* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Finland2012Lec1
Survey
Document related concepts
Transcript
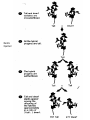
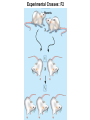
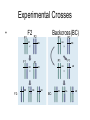
Bayesian Variable Selection in Semiparametric Regression Modeling with Applications to Genetic Mappping Fei Zou Department of Biostatistics University of North Carolina-Chapel Hill Email: [email protected] June 2012 Finland http://www.cs.unc.edu/Courses/comp590-090-f06/Slides/CSclass_Threadgill.ppt The Central Dogma of Molecular Biology tall short •Significant difference in genotype distributions? http://psb.stanford.edu/psb06/presentations/association_mapping.pdf • Copied (with modifications) from psb.stanford.edu/psb06/presentations/association_mapping.pdf Mendel’s Experiment Experimental Crosses: F2 P1 Parents P2 Experimental Crosses • P1 F2 AA F1 F2: P1 BB P1 F1 F1 AA AB AB P2 AA BB AB BB Backcross(BC) P2 AB BC: AB AB AA AB F2 Data Format 0: homozygous AA, 2: homozygous BB, 1: heterozygote AB. Data Structure • For each subject i (i=1,2,…,n) – Phenotype: yi – Genotypes: xij (coded as 0, 1, 2 for genotypes AA, AB and BB, respectively) at marker j (j=1,2,…,m) – Genetic map: locations of markers – Other non-genetic covariates, such as age, sex, environmental conditions Locations of markers Linkage Analysis • Quantitative trait loci (QTL): a particular region of the genome containing one or more genes that are associated with the trait being assayed or measured QTL Mapping of Experimental Crosses • Single QTL Mapping • Single marker analysis • Interval mapping: Lander & Botstein (1989, Genetics) • Multiple QTL mapping • Composite interval mapping • Multiple interval mapping • Bayesian analysis Single Marker Analysis Correlations of marker genotypes in experimental crosses Interval Mapping • Traditional QTL mapping method • Treat QTL position as unknown and use marker genotypes to infer conditional probabilities of QTL genotypes • Profile LOD scores calculated across whole genome – LOD score is a measure for strength of support for QTL – LOD = LRT/4.8 – In any region where the profile exceeds a (genome-wide) significance threshold, a QTL is declared at the position with the highest LOD score. Profile LOD 8 lod 6 4 2 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16171819 X Chromosome QTL • Old believe: one trait one gene – very unlikely • Most traits have a significant environmental exposure component • The vast majority of biological traits are caused by complex polygenic interactions – also context dependent Multiple QTL Mapping • Most complicated traits are caused by multiple (potentially interacting) genes, which also interact with environmental stimuli • Single QTL interval mapping – Ghost QTL – Low power if multiple QTLs affect the trait Two QTL Data Two QTL with opposite effects Two QTL with effects in same direction Multiple QTL Mapping • Available Methods – Composite interval mapping: searching for a putative QTL in a given region while simultaneously fitting partial regression coefficients for "background markers" to adjust the effects of other QTLs outside the region • which background markers to include; window size etc – Multiple interval mapping: fitting multiple QTLs simultaneously • Computationally very intensive; how many QTLs to fit? Multiple QTL Mapping Multiple QTL Mapping Multiple QTL Mapping Bayesian QTL Mapping • Reversible jump Markov chain Monte Carlo (MCMC) (Green 1995): treat the number of QTLs as a parameter – Change of dimensionality, the acceptance probability for such dimension change, which in practice, may not be handled correctly (Ven 2004) • Bayesian variable selection procedures – composite model space (Yi 2004) – stochastic search variable selection (SSVS) (George and McCulloch 1993)