* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Slide 1
SNP genotyping wikipedia , lookup
Neocentromere wikipedia , lookup
Gene expression programming wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Pharmacogenomics wikipedia , lookup
X-inactivation wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Gene expression profiling wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Human genome wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Genomic imprinting wikipedia , lookup
Minimal genome wikipedia , lookup
Genetic drift wikipedia , lookup
History of genetic engineering wikipedia , lookup
Human genetic variation wikipedia , lookup
Heritability of IQ wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Medical genetics wikipedia , lookup
Genome evolution wikipedia , lookup
Behavioural genetics wikipedia , lookup
Pathogenomics wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Designer baby wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Population genetics wikipedia , lookup
Microevolution wikipedia , lookup
Genome (book) wikipedia , lookup
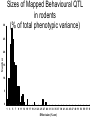
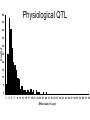
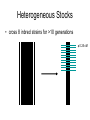
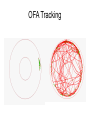
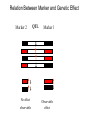
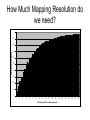
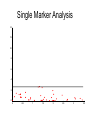
Finding the Molecular Basis of Quantitative Genetic Variation Richard Mott Wellcome Trust Centre for Human Genetics Oxford UK Genetic Traits • Quantitative (height, weight) • Dichotomous (affected/unaffected) • Factorial (blood group) • Mendelian - controlled by single gene (cystic fibrosis) • Complex – controlled by multiple genes*environment (diabetes, asthma) Molecular Basis of Quantitative Traits QTL: Quantitative Trait Locus chromosome genes Molecular Basis of Quantitative Traits QTL: Quantitative Trait Locus chromosome QTG: Quantitative Trait Gene Molecular Basis of Quantitative Traits QTL: Quantitative Trait Locus chromosome SNP: Single Nucleotide Polymorphism QTG: Quantitative Trait Gene QTN: Quantitative Trait Nucleotide Association Studies • Compare unrelated individuals from a population • Phenotypes: – Cases vs Controls – Quantitative measure • Genotypes: state of genome at multiple variable locations (Single Nucleotide Polymorphism = SNP) in each individual • Seek correlation between genotype and phenotype Problems with Association Studies • • • • • Population stratification Linkage Disequilibrium Allele Frequencies Multiple loci Small Effect Sizes • Very few Successes Population Stratification • If the sampling population comprises genetically distinct sub-populations with different disease prevalences • Then - • Any variant that distinguishes the subpopulations is likely to show disease association Admixture Mapping • Population is homogeneous but each individual’s genome is a mosaic of segments from different populations • May be used to map disease loci – multiple sclerosis susceptibility – Reich et al 2005, Nature Genetics Linkage Disequilibrium Mouse Effects of Linkage Disequilibrium • Correlation between nearby SNPs • SNPs near to QTN will show association – Risk of false positive interpretation – But need only genotype “tagging” SNPs – ~ 1 million tagging SNPs will be in LD with ~50% of common variants in the human genome The Common-Disease CommonVariant Hypothesis • Says – disease-predisposing variants will exist at relatively high frequency (i.e. >1%) in the population. – are ancient alleles occurring on specific haplotypes. – detectable in an case-control study using tagging SNPs. • Alternative hypothesis says – disease-predisposing alleles are sporadic new mutations, perhaps around the same genes, on different haplotypes. – families with history of the same disease owe their condition to different mutations events. – Theoretically detectable with family-based strategies which do not assume a common origin for the disease alleles, but are harder to detect with case-control studies (Pritchard, 2001). Power Depends on • Disease-predisposing allele’s – Effect Size (Odds Ratio) – Allele frequency • Sample Size: #cases, #controls • Number of tagging SNPs • To detect an allele with odds ratio of 1.25 and with allele frequency > 1%, at 5% Bonferroni genome-wide significance and 80% power, we require – ~ 6000 cases, 6000 controls – ~ 0.5 million tagging SNPs, one of which must be in perfect LD with the causative variant – [Hirschorn and Daly 2005] WTCCC Wellcome Trust Case-Control Consortium • 2000 cases from each of – – – – – – • • • • Type I Diabetes Type II Diabetes rheumatoid arthritis, susceptibility to TB bipolar depression …. and others … 3000 common controls 0.675 million SNPs ~10 billion genotypes Data expected mid 2006 Mouse Models Map in Human or Animal Models ? • Disease studied directly • Population and environment