothesized that tolerance may favor the development of resistance in bacteria. Because tolerant bacteria can survive in the presence of antibiotics, they have the opportunity to develop resistance. Since a selective pressure exists for bacteria that can survive antibiotic treatment, tolerant strains often become resistant. Thus, it is possible that tolerance may help lead to resistance by encouraging survival of cells tolerant of antibiotics. Species evolution Vertical genetic exchange is when genetic information is passed down through generations as cells divide. Horizontal genetic exchange is the "movement of genetic information between bacteria other than by descent" (Maiden, 1998). Horizontal genetic exchange is the primary mechanism of the evolution of antibiotic resistance. It is a sexual process, which can take place through conjugation, transduction, or transformation. Conjugation is the transfer of DNA from one cell to another by direct cell-cell contact. Transduction is when DNA is transferred from one cell to another by a bacteriophage. Transformation is a process in which "a DNA molecule is taken up from the external environment and incorporated into the genome" (Hartl and Jones, 1998, p308). The three processes yield DNA that can be replicated and passed on to progeny. Therefore, these three mechanisms will be described in further detail, as applied to the evolution of antibiotic resistance. Figure 2 diagrams an overview of the three processes. Figure 2: Transfer of Genetic Material through transformation, translation, and conjugation figure taken from Levy, 1998 see reference in text Conjugation Conjugation plays a large role in the spread of antibiotic resistance through bacteria. This process involves direct cell-to-cell contact of two bacterial cells, and the subsequent transfer of DNA. Conjugation can occur between species that are unrelated; for this reason, a large gene pool is available from which bacteria can exchange and acquire new genetic material. (Guiney, 1984). Sex pili make contact between the donor and the recipient cell. Once the two cell walls are in contact, this allows a mating bridge to form. The plasmid DNA in the donor, possibly containing antibiotic resistance genes, is nicked in one strand; this strand proceeds into the recipient cell by undergoing rollingcircle replication (Hartl and Jones, 1998, p316). Figure 3 demonstrates the process of rolling-circle replication. Complementary copies of the DNA are produced in both the donor and the recipient cells. Finally, the linear plasmid in the recipient becomes circular and is ligated, and then both of the cells have a copy of the plasmid. Figure 3: Rolling Circle Replication see reference in text There are certain barriers to the process of conjugation, which often play a role in the evolution of resistance. First, interactions at the cell surface are involved. Contact must occur for mating to take place. Next, foreign DNA is susceptible to host restriction modification systems which target and cleave foreign DNA. Third, the new plasmid be unable to replicate in the new host. An appropriate origin of replication in the new host may not be available, thus preventing replication. For example, transferred DNA is more likely to be stable in a new host if it contains fewer restriction enzyme sites. This makes it less likely that the DNA will be degraded by restriction enzymes which attack foreign DNA. A selective advantage exists for plasmids with fewer restriction sites. Selective advantages which aid in stability can parallel the beginnings of the evolution of strains that are resistant to antibiotics. Antibiotic resistance will spread more readily if the resistance genes can be transferred into non-resistant cells. The ability to both invade many types of cells and to be maintained in these cells are traits that are advantageous for survival and transfer of plasmids. Transduction Transduction occurs when a bacteriophage carries DNA from one species to another. When a bacteriophage destroys its current host and invades a new one, it may carry pieces of chromosomal DNA or plasmids from the previous host. An occasional phage may carry some bacterial DNA. Recombination can then occur between the phage (carrying bacterial DNA) and the new host's bacterial DNA. Transfer of DNA thus occurs from one bacterial cell to another, carried by a bacteriophage. ( Lacey, 1984 and Hartl and Jones, 1998, p308). This provides another method for the spread of resistance among bacteria. Transformation Transformation is another method of acquiring resistance. During transformation, bacterial cells take up DNA from the surrounding environment. Certain requirements exist in order for transformation to take place. First, exogenous DNA must be present in the immediate environment. Bacteria must have mechanisms that allow the DNA to be taken up through the bacterial cell walls. Also, the DNA must be incorporated into the chromosome of the host, often by homologous recombination. During homologous recombination, parts of the chromosome are replaced with related DNA (Maiden, 1998). Restriction modification systems play a role in transformation as well as in conjugation. However, it is thought that since these modification systems generate both DNA ends and smaller fragments, restriction modification may actually increase the chance of recombination with incorporated fragments. This could occur because recombination occurs more frequently if the ends are homologous. Possible origin of antibiotic resistance genes So now we have some mechanisms by which transfer and uptake of resistance can occur. But where did this resistance originate? The answer is complicated; some resistance is actually the result of random mutations that provide a selective advantage. However, there are other suspected sources of resistance. Many resistance genes may have come from bacteria in soil and water. Soil and water bacteria must live and survive along with fungi/actinomycetes, which produce antibiotics. These bacteria must possess a certain level of resistance in order to survive in their natural environment. Some organisms can exchange plasmids with those in soil and water, as well as with one another. Therefore, antibiotic resistance genes may have originated from bacteria in this environment (Guiney, 1984). When antibiotics are isolated from sources such as fungi, some resistance genes may be isolated as well. Therefore, ironically, antibiotic preparations may be contaminated with DNA encoding resistance genes. This increases the chance that genetic exchange can occur with the bacteria they are trying to kill - the close proximity increases the chance of transformation of DNA (Davies, 1994). There is also some clue into the origin of resistance to -lactam antibiotics. Resistance genes were detected even before the use of antibiotics began. For example, a strain of Staph aureus was found which produced -lactamase--before penicillin was even discovered. It is believed that lyzozyme, which is present in nasal secretions, and