* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download What is PCR? - Cobb Learning
DNA damage theory of aging wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
DNA vaccination wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
DNA profiling wikipedia , lookup
DNA polymerase wikipedia , lookup
Metagenomics wikipedia , lookup
X-inactivation wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Designer baby wikipedia , lookup
Transposable element wikipedia , lookup
Point mutation wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Human genome wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Molecular cloning wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Primary transcript wikipedia , lookup
Epigenomics wikipedia , lookup
DNA supercoil wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Genealogical DNA test wikipedia , lookup
History of genetic engineering wikipedia , lookup
Non-coding DNA wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genome editing wikipedia , lookup
Population genetics wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Genomic library wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Helitron (biology) wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Microsatellite wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genetic drift wikipedia , lookup
Microevolution wikipedia , lookup
SNP genotyping wikipedia , lookup
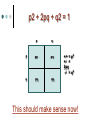
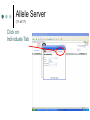
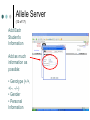
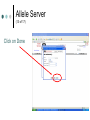
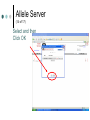
PV92 PCR Informatics Chromosome 16 Day #1: What is PCR? Day #2: Alu Insertion & PCR PCR Polymerase chain reaction (PCR) enables researchers to produce millions of copies of a specific DNA sequence in a relatively short period of time The specific sequence is called the “target sequence” PCR 1st described in 1985 by Kary Mullis He won the Nobel in 1993 for this Made it possible for researchers in a variety of biological fields to incorporate molecular biology (genetics) into their research. •Pathology •Botany •Zoology •Pharmacology What Is It & Why Did it Revolutionize Research PCR produces exponentially large amounts of DNA from trace amounts. Drop of blood Single hair follicle One cheek cell What’s Needed For PCR? Free nucleotides – building blocks DNA primers A strand of nucleic acid that serves as a starting point for DNA replication Complementary to target sequence Taq polymerase Comes from hot springs bacteria Can tolerate high heat of PCR • Discovery of this bacterium made PCR possible PCR Steps 3 Main Steps to PCR Denaturing at ~94 °C Annealing at ~54 °C • Binding of a primer to DNA strand. Extension at ~72 °C • The bases (complementary to the template) are coupled to the primer on the 3' side. Lots of Copies! PCR Animation http://www.sumanasinc.com/webcontent /animations/content/pcr.html Any Questions? PV92 PCR Informatics Chromosome 16 Day #1: What is PCR? Day #2: Alu Insertion & PCR What is PV92? PV92 – A human-specific Alu insertion on C16 Alu’s are a type of transposon Recall that transposons are pieces of DNA that can move around within the genome of a cell Barbara McClintock discovered transposons and won the Nobel in 1983. More About Alu Transposons Also known as ‘jumping genes’ Copies itself and inserts into new locations on the chromosome No evidence that ‘parent’ Alu segments are ever excised…what does this mean in terms of evolution & human population genetics? Alu is a SINE as well as a Retrotransposon Alu is ~300 bp in size Therefore known as a SINE (short interspersed repetitive element) Highly Conserved Alu endonuclease is called so because it was isolated from Anthrobacter luteus Retrotransposon because it uses reverse transcriptase to copy itself But Alu a Defective Transposon Can’t make it’s own reverse transcriptase (RT)…so…it hijacks another gene Hijacked gene is known as L1 (a LINE) and is basically a non-functional retrovirus that can make RT Say WHAT?????? It’s Like This… 1. Alu is transcribed into mRNA via RNA polymerase 2. mRNA hijacks L1 which converts Alu to ds DNA by way of RT 3. DNA copy of Alu is integrated into new chromosome site PV92 Alu Insertion A member of Alu repeat family Human-specific Alu insertion Found in a non-coding region of your DNA Not diagnostic for any disease or disorder (usually) 5’ Alu Amplified Region 3’ Introns, Exons and Alu Inserts Only ~5% of our genome consists of coding DNA (exons). • This is where our functional proteins come from. Out of the remaining 95%, ~40% is intron based, or non-coding. • We pass introns and exons on from one generation to the next. Introns, Exons and Alu Inserts Introns are full of SINES, Alu being one of them. • Origins of Alu unknown. We are targeting a specific locus on C16 that is known to carry the Alu sequence, or not. • Some of you will have the Alu insert and others will not. The Alu Insert is Dimorphic The PV92 Alu is dimorphic so there are two possible PCR products: 641 bp and 941 bp If you have the Alu insert on both of your homologous chromosomes = +,+ If you have Alu on one chromosome = +, If you don’t have the insert = -,- To Alu Or Not… No insertion: 641 bp Approximately 500,000 Alu copies per haploid genome, representing about 5% of the human genome. With Alu: 941 bp 300 bp Alu insert 641 bp Possible PCR Products + - +/- S1 S2 S3 S4 941 bp 641 bp We will use primers that are specific to the Alu region No Alu = 641 bp fragment Alu = 941 bp fragment In the Lab… 1. 2. 3. 4. 5. We will harvest some of your cells… Incubate them with Chelex resin (extract DNA)… Use PCR to amplify the Alu gene… Separate Alu fragments on 2% agarose gel… Use Chi-Square or Hardy Weinberg to calculate population frequency of (+,+), (+,-) and (-,-). Any Questions? Calculating Observed Genotypic Frequencies Hardy-Weinberg Equilibrium The HW equilibrium describes what happens to alleles in an ‘ideal’ population. 1. 2. 3. 4. 5. No selection (= rate of survival for all) No mutation No immigration/emigration Large population Random mating Hardy-Weinberg Equilibrium We know that when we cross two individuals who are heterzygous for Alu we will see: So there is a 25% chance of being ++, 25% being - - and 50% + -. + - + ++ +- - +- -- Hardy Weinberg ? Hardy-Weinberg Equation p2 + 2pq + q2 = 1 p = frequency of + alleles q = frequency of - alleles p q p pp pq q pq qq +/+ = p2 +/- = 2pq -/- = q2 Hardy-Weinberg Equation p2 + 2pq + q2 = 1 p = frequency of + allele q = frequency of - allele p2 = frequency of ++ q2 = frequency of – 2pq = frequency of + Hardy-Weinberg Equilibrium Example +/+ Genotypic frequency = Number with genotype Population total (N) = 25 38 .66 = Genotype +/+ +/– -/- Total (N) # of People 25 5 8 38 0.66 0.13 0.21 1.00 Observed Frequency Calculating Allelic Frequencies p = Frequency of + alleles = Number of + alleles Total number alleles = 55 76 Number of + alleles 25 individuals with two + alleles = 50 + alleles 5 individuals with one + allele = 5 + alleles Total = 55 + alleles Total number of alleles 2N = 2(38) = 76 p = 0.72; therefore q = 0.28 since p + q = 1.00 = 0.72 p2 + 2pq + q2 = 1 p q p pp pq q pq qq +/+ = p2 +/- = 2pq -/- = q2 This should make sense now! (p2, 2pq, q2 values) p2 + 2pq + q2 = 1.00 (0.72)2 + 2(0.72)(0.28) + (0.28)2 = 1.00 0.52 + 0.40 p2 = 0.52 + 0.08 2pq = 0.40 = 1.00 q2 = 0.08 Chi-Square Test Ok…so… Genotype Genotype frequency x Population total (N) = Expected number +/+ (p2) 0.52 x 38 = 20 +/– (2pq) 0.40 x 38 = 15 –/– 0.08 x 38 = 3 (q2) 2 X =∑ Genotype +/+ +/– –/– (Observed – Expected) 2 Expected Observed Expected (O–E)2 E 25 20 1.25 5 15 6.67 8 3 8.33 X2 = 16.25 Allele Server (1 of 17) Cold Springs Harbor Laboratory DNA Learning Center Web site: http://www.dnalc.org/ Allele Server (2 of 17) Scroll through DNALC internet sites until BioServers Link appears Allele Server (3 of 17) Click on Bioservers Allele Server (4 of 17) Enter the Allele Server Allele Server (5 of 17) Click on Manage Groups Allele Server (6 of 17) Select Group Allele Server (7 of 17) Scroll Down to Select “Your Group” Allele Server (8 of 17) Fill Out Form Allele Server (9 of 17) Click on Edit Group Allele Server (10 of 17) Edit Your Group Information Allele Server (11 of 17) Click on Individuals Tab Allele Server (12 of 17) Add Each Student’s Information Add as much information as possible: • Genotype (+/+, +/–. –/–) • Gender • Personal Information Allele Server (13 of 17) Click on Done Allele Server (14 of 17) Select and then Click OK Allele Server (15 of 17) Analyze Data 2: Then Click Here 1: Click Here First Allele Server (16 of 17) Click on the Terse and Verbose Tabs to Review Data Results Any Questions?