* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download HGSS Chapters 11 & 12: Modern Gene Hunting (incomplete)
Epigenetics of diabetes Type 2 wikipedia , lookup
DNA supercoil wikipedia , lookup
Minimal genome wikipedia , lookup
Human genetic variation wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Gene desert wikipedia , lookup
Public health genomics wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Gene expression profiling wikipedia , lookup
Genetic engineering wikipedia , lookup
Human genome wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
SNP genotyping wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Y chromosome wikipedia , lookup
Non-coding DNA wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Neocentromere wikipedia , lookup
Genomic imprinting wikipedia , lookup
Genomic library wikipedia , lookup
Genetic drift wikipedia , lookup
Point mutation wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Quantitative trait locus wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Gene expression programming wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
History of genetic engineering wikipedia , lookup
X-inactivation wikipedia , lookup
Genome (book) wikipedia , lookup
Genome editing wikipedia , lookup
Population genetics wikipedia , lookup
Genome evolution wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Helitron (biology) wikipedia , lookup
Genome-wide association study wikipedia , lookup
Designer baby wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
HLA A1-B8-DR3-DQ2 wikipedia , lookup
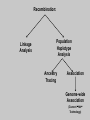
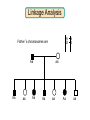
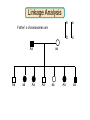
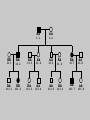
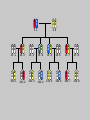
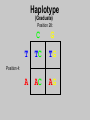
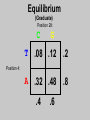
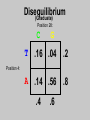
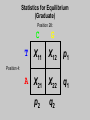
Gene Hunting: Linkage and Association We humans are diploid (i.e., we have two copies of a gene), inheriting one chromosome from mother, the other from father. In transmitting a chromosome to an offspring, however, the physical process of recombination (crossing over) results in a chromosome that contains part of the maternal chromosome and part of the paternal chromosome. Recombination also makes possible a number of different analytical strategies in genetics: linkage, ancestry tracing, and some forms of association. Key terms: polymorphism, recombination, crossing over, linkage, linkage analysis, association design, haplotype, linkage equilibrium/disequilibrium, GWAS (genome-wide association study). Recombination (Crossing Over) In meiosis, homologous chromosomes join together at a section and exchange genetic material. Homologous chromosomes: chromosomes with the same genes on them. E.g., your paternal chromosome number 1 and your maternal chromosome number 1. Recombination: Linkage Analysis Population Haplotype Analysis Ancestry Tracing Association Genome-wide Association (Current “Hot” Technology) Example: A a b B A a b B A a b B C c d D c C D d c C D d Key Point about Recombination: Recombination is a function of physical distance. • If two alleles are separated by 8 nucleotides, then there are “8 chances” of a recombination event between the two.. • If two alleles are separated by 257 nucleotides, then then are “257 chances” of a recombination event between the two. • Therefore, alleles on the same DNA strand that are far away are more likely to be broken up by recombination than alleles that are close together. Original Chromosomes: Allele 1 Allele 2 New Chromosomes: Dad Mom Pair Up A T C G G C T A G C C T G A C A T T A T T G G C T A G C C T G A C G A T A A T T C T G G G G C CC C T TT T A AA A G G C C C C T T G G A A C C A G T A T T Exchange Material A T C G G C T A G C C T G A C G A T A T T G G C T A G C C T G A C A T T A T C G G C T A G C C T G A C A T T G C A T C 3 chances G G C 10 chances T A G C C T G A C A T T G C 17 chances In other words: Alleles close together on the same DNA strand (i.e., the same chromosome) tend to be transmitted as a unit. Alleles far away on the same DNA strand tend to be broken up. Definitions: Linkage: Biological phenomenon that close to one another tyend to transmitted as a unit. Linkage Analysis: (1) tracing the co-segregation of (2) one or more marker genes with a trait gene within pedigrees (3) within families Definitions: Trait gene: A gene that contributes to the trait of interest, e.g., schizophrenia. Marker Gene: A polymorphic “gene” with that does not contribute to the trait but has a known location in the genome. Rationale for Linkage Analysis Can I predict who gets the disorder (trait) by knowing the marker genes in a family? YES: A trait gene is close to a marker. NO: No trait genes are close to the marker. Linkage Analysis A D Father’s chromosomes are aa Aa Aa aa Aa a d Aa aa Aa aa Linkage Analysis A a D d Father’s chromosomes are aa Aa Aa aa Aa Aa aa Aa aa Aa I.1 AA Aa Aa II.1 II.2 II.3 Aa AA Aa III.1 III.2 III.3 aa aa I.2 aa Aa Aa II.4 II.5 II.6 II.7 aa aa Aa Aa III.4 III.5 III.6 III.7 aa II.8 aa III.8 Aa aa I.1 I.2 D d d d A A Aa Aa aa aa Aa Aa a a II.1 II.2 II.3 II.4 II.5 II.6 II.7 II.8 A a AA A a aa a a Aa A a aa III.1 III.2 III.3 III.4 III.5 III.6 III.7 III.8 d d d d D d d D d d d d d d d d d d d d d d d d D d D d d d d d Haplotype Series of alleles along a short section of the same strand of DNA DNA Strand: Haplotype: ATCTGCCTCGCCATAAAGTCATTCGCTCAT ATCTGCCTCGCCATAAAGTCATTCGCTGAT ATCAGCCTCGCCATAAAGTCATTCGCTCAT ATCAGCCTCGCCATAAAGTCATTCGCTGAT position 4: position 28: T A C G allele allele allele allele TC TG AC AG Linkage Equilibrium & Disequilibrium If I know the first allele in a haplotype, can I predict the second allele? Yes Linkage Disequilibrium No Linkage Equilibrium Linkage Equilibrium & Disequilibrium In other words: Equilibrium: Frequency of a haplotype is due to chance. Disequilibrium: Frequency of a haplotype differs from chance frequency. Haplotype (Graduate) Chance: If the frequency of allele T is .2 and the frequency of allele C is .4, then the frequency of haplotype TC is .2*.4 = .08. Nonchance: If the frequency of allele T is .2 and the frequency of allele C is .4, then the frequency of haplotype TC is significantly different from .08. Haplotype (Graduate) Position 28: C G T TC TG A AC AG Position 4: Equilibrium (Graduate) Position 28: C G T .08 .12 .2 A .32 .48 .8 Position 4: .4 .6 Disequilibrium (Graduate) Position 28: C G T .16 .04 .2 A .14 .56 .8 Position 4: .4 .6 Statistics for Equilibrium (Graduate) Position 28: C G T X11 X12 p1 A X21 X22 q1 Position 4: p2 q2 Statistics for Equilibrium (Graduate) d = X11X22 - X12X21= cov(L1, L2) D = X11- p1p2 = X22 - q1q2 If D > 0, D = D/Dmax where Dmax = min(p1q2, p2q1) If D > 0, D = D/Dmax where Dmax = min(p1p2, q1q2) R2 = d2 / (p1p2q1q2) D and R2 are the most often used stats. Formation of Disequilibrium (Graduate) 1. Mutation occurs and creates a new spelling variation (polymorphism). 2. This creates linkage disequilibrium with those polymorphisms along the same DNA strand with the mutation. 3. Over generations, recombination will break up the disequilibrium with polymorphisms that are far away from the mutation. 4. Polymorphisms close to the original mutation, however, will remain in disequilibrium for a longer time. 5. Hence, polymorphisms close to the mutation will be in disequilibrium longer than polymorphisms farther away from the mutation. Disequilibrium: 1. Is the norm rather than the exception for short sections of DNA (100,000 nucleotides). 2. Generates “haplotype blocks” (see next slide). 3. Haplotype Mapping Project (HapMap): provide a map of the haplotype blocks for the human genome. 4. Allows genome-wide association studies. Haplotype Blocks: Section of DNA (vertical bar = polymorphism): Block 1 Block 2 Block 7 • Haplotype Block: Series of adjacent alleles in strong disequilibrium. • Logic: Instead of genotyping all 37 polymorphisms, genotype one in each block. • If there is a “hit,” then go back and genotype the other polymorphisms in that block. Haplotype block structure of the cytochrome P450 CYP2C gene cluster on chromosome 10. From Walton et al. (2005), Nature Genetics 37, 915-0916. Association Design • Begins with KNOWN polymorphism theoretically expected to be associated with the trait (e.g., DRD2 and schizophrenia). • Genotypes people on the gene and phenotypes them on the trait. •Tests whether the genotype is associated with the trait. •Two types: (1) Population-based (controls = general pop) (2) Family-based (controls = genetic relatives) Population-based Association Design Genotype: AA Aa Schiz: Phenotype: Not Schiz: Do c2 test for association. aa Genome-wide Association Study (GWAS) (1) Genotype one locus per haplotype block (2) Do an association test for every gene. (3) Number of genes that can be assayed changes from year to year. GWAS: Genome-wide Association Study 1. DNA arrays with 1,000s of SNPs scattered throughout the genome. (Current chips in 2009 has1,000, 000 different SNPs) 2. Select the SNPs so that they cover ALL the genome. (Some DNA chips concentrate on known protein coding regions rather than trying to cover all the genome) 3. Genotype patients and controls on all the SNPs. 4. Find the SNPs that differ. 5. Problem: number of statistical tests. Problems with GWAS (1) Expensive. (2) Large number of statistical tests. (3) Need very, very large samples (10,000 or more. Results from GWAS (1) Good success in medicine. (2) Limited success for psychiatric disorders (3) Virtually no success for normal behavioral traits (personality, IQ) (4) Genetics of behavior is hyper-polygenic: many, many, many genes