is a "natural bactericidal agent," played a role (Lacey, 1984). Lyzozyme may have imposed a selective pressure, which helped strains to emerge with antibiotic resistance, even before our discovery of antibiotics. Genetic mechanisms The basic mechanisms of transformation, transduction, and conjugation provide a starting point for more complex mechanisms of the development and spread of antibiotic resistance. Some common mechanisms are outlined below and illustrated in figure 4 Point mutations Point mutations are also involved in the development of antibiotic resistance. Spontaneous changes in single nucleotides have produced resistance to some antibiotics. (Lacey, 1984). Point mutations are usually random, and thus occur before exposure to antibiotics. For example, one base change in the -lactamase gene (which cleaves lactam antibiotics such as penicillin) can change this enzyme's substrate specificity (Davies, 1994). Such point mutations may cause changes in the target molecule of the antibiotic. Often however, several random point mutations must occur in order to confer resistance; therefore, this type of resistance development is somewhat less common. Intragenic recombination When smaller fragments of DNA are incorporated through transformation, intragenic recombination and mosaic genes result. Mosaic genes contain DNA of the original allele in some locations, but from different genes or organisms in other locations. This is a result of intragenic recombination. These genes may express proteins that have new phenotypes (Maiden, 1998). Most mosaic genes are lost because they represent only a low frequency of the genes. However, some mosaic genes may express a phenotype that helps the organism to survive. An example of this phenomenon is a mosaic gene encoding an altered penicillin binding protein with low affinity for -lactam antibiotics. This mosaic gene allows survival in the presence the antibiotic, and may be favored by selection. Thus, the selective pressure of antibiotics may be promoting the maintenance and spread of mosaic genes (Maiden, 1998). Transposons Transposable elements are small regions of DNA that can move from one place to another in the genome. Therefore, these play a role in evolution of antibiotic resistance as well, by providing yet another method of genetic exchange. Transposable elements that contain genes in the central region are called transposons. This central sequence may contain resistance to one or more antibiotics for example (making multiple antibiotic resistance probable). Genes in transposons can be transferred between bacterial hosts by transposition into bacterial plasmids, which can then undergo conjugation (Hartl and Jones, 1998, p347). Transposons make multiple antibiotic resistance not only possible, but very likely; this mechanism provides an easy and efficient way for the transfer of resistance to several antibiotics to be spread at one time. Figure 4: Mutations which may Confer Antibiotic Resistance see reference in text Several mechanisms exist for the transfer of resistance between bacteria. When the selective pressure of antibiotics is imposed, bacteria have a large population of resistance genes available to them (Davies, 1994). This provides an environment where the development and spread of antibiotic resistance is likely to continue indefinitely, both due to the selective pressure of antibiotics and to the large existence of resistance in the population currently. In other words, discontinuing the use of antibiotics would probably not stop the spread of resistance entirely. The selective pressure would be lower, but the resistance could still be spread throughout the population in ways such as those described above. In addition, continued use of antibiotics will only worsen the problem. For this reason, close attention to the mechanisms of antibiotic resistance is necessary in order to gain an understanding of how to develop more effective antibiotics. Plasmid From Wikipedia, the free encyclopedia (Redirected from Plasmids) Jump to: navigation, search Figure 1: Schematic drawing of a bacterium with plasmids enclosed. (1)Chromosomal DNA. (2) Plasmids Plasmids are (typically) circular double-stranded DNA molecules that are separate from the chromosomal DNA (Fig. 1). They usually occur in bacteria, sometimes in eukaryotic organisms (e.g., the 2-micrometre-ring in Saccharomyces cerevisiae). Their size varies from 1 to over 400 kilobase pairs (kbp). There are anywhere from one copy, for large plasmids, to hundreds of copies of the same plasmid present in a single cell. Contents [hide] 1 Antibiotic resistance 2 Episomes 3 Vectors 4 Types of plasmid 5 Applications of plasmids 6 Plasmid DNA extraction 7 Conformations 8 See also [edit] Antibiotic resistance Figure 2: Schematic drawing of a plasmid with antibiotic resistances Plasmids often contain genes or gene-cassettes that confer a selective advantage to the bacterium harboring them, e.g., the ability to make the bacterium antibiotic resistant. Every plasmid contains at least one DNA sequence that serves as an origin of replication or ori (a starting point for DNA replication), which enables the plasmid DNA to be duplicated independently from the chromosomal DNA (Fig. 2) [edit] Episomes Episomes are plasmids that can integrate themselves into the chromosomal DNA of the host organism (Fig. 3). For this reason, they can stay intact for a long time, be duplicated with every cell division of the host, and become a basic part of its genetic makeup. This term is no longer commonly used for plasmids, since it is now clear that a region of homology with the chromosome such as a transposon makes a plasmid into an episome. [edit] Vectors Figure 3: Comparison of non-integrating plasmids (top) and episomes (bottom). 1 Chromosomal DNA. 2 Plasmids. 3 Cell division. 4 Chromosomal DNA with integrated plasmids Plasmids used in genetic engineering are called vectors. They are used to transfer genes from one organism to another and typically contain a genetic marker conferring a phenotype that can be selected for or against. Most also contain a polylinker or multiple cloning site (MCS), which is a short region containing several commonly used restriction sites allowing the easy insertion of DNA fragments at this location. See also 'Applications of plasmids', below. [edit] Types of plasmid One way of grouping plasmids is by their ability to transfer to other bacteria. Conjugative plasmids contain so-called tra-genes, which perform the complex process of conjugation, the sexual transfer of plasmids to another bacterium (Fig. 4). Non-conjugative plasmids are incapable of initiating conjugation, hence they can only be transferred with the assistance of conjugative plasmids, by 'accident'. An intermediate class of plasmids are mobilisable, and carry only a subset of the genes required for transfer. These plasmids can 'parasitise' another plasmid, transferring at high frequency in the presence of a conjugative plasmid. It is possible for several different types of plasmids to coexist in a single cell, e.g., seven different plasmids have been found in E. coli. On the other hand, related plasmids are often 'incompatible', resulting in the loss of one of them from the cell line. Therefore, plasmids can Figure 4 : Schematic drawing of bacterial conjugation. 