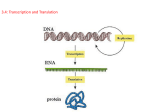
* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download video slide - Independent School District 196
Histone acetylation and deacetylation wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Gene regulatory network wikipedia , lookup
Molecular evolution wikipedia , lookup
RNA interference wikipedia , lookup
List of types of proteins wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Biochemistry wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Bottromycin wikipedia , lookup
Transcription factor wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Non-coding DNA wikipedia , lookup
RNA silencing wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Point mutation wikipedia , lookup
Polyadenylation wikipedia , lookup
Expanded genetic code wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Genetic code wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Non-coding RNA wikipedia , lookup
Gene expression wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Chapter 17 From Gene to Protein PowerPoint TextEdit Art Slides for Biology, Seventh Edition Neil Campbell and Jane Reece Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.1 A ribosome, part of the protein synthesis machinery Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.3 Overview: the roles of transcription and translation in the flow of genetic information (layer 1) TRANSCRIPTION DNA (a) Prokaryotic cell. In a cell lacking a nucleus, mRNA produced by transcription is immediately translated without additional processing. (b) Eukaryotic cell. The nucleus provides a separate compartment for transcription. The original RNA transcript, called pre-mRNA, is processed in various ways before leaving the nucleus as mRNA. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.3 Overview: the roles of transcription and translation in the flow of genetic information (layer 2) TRANSCRIPTION DNA mRNA TRANSLATION Ribosome Polypeptide (a) Prokaryotic cell. In a cell lacking a nucleus, mRNA produced by transcription is immediately translated without additional processing. (b) Eukaryotic cell. The nucleus provides a separate compartment for transcription. The original RNA transcript, called pre-mRNA, is processed in various ways before leaving the nucleus as mRNA. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.3 Overview: the roles of transcription and translation in the flow of genetic information (layer 3) TRANSCRIPTION DNA mRNA Ribosome TRANSLATION Polypeptide (a) Prokaryotic cell. In a cell lacking a nucleus, mRNA produced by transcription is immediately translated without additional processing. Nuclear envelope TRANSCRIPTION DNA (b) Eukaryotic cell. The nucleus provides a separate compartment for transcription. The original RNA transcript, called pre-mRNA, is processed in various ways before leaving the nucleus as mRNA. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.3 Overview: the roles of transcription and translation in the flow of genetic information (layer 4) TRANSCRIPTION DNA mRNA Ribosome TRANSLATION Polypeptide (a) Prokaryotic cell. In a cell lacking a nucleus, mRNA produced by transcription is immediately translated without additional processing. Nuclear envelope TRANSCRIPTION DNA RNA PROCESSING Pre-mRNA mRNA (b) Eukaryotic cell. The nucleus provides a separate compartment for transcription. The original RNA transcript, called pre-mRNA, is processed in various ways before leaving the nucleus as mRNA. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.3 Overview: the roles of transcription and translation in the flow of genetic information (layer 5) TRANSCRIPTION DNA mRNA Ribosome TRANSLATION Polypeptide (a) Prokaryotic cell. In a cell lacking a nucleus, mRNA produced by transcription is immediately translated without additional processing. Nuclear envelope TRANSCRIPTION DNA RNA PROCESSING Pre-mRNA mRNA Ribosome TRANSLATION Polypeptide Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings (b) Eukaryotic cell. The nucleus provides a separate compartment for transcription. The original RNA transcript, called pre-mRNA, is processed in various ways before leaving the nucleus as mRNA. Figure 17.4 The triplet code DNA molecule Gene 2 Gene 1 Gene 3 DNA strand (template) 5 3 A C C A A A C C G A G T U G G U U U G G C U C A TRANSCRIPTION mRNA 5 Codon TRANSLATION Protein Trp Amino acid Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Phe Gly Ser 3 Figure 17.5 The dictionary of the genetic code Second mRNA base C UUU UUC U UUA First mRNA base (5 end) A UAU UCU Phe UCC UCA UAC Ser U UGU Tyr UGC Cys C UCG CUU CCU CAU CUC CCC CAC CUA Leu Leu CCA Pro CAA CUG CCG CAG AUU ACU AAU ACC AAC