* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Molecular Biology of the Peribacteroid Membrane
Epigenetics of neurodegenerative diseases wikipedia , lookup
Primary transcript wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Heritability of IQ wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Public health genomics wikipedia , lookup
Genetically modified crops wikipedia , lookup
Pathogenomics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Transcription factor wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
Microevolution wikipedia , lookup
Essential gene wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Gene expression programming wikipedia , lookup
Genome evolution wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genome (book) wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Genomic imprinting wikipedia , lookup
Minimal genome wikipedia , lookup
Ridge (biology) wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
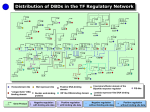
A reverse-genetic screen for N-regulators • UNDERLYING ASSUMPTIONS – Transcription factors (TFs) are involved in N-regulation – Some of these TFs are regulated at the transcriptional level by N availability • SCREENING STRATEGY – Identify N-regulated TF genes by real-time RT-PCR – Over-express suspect TF genes in Arabidopsis (using constitutive and inducible promoters) – Obtain TF knockout mutants for genes of interest – Determine the phenotype of mutant and transgenic lines – Test expression of N-assimilation genes in mutants and transgenic lines – Confirm physical interaction between TF and promoters (e.g. ChIP-PCR) Real-time RT-PCR profiling of >1400 Arabidopsis transcription factor genes 90% coverage with high specificity, sensitivity, dynamic range, and robustness 45 A CT Value 40 35 30 25 2 R =0.999 20 2 R =0.997 15 100 101 102 103 104 105 106 Double-stranded Template Copy Number Fraction of Shoot cDNA 1.0 Expression Level (2 (40-CT) ) 125000 0.8 0.6 0.4 0.2 0.0 B 2 100000 R =0.996 75000 2 R =0.998 50000 2 R =0.993 25000 2 R =0.994 0 0.0 0.2 0.4 0.6 Fraction of Root cDNA Czechowski et al. (2004) Plant J. 38, 366-379. 0.8 1.0 Real-time RT-PCR is more precise than Affy chips Real-time RT-PCR Intra-assay (A) and Interassay (B) variation Affy Chips Interassay variation N-regulated TF genes: Suspects for global control of N acquisition Axenic culture: +N to –N to NO3- Log2 TF ratio X/+N 10 6 2 0 -2 -6 -10 Inducible over-expression to identify target genes/phenotypes +N -N2d NO3-30min 40 N-regulated TFs, including regulators of anthacyanin biosynthesis and flowering (FD). Scheible et al. (2004) Plant Physiol. (in press). The role of transcription factors in abiotic stress tolerance and plant development Overview of TF projects -N NO3- -P Redox Real-time RT-PCR/ Reverse genetics -S Heterosis Salt+H2O stress Seed Develop. Seed/silique-specific TFs: potential tools for seed biotechnology Shoots (log2) TF gene expression level relative to ubi10 50-fold induced: candidates Siliques (log2)