1 Chromosomal DNA. 2 Plasmids. 3 Pilus. be assigned into incompatibility groups, depending on their ability to coexist in a single cell. These incompatibility groupings are due to the regulation of vital plasmid functions. An obvious way of classifying plasmids is by function. There are five main classes: Fertility-(F)plasmids, which contain tra-genes. They are capable of conjugation. Resistance-(R)plasmids, which contain genes that can build a resistance against antibiotics or poisons. Historically known as R-factors, before the nature of plasmids was understood. Col-plasmids, which contain genes that code for (determine the production of) colicines, proteins that can kill other bacteria. Degrative plasmids, which enable the digestion of unusual substances, e.g., toluene or salicylic acid. Virulence plasmids, which turn the bacterium into a pathogen. Plasmids can belong to more than one of these functional groups. Plasmids that exist only as one or a few copies in each bacterium are, upon cell division, in danger of being lost in one of the segregating bacteria. Such single-copy plasmids have systems which attempt to actively distribute a copy to both daughter cells. Some plasmids include an addiction system. These plasmids produce both a long-lived poison and a short-lived antidote. Daughter cells that retain a copy of the plasmid survive, while a daughter cell that fails to inherit the plasmid dies or suffers a reduced growth-rate because of the lingering poison from the parent cell. This is an example of plasmids as selfish DNA. [edit] Applications of plasmids Plasmids serve as important tools in genetics and biochemistry labs, where they are commonly used to multiply (make many copies of) or express particular genes. There are many plasmids that are commercially available for such uses. Initially, the gene to be replicated is inserted in a plasmid. These plasmids contain, in addition to the inserted gene, one or more genes capable of providing antibiotic resistance to the bacterium that harbors them. The plasmids are next inserted into bacteria by a process called transformation, which are then grown on specific antibiotic(s). Bacteria which took up one or more copies of the plasmid then express (make protein from) the gene that confers antibiotic resistance. This is typically a protein which can break down any antibiotics that would otherwise kill the cell. As a result, only the bacteria with antibiotic resistance can survive, the very same bacteria containing the genes to be replicated. The antibiotic(s) will, however, kill those bacteria that did not receive a plasmid, because they have no antibiotic resistance genes. In this way the antibiotic(s) acts as a filter selecting out only the modified bacteria. Now these bacteria can be grown in large amounts, harvested and lysed to isolate the plasmid of interest. Another major use of plasmids is to make large amounts of proteins. In this case you grow the bacteria containing a plasmid harboring the gene of interest. Just as the bacteria produces proteins to confer its antibiotic resistance, it can also be induced to produce large amounts of proteins from the inserted gene. This is a cheap and easy way of massproducing a gene or the protein it then codes for--for example, insulin or even antibiotics. [edit] Plasmid DNA extraction As alluded to above, plasmids are often used to purify a specific sequence, since they can easily be purified away from the rest of the genome. For their use as vectors, and for molecular cloning, plasmids often need to be isolated. There are several methods to isolate plasmid DNA from bacteria, the archaetypes of which are the miniprep and the maxiprep. The former can be used to quickly find out whether the plasmid is correct in any of several bacterial clones. The yield is a small amount of impure plasmid DNA, which is sufficient for analysis by restriction digest and for some cloning techniques. In the latter, much larger volumes of bacterial suspension are grown from which a maxi-prep can be performed. Essentially this is a scaled-up miniprep followed by additional purification. This results in relatively large amounts (several ug) of very pure plasmid DNA. In recent times many commercial kits have been created to perform plasmid extraction at various scales, purity and levels of automation. [edit] Conformations When performing DNA_electrophoresis, plasmid DNA may appear in the following five conformations: "Supercoiled" (or "Covalently Closed-Circular") DNA is fully intact with both strands uncut. "Relaxed Circular" DNA is fully intact with both strands uncut, but has been enzymatically "relaxed" (supercoils removed). "Supercoiled Denatured" DNA, is not a "natural" form present in vivo. It is a contaminent often produced in small quantities following excessive alkaline lysis; both strands are uncut but are not correctly paired, resulting in a compacted plasmid form. "Nicked Open-Circular" DNA has one strand cut. "Linearized" DNA has both strands cut site at only one site. The relative electrophoretic mobility (speed) of these DNA conformations in a gel are as follows: Nicked Open Circular (slowest) Linear Relaxed Circular Supercoiled Denatured Supercoiled (fastest) The rate of migration for small linear fragments is directly proportional to the voltage applied at low voltages. At higher voltages, larger fragments migrate at continually increasing yet different rates. Therefore the resolution of a gel decreases with increased voltage. At a specified, low voltage, the migration rate of small linear DNA fragments is a function of their length. Large linear fragments (over 20kb or so) migrate at a certain fixed rate regardless of length. This is because the molecules 'reptate', with the bulk of the molecule following the leading end through the gel matrix. Restriction digests are frequently used to analyse purified plasmids. Enzymes specifically break the DNA at certain short sequences. The resulting linear fragments form 'bands' after gel electrophoresis. [edit] See also Bacterial artificial chromosome Retrieved from "http://en.wikipedia.org/wiki/Plasmid" Category: Molecular biology Views Article Discussion Edit this page History Personal tools Sign in / create account Navigation Main Page Community Portal Current events Recent changes Random article Help Contact Wikipedia Donations Search Go Search Toolbox What links here Related changes Upload file Special pages Printable version Permanent link Cite this article In other languages Česky Dansk Deutsch Español Français עברית Македонски Nederlands 日本語 Polski Português Suomi Русский Svenska Tiếng Việt 中文 This page was last modified 23:31, 9 February 2006. All text is available under the terms of the GNU Free Documentation License (see Copyrights for details). Wikipedia® is a registered trademark of the Wikimedia Foundation, Inc. Privacy policy About Wikipedia Disclaimers Your continued donations keep Wikipedia running! Bacterial artificial chromosome From Wikipedia, the free encyclopedia Jump to: navigation, search A bacterial artificial chromosome (BAC) is a DNA construct, based on a fertility plasmid, used for transforming and cloning in bacteria, usually E. coli. Its usual insert size is 150 kbp, with a range from 100 to 300 kbp. BACs are often used to sequence the genetic code of organisms in genome projects, for example the Human Genome Project. A short piece of the organism's DNA is amplified as an insert in BACs, and then sequenced. Finally, the sequenced parts are rearranged in silico, resulting in the genomic sequence of the organism. Retrieved from "http://en.wikipedia.org/wiki/Bacterial_artificial_chromosome" Category: Molecular biology Views Article Discussion Edit this page History Personal tools Sign in / create account Navigation Main Page Community Portal Current events Recent changes Random article Help Contact Wikipedia Donations Search Go Search Toolbox What links here Related changes Upload file Special pages Printable version Permanent link Cite this article In other languages Deutsch This page was last modified 14:44, 14 January 2006. All text is available under the terms of the GNU Free Documentation License (see Copyrights for details). Wikipedia® is a registered trademark of the Wikimedia Foundation, Inc. Privacy policy About Wikipedia Disclaimers Your continued donations keep Wikipedia running! Category:Molecular biology From Wikipedia, the free encyclopedia Jump to: navigation, search Molecular biology is the study of biology at a molecular level. The field overlaps with other areas of biology, particularly genetics and biochemistry. Molecular biology chiefly concerns itself with understanding the interactions between the various systems of a cell, including the interrelationship of DNA, RNA and protein synthesis and learning how these interactions are regulated. The main article for this category is Molecular biology. Subcategories There are 6 subcategories to this category. E Electrophoresis Gene expression G M Molecular biologists Molecular genetics Peptides Proteins P Articles in category "Molecular biology" There are 153 articles in this section of this category. * G cont. P cont. Molecular biology List of molecular biology topics A Affymetrix H Agarose gel electrophoresis Amplified fragment length polymorphism Antisense Bacterial artificial I chromosome Bacterial conjugation Bonnie Bassler Biochip Biopolymer Blot (biology) C-myc CDNA library Central dogma of molecular biology Chromosome Immunoelectrophoresis Immunomagnetic separation Insertion sequence International Nucleotide Sequence Database Collaboration K Protein-fragment Complementation Assay Proteinoid Pyrenoid Pyrosequencing Q C Helicase-dependent amplification Henderson limit R Hybridisation (molecular biology) Hybridization probe B Gene copy number Gene gun Gene therapy Genetic fingerprinting Genetically modified organism Knockout mouse Q-PCR RNase H RT-PCR Real-time PCR Replica plating Reporter gene Restriction digest Restriction enzyme Restriction fragment length polymorphism Restriction sites Retrotransposon Retrovirus Reverse transcriptase walking Cloning Cloning vector Competent cell Complementarity (molecular biology) Cosmid Cycling probe technology D Krüppel associated box S L Library (biology) Lipofection List of proteins List of publications in biology M DNA bank DNA computing DNA electrophoresis DNA extraction DNA ligase DNA microarray N DNA sequencing DNA topology DNA-DNA hybridisation DNase footprinting assay Dideoxynucleotides Downstream (molecular biology) N-formylmethionine Neuropeptide Nick translation Nonribosomal peptide Northern blot Nucleic acid Nucleosome O E EMBO Journal EcoRI P Edman degradation Electrophoresis Electrophoretic mobility shift assay Electroporation Endogenous retrovirus Expression profiling F F1 generation Fluorescence resonance energy Minipreparation Molecular genetics Molecular lesion Molecular modelling Optical tweezers T P element Percentage solution Phage display Phagemid Phosphodiester bonds Phosphodiesterase Pilus Plant breeding Plant virus Plasmid Polyhistidine-tag Polyketide SDS-PAGE STING Salvage synthesis Sequencing Serial Analysis of Gene Expression Shotgun sequencing Single nucleotide polymorphism Southern blot Southwestern blot Standard curve Structural biology Subcloning Superhelix Tachykinin peptides Taffazin Temperature gradient gel electrophoresis Thermal cycler Tilling Transduction (genetics) Transfection Transformation (genetics) Transgenic plants Transposase Transposon Triparental mating Two-dimensional gel electrophoresis Two-hybrid screening transfer Fluorescent in situ hybridization Fluorescent tag Functional genomics G Gel electrophoresis Gel extraction Gel filtration chromatography Gene Polymerase chain U reaction Post transcriptional gene silencing Primer (molecular biology) Protein Protein Information W Resource Protein Misfolding Cyclic Amplification Protein electrophoresis Y Protein microarray Protein tag UniProt Upstream (molecular biology) Western blot Yeast artificial chromosome Zinc finger protein Zymography Z Retrieved from "http://en.wikipedia.org/wiki/Category:Molecular_biology" Categories: Biochemistry | Biology Views Category Discussion Edit this page History Personal tools Sign in / create account Navigation Main Page Community Portal Current events Recent changes Random article Help Contact Wikipedia Donations Search Go Search Toolbox What links here Related changes Upload file Special pages Printable version Permanent link In other languages Català Deutsch Suomi Français Bahasa Indonesia Íslenska 日本語 한국어 Polski Русский Tiếng Việt This page was last modified 22:58, 29 January 2006. All text is available under the terms of the GNU Free Documentation License (see Copyrights for details). Wikipedia® is a registered trademark of the Wikimedia Foundation, Inc. Privacy policy About Wikipedia Disclaimers Your continued donations keep Wikipedia running! Category:Electrophoresis From Wikipedia, the free encyclopedia Jump to: navigation, search Electrophoresis is a method of moving charged particles through a medium by using an electric field induced by electrodes. It is also used to separate molecules with different physical characteristics using electrical charges. The main article for this category is Electrophoresis. Articles in category "Electrophoresis" There are 13 articles in this section of this category. A E P Agarose gel electrophoresis Electrophoresis Electrophoresis (disambiguation) Protein electrophoresis Serum protein electrophoresis Temperature gradient gel electrophoresis Two-dimensional gel electrophoresis S C G Capillary electrophoresis Gel electrophoresis T D I DNA electrophoresis Difference gel electrophoresis Iontocaine Iontophoresis Retrieved from "http://en.wikipedia.org/wiki/Category:Electrophoresis" Categories: Molecular biology | Analytical chemistry Views Category Discussion Edit this page History Personal tools Sign in / create account Navigation Main Page Community Portal Current events Recent changes Random article Help Contact Wikipedia Donations Search Go Search Toolbox What links here Related changes Upload file Special pages Printable version Permanent link In other languages Français This page was last modified 16:28, 25 January 2006. All text is available under the terms of the GNU Free Documentation License (see Copyrights for details). Wikipedia® is a registered trademark of the Wikimedia Foundation, Inc. Privacy policy About Wikipedia Disclaimers Your continued donations keep Wikipedia running! Category:Gene expression From Wikipedia, the free encyclopedia Jump to: navigation, search Gene expression is the multi-step process by which a gene's information is converted into the structures and functions of a cell, following the central dogma of molecular biology. The main article for this category is Gene expression. Subcategories There is 1 subcategory to this category. T Transcription factors Articles in category "Gene expression" There are 67 articles in this section of this category. A I cont. R cont. Regulatory sequence Repressor SECIS element STAT protein Shine-Dalgarno sequence Signal peptide Silencer (DNA) Sox2 Spliceosome Splicing (genetics) Activator (genetics) Alternative splicing Anisomycin Inducible gene Intron K S B Basic-helix-loophelix CCAAT box Cis-acting element Coactivator (genetics) Constitutive gene Krüppel associated box Lac operon Lac repressor MSin3 interaction domain Messenger RNA Monocistronic L C M D DNA microarray Dicistronic N E Enhancer Exon trapping O Expressed sequence tag Expression vector G P Gene expression Gene regulatory network Gene silencing Genetic code H Nanog (transcription factor) Noncoding DNA Oct-4 Operon T Paramutation U Polyadenylation Polycistronic Posttranslational modification Preliminary messenger Z RNA Pribnow box TATA box Three prime untranslated region Trans-acting factor Transcription (genetics) Transcription Factor II D Transcription factor Transfer RNA Translation (genetics) Translational errors Transvection (genetics) Upstream transcription factor Housekeeping gene Hypoxia inducible factors I Promoter Protein biosynthesis Protein synthesis RNA interference Regulation of gene expression Zif268 R Icsbp Imprinting (genetics) Retrieved from "http://en.wikipedia.org/wiki/Category:Gene_expression" Categories: Biochemistry | Molecular genetics | Molecular biology | Cell biology Views Category Discussion Edit this page History Personal tools Sign in / create account Navigation Main Page Community Portal Current events Recent changes Random article Help Contact Wikipedia Donations Search Go Search Toolbox What links here Related changes Upload file Special pages Printable version Permanent link This page was last modified 07:04, 15 September 2004. All text is available under the terms of the GNU Free Documentation License (see Copyrights for details). Wikipedia® is a registered trademark of the Wikimedia Foundation, Inc. Privacy policy About Wikipedia Disclaimers Your continued donations keep Wikipedia running! Category:Molecular genetics From Wikipedia, the free encyclopedia Jump to: navigation, search Molecular genetics is the field of biology which studies the structure and function of genes at a molecular level. Molecular genetics employs the methods of genetics and molecular biology. The main article for this category is Molecular genetics. Subcategories There are 4 subcategories to this category. D DNA repair DNA replication Gene expression Repetitive DNA sequences G R Articles in category "Molecular genetics" There are 55 articles in this section of this category. * C cont. M cont. Mitotic crossover Molecular clock Non-coding RNA Nuclear localization signal Nucleotide diversity Open reading frame PMal-C2 RNA interference RNA polymerase Molecular genetics A CpG site D N AP site Array comparative genomic hybridization Auxotrophy E B Biopolymer C DNA barcoding DNA repair Dyad symmetry Electrophoretic mobility shift assay Euchromatin G C7 protein C7.GAT protein CAAT box CCAAT box Central dogma of molecular biology O P Gene duplication Gene family Gene gun Genetic code Genetic engineering Genetic screen R Chromatid Chromatin Chromosomal crossover Coactivator (genetics) Comparative genomic hybridization Conservation (genetics) Consortium for the Barcode of Life CpG island Germinal choice technology Recombinant DNA Retrotransposon Reverse genetics Sigma factor Site-directed mutagenesis Sp1 (biology) Sp1C Synthetic biology H S Heterochromatin I Imprinting (genetics) Intein K T Krüppel associated box Transcription Factor II D M MSin3 interaction domain Mitochondrial genome Retrieved from "http://en.wikipedia.org/wiki/Category:Molecular_genetics" Categories: Genetics | Molecular biology | Biotechnology Views Category Discussion Edit this page History Personal tools Sign in / create account Navigation Main Page Community Portal Current events Recent changes Random article Help Contact Wikipedia Donations Search Go Search Toolbox What links here Related changes Upload file Special pages Printable version Permanent link This page was last modified 12:58, 21 October 2004. All text is available under the terms of the GNU Free Documentation License (see Copyrights for details). Wikipedia® is a registered trademark of the Wikimedia Foundation, Inc. Privacy policy About Wikipedia Disclaimers Bugs Fighting Back Basics of Bacterial Resistance How Bacteria Become Resistant Once it was thought that antibiotics would help us wipe out forever the diseases caused by bacteria. But the bacteria have fought back by developing resistance to many antibiotics. Bacterial resistance to antibiotics can be acquired in four ways: 1. Through a spontaneous change in the bacterium's DNA: Changes like this are called mutations. Mutations happen at random in all living things, and they can result in all kinds of changes in the bacterium. Antibiotic resistance is just one of many changes that can result from a random mutation. 2. Transformation: This happens when one bacterium takes up some DNA from the chromosomes of another bacterium. 3. Plasmid exchange: Antibiotic resistance can be coded for in the DNA found in a small circle known as a plasmid in a bacterium. The plasmids can randomly pass between bacteria (usually touching). 4. Sharing of mutations, some of which control resistance to antibiotics. Two examples are: Plasmid transfer between different kinds of bacteria. This can happen between similar bacteria and between very dissimilar bacteria. Gene cassettes are a small group of genes that can be added to a bacterium's chromosomes. The bacteria can then accept a variety of gene cassettes that give the bacterium resistance to a variety of antibiotics. The cassettes also can confirm resistance against disinfectants and pollutants. The acquired genetically based resistance is permanent and inheritable through the reproductive process of bacteria, called fission. Some bacteria produce their own antibiotics to protect themselves against other microorganisms. Of course, a bacterium will be resistant to its own antibiotic! But sometimes the DNA that gives that bacterium resistance to its own antibiotic can be transferred to a bacterium of another species. Then that other bacterium could be resistant to the first bacterium's antibiotic! Scientists think, but haven't proved, that the genes for resistance in this case have been transferred between bacteria of different species through plasmid or cassette transfer. Laboratory analysis of commercial antibiotic preparations has shown that they contain DNA from antibiotic-producing organisms. The DNA includes the antibiotic-resistance gene sequence. Genetic transfer may be induced by the bacteria involved, that is the source and the destination bacteria. One model suggests that when a DNA resistance plasmid released by one bacterium is accepted by a different species of bacterium, the recipient may be stimulated to release its own plasmid. The process is known as retrotransfer. Resistant genes occur not only in bacteria that carry disease, but also in commensal bacteria (those living within the same environment—soil, water, digestive tract—benefiting from each other). Eating meat or milk from animals that have been exposed to antibiotics, or plants that have been exposed to pesticides, brings the antibiotic and/or resistant bacteria in contact with bacteria in your digestive tract. The interaction between bacteria can then allow for transfer of genes for antibiotic resistance to the bacteria in your intestines. How Resistance Works Some mechanisms for resistance include: o o o o o Changing the target molecule: For example, if the antibiotic attacks a certain enzyme in a bacterium, the bacterium can adapt by using a different enzyme to accomplish the same function. Enzymatically inactivating or decomposing the antibiotic. Sequestering (storing) the drug by creating alternative pathways within the bacterium. Preventing the drug from entering the bacterium. Pumping out the antibiotic as quickly as it enters the bacterium. Recommended Actions for Consumers and Physicians to Limit Resistance (from Levy, S.B. Scientific American, March 1998; p 53.) Consumers: o o o o o Do not demand antibiotics. Never use antibiotics unless they are prescribed by your doctor. When given antibiotics, take them exactly as prescribed and complete the full course of treatment: continue taking the antibiotic even after you start to feel well, and do not hoard pills for later use. Wash fruits and vegetables thoroughly; avoid raw eggs and undercooked meat, especially in ground form. Avoid antibacterial soaps and other products unless you are caring for a sick person whose defenses are weakened. Physicians (Who knows? You might be a doctor someday!): o o Wash hands thoroughly between patient visits. Do not give in to patients' demands for unneeded antibiotics. o o o When possible, prescribe antibiotics that target only a narrow range of bacteria. Isolate hospital patients with multidrug-resistant infections. Familiarize yourself with local data on antibiotic resistance. For Further Reading 20. Nordenberg, Tamar. “Miracle Drugs vs. Superbugs,” FDA Consumer, November/December 1998; pp 22-25. 21. Radetsky, Peter “Last Days of the Wonder Drugs,” Discover, November 1998; pp 76-85. 22. Levy, S.B. “The Challenge of Antibiotic Resistance,” Scientific American, March 1998; pp. 46-53. 23. Miller, R.V. “Bacterial Gene Swapping in Nature,” Scientific American, January 1998. For more information, at other Web sites... Agricultural Antibiotic Use Could Contribute to Drug Resistance — from Scientific American, 23 April 2002. The Alliance for the Prudent Use of Antibiotics (APUA) — official site with information for patients and health care professionals. Ribosome Research Encompasses Early Life on Earth and Antibiotic Resistance — news article from Argonne National Laboratory, 26 October 2001. The Rise of Antibiotic-Resistant Infections — from FDA Consumer, September 1995, published by the U.S. Food and Drug Administration. Back to: Antibiotics in Action Directory | Site Map | Pharmaceutical Achievers Home References 24. Devitt, Terry, ed. Microbes: What Doesn't Kill them Makes them Stronger, The Why Files, University of Wisconsin, May 1997. 25. Hawkey, Peter M. “The Origins and Molecular Basis of Antibiotic Resistance,” British Medical Journal, September 5, 1998, pp. 657-660. 26. Levy, S.B. "The Challenge of Antibiotic Resistance," Scientific American, March 1998; pp. 46-53. Copyright ©2002 The Chemical Heritage Foundation Recombinant DNA and Gene Cloning Recombinant DNA is DNA that has been created artificially. DNA from two or more sources is incorporated into a single recombinant molecule. Index to this page An Overview Plasmids An Example o pAMP o pKAN o Ligation Possibilities Transforming E. coli Cloning other Genes Recombinant DNA products for human therapy Making Recombinant DNA (rDNA): An Overview Treat DNA from both sources with the same restriction endonuclease (BamHI in this case). BamHI cuts the same site on both molecules 5' GGATCC 3' 3' CCTAGG 5' The ends of the cut have an overhanging piece of single-stranded DNA. These are called "sticky ends" because they are able to base pair with any DNA molecule containing the complementary sticky end. In this case, both DNA preparations have complementary sticky ends and thus can pair with each other when mixed. DNA ligase covalently links the two into a molecule of recombinant DNA. To be useful, the recombinant molecule must be replicated many times to provide material for analysis, sequencing, etc. Producing many identical copies of the same recombinant molecule is called cloning. Cloning can be done in vitro, by a process called the polymerase chain reaction (PCR). Here, however, we shall examine how cloning is done in vivo. Cloning in vivo can be done in unicellular prokaryotes like E. coli unicellular eukaryotes like yeast and in mammalian cells grown in tissue culture. In every case, the recombinant DNA must be taken up by the cell in a form in which it can be replicated and expressed. This is achieved by incorporating the DNA in a vector. A number of viruses (both bacterial and of mammalian cells) can serve as vectors. But here let us examine an example of cloning using E. coli as the host and a plasmid as the vector. Plasmids Plasmids are molecules of DNA that are found in bacteria separate from the bacterial chromosome. They: are small (a few thousand base pairs) usually carry only one or a few genes are circular have a single origin of replication Plasmids are replicated by the same machinery that replicates the bacterial chromosome. Some plasmids are copied at about the same rate as the chromosome, so a single cell is apt to have only a single copy of the plasmid. Other plasmids are copied at a high rate and a single cell may have 50 or more of them. Genes on plasmids with high numbers of copies are usually expressed at