AUC lle AUA ACA Thr AAG GUU GCU GAU GUC GCC GAC GUA GUG Val GCA Ala GCG Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings His Gln Asn AAA AUGMet or ACG start G G UAA Stop UGA Stop A UAG Stop UGG Trp G UUG C A Lys CGC CGA Arg Asp CGG G AGU U AGC Ser C AGA A AGG Arg G GGC GGA Glu C A U GGU GAA GAG U CGU GGG Gly C A G Third mRNA base (3 end) U Figure 17.6 A tobacco plant expressing a firefly gene Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.7 The stages of transcription: initiation, elongation, and termination (layer 1) Promoter Transcription unit 5 3 3 5 Start point RNA polymerase Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings DNA Figure 17.7 The stages of transcription: initiation, elongation, and termination (layer 2) Promoter Transcription unit 5 3 3 5 Start point DNA 1 Initiation. After RNA polymerase binds to RNA polymerase the promoter, the DNA strands unwind, and the polymerase initiates RNA synthesis at the start point on the template strand. 5 3 3 5 Template strand of DNA Unwound RNA DNA transcript Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.7 The stages of transcription: initiation, elongation, and termination (layer 3) Promoter Transcription unit 5 3 3 5 Start point DNA 1 Initiation. After RNA polymerase binds to RNA polymerase the promoter, the DNA strands unwind, and the polymerase initiates RNA synthesis at the start point on the template strand. 5 3 3 5 Template strand of DNA Unwound RNA DNA transcript 2 Elongation. The polymerase moves downstream, unwinding the Rewound DNA and elongating the RNA transcript 5 3 . In the wake of transcription, the DNA strands re-form a double helix. RNA 5 3 3 5 RNA transcript Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 3 5 Figure 17.7 The stages of transcription: initiation, elongation, and termination (layer 4) Promoter Transcription unit 5 3 3 5 Start point DNA 1 Initiation. After RNA polymerase binds to RNA polymerase the promoter, the DNA strands unwind, and the polymerase initiates RNA synthesis at the start point on the template strand. 5 3 3 5 Template strand of DNA Unwound RNA DNA transcript 2 Elongation. The polymerase moves downstream, unwinding the Rewound DNA and elongating the RNA transcript 5 3 . In the wake of transcription, the DNA strands re-form a double helix. RNA 5 3 3 5 3 5 RNA transcript 3 Termination. Eventually, the RNA transcript is released, and the polymerase detaches from the DNA. 5 3 3 5 3 5 Completed RNA transcript Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Non-template strand of DNA Elongation RNA nucleotides RNA polymerase A T C C A A 3 3 end U 5 A E G C A T A G G T T Direction of transcription (“downstream) 5 Newly made RNA Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Template strand of DNA Figure 17.8 The initiation of transcription at a eukaryotic promoter TRANSCRIPTION 1 Eukaryotic promoters DNA RNA PROCESSING Pre-mRNA mRNA TRANSLATION Ribosome Polypeptide Promoter 5 3 3 5 T A T A A AA ATAT T T T TATA box Start point Template DNA strand 2 Several transcription factors Transcription factors 5 3 3 5 3 Additional transcription factors RNA polymerase II Transcription factors 5 3 3 5 5 RNA transcript Transcription initiation complex Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.9 RNA processing: addition of the 5 cap and poly-A tail A modified guanine nucleotide added to the 5 end TRANSCRIPTION RNA PROCESSING 50 to 250 adenine nucleotides added to the 3 end DNA Pre-mRNA 5 mRNA Protein-coding segment Polyadenylation signal 3 G P P P AAUAAA AAA…AAA Ribosome TRANSLATION 5 Cap 5 UTR Start codon Stop codon Polypeptide Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 3 UTR Poly-A tail Figure 17.10 RNA processing: RNA splicing TRANSCRIPTION RNA PROCESSING DNA Pre-mRNA 5 Exon Intron Pre-mRNA 5 Cap 30 31 1 Coding segment mRNA Ribosome Intron Exon Exon 3 Poly-A tail 104 105 146 Introns cut out and exons spliced together TRANSLATION Polypeptide mRNA 5 Cap 1 3 UTR Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Poly-A tail 146 3 UTR Figure 17.11 The roles of snRNPs and spliceosomes in pre-mRNA splicing RNA transcript (pre-mRNA) 5 Intron Exon 1 Exon 2 Protein 1 Other proteins snRNA snRNPs Spliceosome 2 5 Spliceosome components 3 mRNA 5 Exon 1 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Exon 2 Cut-out intron Figure 17.12 Correspondence between exons and protein domains Gene DNA Exon 1 Intron Exon 2 Intron Exon 3 Transcription RNA processing Translation Domain 3 Domain 2 Domain 1 Polypeptide Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.13 Translation: the basic concept TRANSCRIPTION DNA mRNA Ribosome TRANSLATION Polypeptide Amino acids Polypeptide