* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Chapter 14 – RNA molecules and RNA processing
X-inactivation wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Non-coding DNA wikipedia , lookup
Microevolution wikipedia , lookup
Genome evolution wikipedia , lookup
Hammerhead ribozyme wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Frameshift mutation wikipedia , lookup
Human genome wikipedia , lookup
Gene expression profiling wikipedia , lookup
Point mutation wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Alternative splicing wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Expanded genetic code wikipedia , lookup
RNA interference wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Transfer RNA wikipedia , lookup
Genetic code wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
RNA silencing wikipedia , lookup
Polyadenylation wikipedia , lookup
History of RNA biology wikipedia , lookup
Messenger RNA wikipedia , lookup
Non-coding RNA wikipedia , lookup
RNA-binding protein wikipedia , lookup
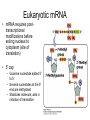
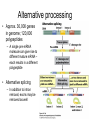
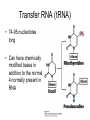
Chapter 14 – RNA molecules and RNA processing Gene organization • Francis Crick – 1958 – Nucleotide sequence of a gene directly codes for amino acid sequence of polypeptide • Gene contains interruptions of nucleotides that do not code for amino acids – Eukaryotic genes Gene organization • DNA and RNA transcripts within the nucleus are larger than transcripts found in the cytoplasm – Exons are coding regions – Introns are intervening sequences in eukaryotic genes • Occasionally seen in prokaryotes as well • Are spliced out of premRNA/primary transcript before leaving the nucleus Messenger RNA (mRNA) • Structure – 5′ untranslated region • Bacteria contain ShineDalgarno sequence – Serves as ribosome attachment site – Protein coding region • Codes for amino acids • Codon – 3 nucleotide sequence that codes for one amino acid – 3′ untranslated region • Aids in stability of molecule Prokaryotic mRNA • Since introns are rare, mRNA can begin to be translated before transcription is complete – Ribosome associates with Shine-Dalgarno sequence, and moves down mRNA molecule in 5′→3′ direction Eukaryotic mRNA • mRNA requires posttranscriptional modifications before exiting nucleus to cytoplasm (site of translation) • 5′ cap – Guanine nucleotide added 5′ to 5′ – Several nucleotides at the 5′ end are methylated – Stabilizes molecule; aids in initiation of translation Post-transcriptional modifications cont • polyA tail – At 3′ end there is at least one possible cleavage site where nucleotides are removed • After removal, 50-250 adenine nucleotides are added – Polyadenylation – With associated proteins, stabilizes molecule Post-transcriptional modifications cont • RNA splicing – Splicing out of introns requires 5′ splice site, 3′ splice site, and branch point – Spliceosome • 5 snRNPs and several other proteins – Process of splicing • 5′ end of intron is cut and folds back on itself to attach to branch point sequence – Forms a lariat • 3′ end of intron is cut and intron is released • 3′ of exon #1 is ligated to 5′ end of exon #2 • Intron reverts to linear form and is degraded Alternative processing • Approx. 30,000 genes in genome; 120,000 polypeptides – A single pre-mRNA molecule can give rise to different mature mRNA – each results in a different polypeptide • Alternative splicing – In addition to intron removal, exons may be removed as well Alternative processing cont • Multiple 3′ cleavage sites – Cleavage may occur at different sites before polyA tail is added – Any exons not included will yield a different polypeptide Transfer RNA (tRNA) • 74-95 nucleotides long • Can have chemically modified bases in addition to the normal 4 normally present in RNA tRNA cont • Complementary base pairs form a cloverleaf shape (folds into an “L” 3D) • 3′ end is the acceptor arm – where a specific amino acid attaches • Anticodon arm contains 3 nucleotides (anticodon) that recognize codon of mRNA • Initial transcript contains introns that are removed Ribosomal RNA (rRNA) • Original transcript for ribosomal RNA is cleaved by snoRNA – snoRNA + proteins form snoRNPs • Sizes of rRNA measured in Svedberg untis (S) – how fast substances sediment out in a centrifugal field) – Based on molecular weight and structure – not cumulative • Ribosomes – 1 or more molecules of rRNA and approx 50 proteins – Complete ribosome consists of 2 subunits – large and small RNA interference (RNAi) • miRNA (microRNA) and siRNA (small interfering RNA) • Both arise from doublestranded RNA, which is cute by enzyme Dicer – fragments are miRNA and siRNA • Small fragments bind to mRNA – miRNA inhibits translation – siRNA – degrades mRNA