high levels. In nature, these genes often encode proteins (e.g., enzymes) that protect the bacterium from one or more antibiotics. Plasmids enter the bacterial cell with relative ease. This occurs in nature and may account for the rapid spread of antibiotic resistance in hospitals and elsewhere. Plasmids can be deliberately introduced into bacteria in the laboratory transforming the cell with the incoming genes. An Example (courtesy of David Miklos and Greg Freyer of the Cold Spring Harbor Laboratory, who used these plasmids as the basis of a laboratory introduction to recombinant DNA technology that every serious biology student — high school or college — should experience!) pAMP 4539 base pairs a single replication origin a gene (ampr)conferring resistance to the antibiotic ampicillin (a relative of penicillin) a single occurrence of the sequence 5' GGATCC 3' 3' CCTAGG 5' that, as we saw above, is cut by the restriction enzyme BamHI a single occurrence of the sequence 5' AAGCTT 3' 3' TTCGAA 5' that is cut by the restriction enzyme HindIII Treatment of pAMP with a mixture of BamHI and HindIII produces: a fragment of 3755 base pairs carrying both the ampr gene and the replication origin a fragment of 784 base pairs both fragments have sticky ends pKAN 4207 base pairs a single replication origin a gene (kanr) conferring resistance to the antibiotic kanamycin. a single site cut by BamHI a single site cut by HindIII Treatment of pKAN with a mixture of BamHI and HindIII produces: a fragment of 2332 base pairs a fragment of 1875 base pairs with the kanr gene (but no origin of replication) both fragments have sticky ends These fragments can be visualized by subjecting the digestion mixtures to electrophoresis in an agarose gel. Because of its negatively-charged phosphate groups, DNA migrates toward the positive electrode (anode) when a direct current is applied. The smaller the fragment, the farther it migrates in the gel. Ligation Possibilities If you remove the two restriction enzymes and provide the conditions for DNA ligase to do its work, the pieces of these plasmids can rejoin (thanks to the complementarity of their sticky ends). Mixing the pKAN and pAMP fragments provides several (at least 10) possibilities of rejoined molecules. Some of these will not produce functional plasmids (molecules with two or with no replication origin cannot function). One interesting possibility is the joining of the 3755-bp pAMP fragment (with ampr and a replication origin) with the 1875-bp pKAN fragment (with kanr) Sealed with DNA ligase, these molecules are functioning plasmids that are capable of conferring resistance to both ampicillin and kanamycin. They are molecules of recombinant DNA. Because the replication origin, which enables the molecule to function as a plasmid, was contributed by pAMP, pAMP is called the vector. Transforming E. coli Treatment of E. coli with the mixture of religated molecules will produce some colonies that are able to grow in the presence of both ampicillin and kanamycin. A suspension of E. coli is treated with the mixture of religated DNA molecules. The suspension is spread on the surface of agar containing both ampicillin and kanamycin. The next day, a few cells — resistant to both antibiotics — will have grown into visible colonies containing billions of transformed cells. Each colony represents a clone of transformed cells. However, E. coli can be simultaneously transformed by more than one plasmid, so we must demonstrate that the transformed cells have acquired the recombinant plasmid. Electrophoresis of the DNA from doubly-resistant colonies (clones) tells the story. Plasmid DNA from cells that acquired their resistance from a recombinant plasmid only show only the 3755-bp and 1875-bp bands (Clone 1, lane 3). Clone 2 (Lane 4) was simultaneous transformed by religated pAMP and pKAN. (We cannot tell if it took up the recombinant molecule as well.) Clone 3 (Lane 5) was transformed by the recombinant molecule as well as by an intact pKAN. Cloning other Genes The recombinant vector described above could itself be a useful tool for cloning other genes. Let us assume that within its kanamycin resistance gene (kanr) there is a single occurrence of the sequence 5' GAATTC 3' 3' CTTAAG 5' This is cut by the restriction enzyme EcoRI, producing sticky ends. If we treat any other sample of DNA, e.g., from human cells, with EcoRI, fragments with the same sticky ends will be formed. Mixed with EcoRI-treated plasmid and DNA ligase, a small number of the human molecules will become incorporated into the plasmid which can then be used to transform E. coli. But how to detect those clones of E. coli that have been transformed by a plasmid carrying a piece of human DNA? The key is that the EcoRI site is within the kanr gene, so when a piece of human DNA is inserted there, the gene's function is destroyed. All E. coli cells transformed by the vector, whether it carries human DNA or not, can grow in the presence of ampicillin. But E. coli cells transformed by a plasmid carrying human DNA will be unable to grow in the presence of kanamycin. So, Spread a suspension of treated E. coli on agar containing ampicillin only grow overnight with a sterile toothpick transfer a small amount of each colony to an identified spot on agar containing kanamycin (do the same with another ampicillin plate) Incubate overnight All those clones that continue to grow on ampicillin but fail to grow on kanamycin (here, clones 2, 5, and 8) have been transformed with a piece of human DNA. Some recombinant DNA products being used in human therapy Using procedures like this, many human genes have been cloned in E. coli or in yeast. This has made it possible — for the first time — to produce unlimited amounts of human proteins in vitro. Cultured cells (E. coli, yeast, mammalian cells) transformed with the human gene are being used to manufacture: insulin for diabetics factor VIII for males suffering from hemophilia A factor IX for hemophilia B human growth hormone (GH) erythropoietin (EPO) for treating anemia three types of interferons several interleukins granulocyte-macrophage colony-stimulating factor (GM-CSF) for stimulating the bone marrow after a bone marrow transplant granulocyte colony-stimulating factor (G-CSF) for stimulating neutrophil production, e.g., after chemotherapy and for mobilizing hematopoietic stem cells from the bone marrow into the blood. tissue plasminogen activator (TPA) for dissolving blood clots adenosine deaminase (ADA) for treating some forms of severe combined immunodeficiency (SCID) angiostatin and endostatin for trials as anti-cancer drugs parathyroid hormone leptin hepatitis B surface antigen (HBsAg) to vaccinate against the hepatitis B virus Many more examples are in the pipeline. Welcome&Next Search 12 February 2006 Bacterial Transformation Introduction: This is a very basic technique that is used on a daily basis in a molecular biological laboratory. The purpose of this technique is to introduce a foreign plasmid into a bacteria and to use that bacteria