Ribosome tRNA with amino acid attached Gly tRNA Anticodon A A A U G G U U U G G C Codons 5 mRNA Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 3 Figure 17.14 The structure of transfer RNA (tRNA) 3 A Amino acid C attachment site C A 5 C G G C C G U G U A A U A U U C * G U AG * CACA A CUC * G * U G U G G * C CGAG C * * AGG U * * GA G C Hydrogen G C bonds U A * G * A A C * U A A G Anticodon (a) Two-dimensional structure. The four base-paired regions and three loops are characteristic of all tRNAs, as is the base sequence of the amino acid attachment site at the 3 end. The anticodon triplet is unique to each tRNA type. (The asterisks mark bases that have been chemically modified, a characteristic of tRNA.) Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 5 3 Amino acid attachment site Hydrogen bonds A 3 Anticodon (b) Three-dimensional structure Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings A G Anticodon 5 (c) Symbol used in this book Figure 17.15 An aminoacyl-tRNA synthetase joins a specific amino acid to a tRNA Amino acid Aminoacyl-tRNA synthetase (enzyme) 1 Active site binds the amino acid and ATP. P P P Adenosine ATP 2 ATP loses two P groups and joins amino acid as AMP. P Adenosine Pyrophosphate Pi Phosphates P Pi Pi tRNA 3 Appropriate tRNA covalently Bonds to amino Acid, displacing AMP. P Adenosine AMP 4 Activated amino acid is released by the enzyme. Aminoacyl tRNA (an “activated amino acid”) Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.16 The anatomy of a functioning ribosome TRANSCRIPTION DNA mRNA Ribosome TRANSLATION Polypeptide Exit tunnel Growing polypeptide tRNA molecules Large subunit E P A Small subunit 5 mRNA 3 (a) Computer model of functioning ribosome. This is a model of a bacterial ribosome, showing its overall shape. The eukaryotic ribosome is roughly similar. A ribosomal subunit is an aggregate of ribosomal RNA molecules and proteins. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings P site (Peptidyl-tRNA binding site) A site (AminoacyltRNA binding site) E site (Exit site) Large subunit E mRNA binding site P A Small subunit (b) Schematic model showing binding sites. A ribosome has an mRNA binding site and three tRNA binding sites, known as the A, P, and E sites. This schematic ribosome will appear in later diagrams. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Amino end Growing polypeptide Next amino acid to be added to polypeptide chain tRNA 3 mRNA 5 Codons (c) Schematic model with mRNA and tRNA. A tRNA fits into a binding site when its anticodon base-pairs with an mRNA codon. The P site holds the tRNA attached to the growing polypeptide. The A site holds the tRNA carrying the next amino acid to be added to the polypeptide chain. Discharged tRNA leaves via the E site. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.17 The initiation of translation Large ribosomal subunit P site 3 U A C 5 5 A U G 3 Initiator tRNA GTP GDP E A mRNA 5 Start codon 3 mRNA binding site Small ribosomal subunit 1 A small ribosomal subunit binds to a molecule of mRNA. In a prokaryotic cell, the mRNA binding site on this subunit recognizes a specific nucleotide sequence on the mRNA just upstream of the start codon. An initiator tRNA, with the anticodon UAC, base-pairs with the start codon, AUG. This tRNA carries the amino acid methionine (Met). Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 5 3 Translation initiation complex 2 The arrival of a large ribosomal subunit completes the initiation complex. Proteins called initiation factors (not shown) are required to bring all the translation components together. GTP provides the energy for the assembly. The initiator tRNA is in the P site; the A site is available to the tRNA bearing the next amino acid. Figure 17.18 The elongation cycle of translation 1 Codon recognition. The anticodon TRANSCRIPTION Amino end of polypeptide DNA mRNA Ribosome of an incoming aminoacyl tRNA base-pairs with the complementary mRNA codon in the A site. Hydrolysis of GTP increases the accuracy and efficiency of this step. TRANSLATION Polypeptide E mRNA Ribosome ready for next aminoacyl tRNA 5 3 P A site site 2 GTP 2 GDP E E P P A 2 Peptide bond formation. An GDP 3 Translocation. The ribosome translocates the tRNA in the A site to the P site. The empty tRNA in the P site is moved to the E site, where it is released. The mRNA moves along with its bound tRNAs, bringing the next codon to be translated into the A site. GTP Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings A E P A rRNA molecule of the large Subunit catalyzes the formation of a peptide bond between the new amino acid in the A site and the carboxyl end of the growing polypeptide in the P site. This step attaches the polypeptide to the tRNA in the A site. Figure 17.19 The termination of translation Release factor Free polypeptide 5 3 3 3 5 5 Stop codon (UAG, UAA, or UGA) 1 When a ribosome reaches a stop 2 The release factor hydrolyzes codon on mRNA, the A site of the ribosome accepts a protein called a release factor instead of tRNA. the bond between the tRNA in the P site and the last amino acid of the polypeptide chain. The polypeptide is thus freed from the ribosome. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 3 The two ribosomal subunits and the other components of the assembly dissociate. Figure 17.20 Polyribosomes Completed polypeptide Growing polypeptides Incoming ribosomal subunits Start of mRNA (5 end) End of mRNA (3 end) (a) An mRNA molecule is generally translated simultaneously by several ribosomes in clusters called polyribosomes. Ribosomes mRNA 0.1 µm (b) This micrograph shows a large polyribosome in a prokaryotic cell (TEM). Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.21 The signal mechanism for targeting proteins to the ER 1 Polypeptide synthesis begins on a free ribosome in the cytosol. 2 An SRP binds to the signal peptide, halting synthesis momentarily. 3 The SRP binds to a receptor protein in the ER membrane. This receptor is part of a protein complex (a translocation complex) that has a membrane pore and a signal-cleaving enzyme. 4 The SRP leaves, and the polypeptide resumes growing, meanwhile translocating across the membrane. (The signal peptide stays attached to the membrane.) 5 The signalcleaving enzyme cuts off the signal peptide. 6 The rest of the completed polypeptide leaves the ribosome and folds into its final conformation. Ribosome mRNA Signal peptide Signalrecognition particle (SRP) SRP receptor CYTOSOL protein Translocation complex Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Signal peptide removed ER membrane Protein Figure 17.22 Coupled transcription and translation in bacteria RNA polymerase DNA mRNA Polyribosome RNA polymerase Direction of transcription 0.25 m DNA Polyribosome Polypeptide (amino end) Ribosome mRNA (5 end) Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.23 The molecular basis of sickle-cell disease: a point mutation Wild-type hemoglobin DNA 3 Mutant hemoglobin DNA 5 C T T In the DNA, the mutant template strand has an A where the wild-type template has a T. A The mutant mRNA has a U instead of an A in one codon. 3 5 T C mRNA A mRNA G A A 5 G 3 U 5 3 Normal hemoglobin Sickle-cell hemoglobin Glu Val Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings The mutant (sickle-cell) hemoglobin has a valine (Val) instead of a glutamic acid (Glu). Figure 17.24 Base-pair substitution Wild type mRNA A U G A A G U U U G G C U A A 5 Protein Met Lys 3 Phe Amino end Gly Stop Carboxyl end Base-pair substitution No effect on amino acid sequence U instead of C A U G A A G U U U G G U U A A Met Lys Missense Phe Gly Stop A instead of G A U G A A G U U U A G U U A A Met Lys Phe Ser Stop Nonsense U instead of A A U G U A G U U U G G C U A A Met Stop Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Figure 17.25 Base-pair insertion or deletion Wild type mRNA 5 Protein A UG A A GU U U GG C U A A Met Lys Gly Phe 3 Stop Amino end Carboxyl end Base-pair insertion or deletion Frameshift causing immediate nonsense Extra U A U GU A A G U U U GG C U A Met Stop Frameshift causing extensive missense U Missing A U G A A G U U G G C U A A Met Lys Leu Ala Insertion or deletion of 3 nucleotides: no frameshift but extra or missing amino acid A A G Missing A U G U U UG G C U A A Met Phe Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Gly Stop Figure 17.26 A summary of transcription and translation in a eukaryotic cell DNA TRANSCRIPTION 1 RNA is transcribed from a DNA template. 3 5 RNA transcript RNA polymerase RNA PROCESSING Exon 2 In eukaryotes, the RNA transcript (premRNA) is spliced and modified to produce mRNA, which moves from the nucleus to the cytoplasm. RNA transcript (pre-mRNA) Intron Aminoacyl-tRNA synthetase NUCLEUS Amino acid tRNA FORMATION OF INITIATION COMPLEX CYTOPLASM 3 After leaving the nucleus, mRNA attaches to the ribosome. mRNA AMINO ACID ACTIVATION 4 Each amino acid attaches to its proper tRNA with the help of a specific enzyme and ATP. Growing polypeptide Activated amino acid Ribosomal subunits 5 TRANSLATION A succession of tRNAs add their amino acids to the polypeptide chain Anticodon as the mRNA is moved through the ribosome one codon at a time. (When completed, the polypeptide is released from the ribosome.) 5 E A AAA UG GU U U A U G Codon Ribosome Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings