* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Microarray Image Data Analysis
Epigenetics in learning and memory wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Gene desert wikipedia , lookup
X-inactivation wikipedia , lookup
Heritability of IQ wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Metagenomics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Public health genomics wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Gene expression programming wikipedia , lookup
Oncogenomics wikipedia , lookup
History of genetic engineering wikipedia , lookup
Pathogenomics wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Designer baby wikipedia , lookup
Essential gene wikipedia , lookup
Genome evolution wikipedia , lookup
Microevolution wikipedia , lookup
Genome (book) wikipedia , lookup
Genomic imprinting wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Ridge (biology) wikipedia , lookup
Minimal genome wikipedia , lookup
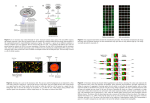
Gene Discovery from Microarray Images 陳朝欽、 高成炎、張春梵 ARCNTU, NTU-Hospital [email protected] [email protected] Project#: 93-EC-17-A-19-S1-0016 Motivation and Data Acquisition • Parts of our current works attempt to investigate and discover “a subset of genes” related to some specific diseases such as Hepatoma and Gastric Cancers by microarray experiments. Hence, we collect data from cDNA microarray images which are “spot signal intensities” via a sequence of biological experiments A Paradigm for Microarray Image Data Analysis Outline • • • • • • Microarray Image Data Acquisition Gridding for Image Segmentation Normalization from MA-Plot Finding Differentially Expressed Genes Finding Discriminative Genes Performance Evaluation by Dendrogram and K-means Algorithms A Look at a Microarray Slide Examples of Microarray Images Gridding for Spot Segmentation Gridding for a Block of 30*9 Spots Spot Feature Computation • Cy3 (for Column 1) 639 54879 5980 1984 324 910 2153 236 • Cy5 (for Column 6) 104 52858 567 189 36 1489 5083 407 M-A plot and Piecewise Normalization Normalized Ratio from MA-Plot Pre-Processing / Normalization • Due to the process of measurements or some unavoidable factors, “Raw Data” directly collected from experiments may contain noise and may have different scales, or have missing items. Thus, a pre-processing step for filtering out some inappropriate data, or normalization may be done. Spot Features for Gene Discovery Cy3 201 520 28276 4072 14807 1058 572 Cy5 67 153 21747 6324 690 1451 524 M=(log2Cy3 − log2Cy5) A= (log2Cy3+log2Cy5)/2 Program compustt.c computes spot features and pieceline.c does normalization and maplot.c does M-A plot Microarray Pattern Analysis • Microarrays consisting of 13574 effected genes from 18564 in a chip with tumor dyed in Cy3 and normal dyed in Cy5 • 12 HCV, 27 HBV, 1 HCV+HBV, 4 neither HCV nor HBV patients • Criterion for Differentially Expressed is defined as log2(Lowess normalized ratio of Cy3/Cy5) is greater than T (↑) or less than -T (↓) Feature Selection/Extraction (1) • Given a set of N patterns from K categories (K=2, a problem of dichotomy) with Ni , 1≤ i ≤ K, patterns belonging to category i, each pattern consists of M redundant features, e.g., a microarray can be represented as a pattern consisting of 13574 features corresponding to 13574 effected genes. The goal is to select a small subset of features for “Recognition” Feature Selection/Extraction (2) • Given a set of N patterns from K categories (K=2, a problem of dichotomy) with Ni , 1≤ i ≤ K, patterns belonging to category i. The goal of extraction is to transform an M-dimensional pattern into an m-dimensional pattern with m<<M for classification. A selected feature preserves the original meaning but an extraction usually does not preserve the original one. 16 Most Discriminative Genes to distinguish HCV from HBV [YCT39] Index 13796 7197 2918 8495 11189 11087 9443 9546 Accession# U35376 BG259957 BI520001 AJ012159 AB008549 BC006496 CAC51145 X52125 Index 16144 16496 17213 14579 587 113 17215 16760 Accession# AK024601 Y00083 BC007437 BC011568 AF386492 Y16961 AF195766 AI022747 Next 16 Most Discriminative Genes to distinguish HCV from HBV Index 5947 4885 11291 1262 8055 10965 4164 8088 Accession# BG207354 AK021818 AF155110 BI861005 AJ224741 AAF36120 NM_000423 BC000187 Index 7353 5434 12727 14993 4182 5341 10052 8140 Accession# AF070641 AB050785 AB062987 AA974308 AI970531 X65882 AB011542 AK026068 32 Discriminative Genes by Fisher’s Ratios for a Dendrogram 32 Discriminative Genes by Chuang+Kao’s for a Dendrogram Dendrogram from Chen’s 32 Most Discriminative Genes [CC39] Dendrogram from Genasia’s 32 Most Discriminative Genes K-means Clustering Results by using 32 Best Discriminative Genes • G45 from Genasia: distortion 341.26 1222221222 2211111111 111111111111111111 • X47 from C. Chen: distortion 302.33 1222221222 2211111111 112111111111111111 • Y48 by Fisher’s Ratio on YCT39: distortion 307.49 1222221222 2211111111 112111111111111111 • PY50 by Chuang+Kao’s on YCT39: distortion 290.06 2222222222 2211211111 112111111111111111 Leave-one-out errors by 1-nn : 4, 3, 2, 1 (/39) Leave-one-out errors by Fisher : 15, 7, 8, 9 (/39) Up (Down) Regulated Genes for Gastric Cancers • 5 Advanced and 5 Early Stage of Patients with Gastric Cancer • We find the following genes which can completely discriminate Patients of “Advanced Stage” from “Early Stage” under clinical diagnosis Dengrogram for Gastric Patients Top 16 Discriminative Genes for Advanced and Early Stages Index 15843 12994 18370 2070 1118 9661 2017 1128 Accession# AF316855 BF868865 BC002996 AK021788 BC000249 AP000350 U53530 AF035281 Index 8728 494 10990 342 10425 6052 170 1016 Accession# AL591713 AB014526 L77570 BC007848 BG745129 AF073362 AK000278 BF526386 Thank You • http://www.bioinfo.ntu.edu.tw • http://www.cs.nthu.edu.tw/~cchen • Tel: (02) 2312 3456 ~ 5917 • Tel: (02) 2362 5336 ~ 418 • Tel: (03) 573 1078