to amplify the plasmid in order to make large quantities of it. This is based on the natural function of a plasmid: to transfer genetic information vital to the survival of the bacteria. The plasmid: A plasmid is a small circular piece of DNA (about 2,000 to 10,000 base pairs) that contains important genetic information for the growth of bacteria. In nature, this information is often a gene that encodes a protein that will make the bacteria resistant to an antibiotic. Plasmids probably came about as a result of bacteria evolving in close proximity to other heterotrophs. Bacteria often grow in the same environment as molds and fungi and compete with them for food (complex organic material). As a result, molds and fungi have evolved to make toxins that kill bacteria (which we now use as antibiotics in medicine) in order to win in the competition for food. Bacteria, in turn, evolved to make proteins that inactivate the toxins. The bacteria share this vital information by passing it among themselves in the form of genes in plasmids. Plasmids were discovered in the late sixties, and it was quickly realized that they could be used to amplify a gene of interest. A plasmid containing resistance to an antibiotic (usually ampicillin) is used as a vector. The gene of interest is inserted into the vector plasmid and this newly constructed plasmid is then put into E. coli that are sensitive to ampicillin. The bacteria are then spread over a plate that contains ampicillin. The ampicillin provides a selective pressure because only bacteria that have acquired the plasmid can grow on the plate. Therefore, as long as you grow the bacteria in ampicillin, it will need the plasmid to survive and it will continually replicate it, along with your gene of interest that has been inserted to the plasmid. There are many different kinds of plasmids commercially available. All of them contain 1) a selectable marker (i.e., a gene that encodes for antibiotic resistance), 2) an origin of replication (which is used by the DNA making machinery in the bacteria as the starting point to make a copy of the plasmid) and 3) a multiple cloning site. The multiple cloning site has many restriction enzyme sites (to be discussed in a later lab) and is used to insert the DNA of interest. The multiple cloning site is usually in the middle of a reporter gene like Lac Z. A commonly used plasmid is pBluescript: Figure 1 The main differences among commercially available plasmids are the number of restriction enzyme sites, their order in the multiple cloning site, the type of antibiotic resistance that the plasmid confers, and some other genetic information that makes the plasmid useful for a specific purpose. Bacteria transformed with pBluescript will survive in ampicillin containing media and will replicate the plasmid, including any gene that is placed in the multiple cloning site. Competent Cells: Since DNA is a very hydrophilic molecule, it won't normally pass through a bacterial cell's membrane. In order to make bacteria take in the plasmid, they must first be made "competent" to take up DNA. This is done by creating small holes in the bacterial cells by suspending them in a solution with a high concentration of calcium. DNA can then be forced into the cells by incubating the cells and the DNA together on ice, placing them briefly at 42oC (heat shock), and then putting them back on ice. This causes the bacteria to take in the DNA. The cells are then plated out on antibiotic containing media. For a short animation on E. coli transformation click here. Competency The procedure to prepare competent cells can sometimes be tricky. Bacteria aren't very stable when they have holes put in them, and they die easily. A poorly performed procedure can result in cells that aren't very competent to take up DNA. A wellperformed procedure will result in very competent cells. The competency of a stock of competent cells is determined by calculating how many E. coli colonies are produced per microgram (10 -6 grams) of DNA added. An excellent preparation of competent cells will give ~108 colonies per ug. A poor preparation will be about 10 4 / ug or less. Our preps should be in the range of 10 5 to 10 6. In this experiment you will be making competent cells, transforming them with a plasmid and calculating their competency. There will be a lab report due for this lab. Procedure: Important This procedure must be performed under sterile conditions. Use only autoclaved plasticware and always work with a flame in front of you. Also, bacteria are very labile in high calcium, so keep the bacteria on ice and away from the flame at all times to keep them viable. Competent Cells: 1. (This step will be performed for you before you come into lab). Pick a single colony from a freshly grown plate of E. coli and disperse it in 100 ml of LB media in a 1 L flask. Incubate the culture at 37oC with vigorous shaking for approximately 3 hours. Cell density is monitored by determining OD600 and should be less than 10 8 cells / ml (log phase of growth - the most healthy bacteria). 2. Transfer 50 ml of this culture to a 50 ml conical tube and centrifuge at 2,000 rpm for 10 min. (Sorvall HS-4 rotor) in room 312. 3. Decant the supernatant into waste beaker (this must be sterilized before being dumped down the drain). Resuspend the pellet in 10 ml of ice cold 0.1 M CaCl2. This is most easily done by resuspending in 1 ml, using the P1000 pipette and then adding 9.0 ml. Cut about 0.5 cm from the end of the blue tip before pulling E. coli through it, since the cells are fragile in this high calcium solution and may lyse if sheared. After they have been resuspended, centrifuge at 2,000 rpm for 10 min. (Sorvall HS-4 rotor). 4. Decant the supernatant into waste beaker (to be autoclaved later). Resuspend the pellet in 1.0 ml of ice cold 0.1 M CaCl2 Bacterial Transformation: -3 1. Pipette 200ul competent cells into each of 3 ice cold Eppendorf tubes. Label the tubes Control, 1 ng, and 10 ng (1 ng is 10 ug, or -9 10 milligrams). The unknown plasmid is at a concentration of 1 ng/ul. Add 1 ng of your unknown plasmid to one tube and 10 ng to the other. Place the tubes on ice for 30 min. o 2. Put the tubes at 42 C for exactly 90 seconds. Return the cells to ice for 1-2 minutes. 3. Pipette the transformation mixtures onto labeled plates containing ampicillin and spread them around using a sterilized, bent glass rod spreader. o 4. Place upside down in the 37 C incubator in room 305 overnight. 5. 16 - 20 hours later, count the number of colonies on the plate with well-isolated colonies. Put parafilm around the edge of a plate and put it in a refrigerator for later use. Check the control plate to see that no colonies grew on it. Dispose of the plate and the control plate in the biohazard bag. Calculations: To calculate the competency of your cells, divide the number of colonies on your plate by the amount of DNA (in ug) you added to the transformation. Back to Lab Syllabus Back to Home Page If you have questions please send me a message at my Email address