* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Will Entrez Find Every Sequence Record?
Public health genomics wikipedia , lookup
Genetic engineering wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genome (book) wikipedia , lookup
Human genome wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Protein moonlighting wikipedia , lookup
Pathogenomics wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Gene therapy wikipedia , lookup
Genome evolution wikipedia , lookup
Gene expression programming wikipedia , lookup
Gene expression profiling wikipedia , lookup
Sequence alignment wikipedia , lookup
Gene desert wikipedia , lookup
Genome editing wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Point mutation wikipedia , lookup
Microevolution wikipedia , lookup
Metagenomics wikipedia , lookup
Gene nomenclature wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Designer baby wikipedia , lookup
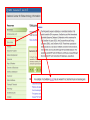
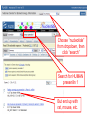
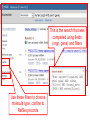
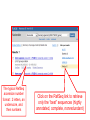
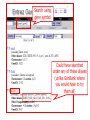
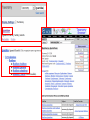
NCBI’s Bioinformatics Resources Michele R. Tennant, Ph.D., M.L.I.S. Health Science Center Libraries January 2016 Entrez Nucleotides Entrez Nucleotides (GenBank) • • • Database of nucleotide sequences (ATGC) Actually contains data from several databases - GenBank, EMBL, DDBJ, RefSeq Hard to search because many submitting scientists send in redundant information and poorly annotated information Nucleotide Data Domain • As of December 15, 2015 • Over 203,939,111,071 bases • Over 189,232,925 sequence records • Some complete genomes and chromosomes So Why So Hard to Search? • • • No controlled vocabulary - lose power of MeSH must OR synonyms. Often miss the records you want. Archival - quality of annotations depends on the submitter (especially features field); little to no quality control; spelling errors! Often miss the records you want. Redundant - lots of records for the same gene; partial records, etc. Often pull up records you don’t want. GenBank Sample Record • • Before searching, we will look at a GenBank sample record Note that the “Features” field provides useful biological information, and may be searched “Definition” field acts as record title – search [titl] Click any link in sample record to access definition of field and search tips Unique identifier; assigned by NCBI; required by journals/grants Link to PubMed citation/abstract The “Features” field provides the most biological information; search as [fkey] Numbers indicate location on the nucleotide sequence …3158 GenBank Identifiers • Accession Number - U49845 [accn] • Unique identifier; does not change • Letter prefix no longer has significance • Version - U49845.1 • If any change to sequence, version U49845.2 created • GenInfo Identifier (GI number) [uid] • Run parallel to accession.version system; change in sequence changes number Searching “Nucleotides” • Database is difficult to search: • Redundant records • Archival - poor or missing annotation • • Best searches are done using commands; need a class to learn all Practice search – search for sequences for human presenilin 1 • Is there anything odd about the some of the retrieved results? Choose “nucleotide” from dropdown, then click “search” Search for HUMAN presenilin 1 But end up with rat, mouse, etc. Searching “Nucleotides” • • We retrieved the non-human and PSEN2 (rather than PSEN1) records because the computer looked for the terms “human” and “presenilin 1” ANYWHERE in the record (click on details tab to see how the computer parsed your search) Use complex boolean searching to clean this up: term [field] AND term [field] Searching “Nucleotides” • • How to get rid of non-human sequences? • Search human [orgn] (this works for any taxon) How to get rid of non-presenilin 1 sequences? • Another trick – search PSEN1 [gene] • Note – you may miss relevant sequences, but should not pick up irrelevant sequences • The sequences that you miss are the ones that have not been annotated with the current official gene symbol in the “gene” field • DO NOT use this method if you need to find every sequence for a particular gene • Human [orgn] AND PSEN1 [gene] This is the search that was completed using fields (orgn, gene) and filters Use these filters to choose molecule type, confine to RefSeq records How Can I Find “Best” Sequences • • • • • • Non-redundant, curated subset of the sequence data domains Contains one record for each gene or splice variant from each organism represented Records can be thought of as “review articles” for sequences “Best” (usually longest) sequence used as seed Value-added annotations provided by experts Easy – a tab now exists to limit retrieval to just RefSeq The typical RefSeq accession number format: 2 letters, an underscore, and then numbers Click on the RefSeq link to retrieve only the “best” sequences (highly annotated, complete, nonredundant) Viewing Formats • • • The “Default” view is the standard GenBank record Researchers often use the “FASTA” format for analysis Change the record format at the “Display” pull-down menu Entrez Proteins Entrez Proteins • • Contains data from several databases: • SwissProt, PIR, PRF, PDB • Translations from annotated coding regions in GenBank and RefSeq Redundant archival data domain of publicly available protein sequences Searching Entrez Proteins • • Searched like Entrez Nucleotides “Filters” choices differ; includes molecular weight and sequence length filters Entrez Gene Entrez Gene • • • • Pulls together information (sequences, structures, literature, gene models, pathways, etc.) for genes Best place to start for “gene-centered” info One record per gene per organism Search by names, symbols, accessions, publications, GO terms, chromosome numbers, E.C. numbers, etc. Search using gene symbol Could have searched under any of these aliases (unlike GenBank where you would have to try them all) Official gene symbol as determined by the Human Genome Nomenclature Commission Summary of protein, function and diseasecausing mutations; from RefSeq record Links to PubMed records that provide evidence of function – any researcher can add these Gene Ontology terms form a controlled vocabulary with three components – biological process, molecular function, and cellular component Links to OMIM records of phenotype/ disease Links to protein interactions Links to homology maps Pathway info may be available from the Kyoto Encyclopedia of Genes and Genomes Sequence and domain links Links to GeneReviews – clinical resource Taxonomy Browser Search Taxonomy Browser • • How many genera from the family Iguanidae are represented by sequence data? How many nucleotide and protein sequences are available for the family? Entrez Searching Summary To Find Everything(?) Broaden Search • • • • • • OR together synonyms OR together related terms (gene name, gene symbol, protein name, alternate spellings, disorder) Don’t specify a field- search entire record Truncation - use * at end of word root Click “Similar articles” Try using Taxonomy Browser to pick up all taxa in a particular group Fewer/Best Records Narrow Search • • • Search particular fields: • PubMed - MeSH Browser, subheadings, major MeSH • Nucleotide - features, title, gene, properties, organism Use “Filters” Search only the RefSeq database Will Entrez Find Every Sequence Record? • No!!! • Entrez relies on annotation of records, so you are searching solely on “terminology” • Some records are not annotated, some records are poorly or incorrectly annotated • To find all useful sequences – need to search on sequence itself • BLAST Entrez “Similar Articles” • PubMed similar articles • Based on a “word weight” algorithm – MeSH, title, abstract words • In order by weight (highest weight first)