stratification • Very many SNPs (1,000,000?) required • Hard to detect trait loci – very large sample sizes required to detect loci of small effect (5,000-10,000) • Potentially very high mapping resolution – single gene • Very Expensive • Animal Model required • Population and environment controlled • Fewer SNPs required (~10010,000) • Easy to detect QTL with ~500 animals • Poorer mapping resolution – 1Mb (10 genes) • Relatively inexpensive QTL Mapping in Mice using Inbred Line Crosses • Genetically Homozygous – genome is fixed, breed true. • Standard Inbred Strains available • Haplotype diversity is controlled far more than in human association studies • QTL detection is very easy • QTL fine mapping is hard 30 Sizes of Mapped Behavioural QTL in rodents (% of total phenotypic variance) 25 Number 20 15 10 5 0 1 3 5 7 9 11 13 15 17 19 21 23 25 27 29 31 33 35 37 39 41 43 45 47 49 51 53 55 57 59 Effect size (% var) Physiological QTL 50 45 40 35 Number 30 25 20 15 10 5 0 1 3 5 7 9 11 13 15 17 19 21 23 25 27 29 31 33 35 37 39 41 43 45 47 49 51 53 55 57 59 Effect size (% var) Effect sizes of cloned genes 4 Number 3 2 1 0 1 3 5 7 9 11 13 15 17 19 21 23 25 27 29 31 33 35 37 39 41 43 45 47 49 51 53 55 57 59 Effect size (% var) QTL detection: F2 Intercross X A B QTL mapping: F2 Intercross X A X B F1 QTL mapping: F2 Intercross X A X B F1 F2 QTL mapping: F2 Intercross +1 -1 0 F1 QTL 0 0 F2 +2 -2 QTL mapping: F2 Intercross +1 -1 0 F1 0 0 F2 +2 -2 QTL mapping: F2 Intercross Genotype a skeleton of 20cM markers across genome 0 0 F2 +2 -2 QTL mapping: F2 Intercross AB AA AB BA AB BA AB BA AB BA BA BA BA BA BA AA BA BA BA AA 0 BB BB AB 0 AA F2 +2 -2 QTL mapping: F2 Intercross AB AA AB BA AB BA AB BA AB BA BA BA BA BA BA AA BA BA BA AA 0 BB BB AB 0 AA F2 +2 -2 Single Marker Association • Test of association between genotype and trait at each marker position. • ANOVA • F2 crosses are – good for detecting QTL – bad for fine-mapping – typical mapping resolution 1/3 chromosome – 20-30 cM Increasing mapping resolution • Increase number of recombinants: – more animals – more generations in cross Heterogeneous Stocks • cross 8 inbred strains for >10 generations Heterogeneous Stocks • cross 8 inbred strains for >10 generations Heterogeneous Stocks • cross 8 inbred strains for >10 generations 0.25 cM Mosaic Crosses founders G3 mixing GN inbreeding chopping up F2, diallele F20 HS, AI, outbreds RI (RIHS, CC) Analysis of mosaic crosses chromosome markers alleles 1 1 2 1 1 1 2 1 11 2 2 1 2 2 1 1 1 1 2 1 1 2 111 11 2 2 1 2 1 2 • Want to predict ancestral strain from genotype • We know the alleles in the founder strains • Single marker association lacks power, can’t distinguish all strains • Multipoint analysis – combine data from neighbouring markers Analysis of mosaic crosses chromosome markers alleles 1 1 2 1 1 1 2 1 11 2 2 1 2 2 1 1 1 1 2 1 1 2 111 11 2 2 1 2 1 2 •Hidden Markov model HAPPY •Hidden states = ancestral strains •Observed states = genotypes •Unknown phase of genotypes - analyse both chromosomes simultaneously •Output is probability that a locus is descended from a pair of strains •Mott et al 2000 PNAS Testing for a QTL • piL(s,t) = Prob( animal i is descended from strains s,t at locus L) • piL(s,t) calculated using – genotype data – founder strains’ alleles • Phenotype is modelled yi = Ss,t piL(s,t)T(s,t) + Covariatesi + ei • Test for no QTL at locus L – H0: T(s,t) are all same – ANOVA – partial F test Example: Open Field Avtivity • Mouse Model for Anxiety OFA Tracking multipoint singlepoint significance threshold Talbot et al 1999, Mott et al 2000 Relation Between Marker and Genetic Effect Marker 2 No effect observable QTL Marker 1 Observable effect How Much Mapping Resolution do we need? 1 0.9 0.8 Cumulative Probability 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 1 3 5 7 9 11 13 15 17 19 21 23 25 #Genes per Mb in mouse genome 27 29 31 33 35 37 39 Mapping Resolution in Mouse QTL experiments • F2 – ~25-50 Mb [250-300 genes] • HS – 1-5 Mb [10-50 genes] • Need More Resolution Other Outbred Populations • Commercially available outbreds may contain more historical recombination • Potentially finer mapping resolution • How to exploit it ? MF1 Outbred Mice MF1 Analysis of MF1 Single Marker Analysis 14 12 10 8 6 4 2 0 0 0.5 1 1.5 2 2.5 3 3.5 Unknown progenitors • Sometime in the 1970’s…. LACA x CF MF1 MF1 resemble HS • Sequencing revealed very few new variants in MF1 compared to HS strains • Variants present in HS strains also present in MF1 MF1 as a mosaic of inbred strains Mapping with 30 generation HS Mapping with MF1 mice Yalcin et al 2004 Nature Genetics Acknowledgements • • • • Jonathan Flint Binnaz Yalcin William Valdar Leah Solberg Further Reading • Mouse – Flint et al Nature Reviews Genetics 2005 • Human – Hirschhorn and Daly, Nature Reviews Genetics 2005 – Zondervan and Cardon, Nature Reviews Genetics 2004