* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Deamination of Cytosine and 5
Gene expression profiling wikipedia , lookup
Minimal genome wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Epitranscriptome wikipedia , lookup
Expanded genetic code wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Gene expression programming wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Koinophilia wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Genome (book) wikipedia , lookup
Non-coding DNA wikipedia , lookup
DNA polymerase wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Population genetics wikipedia , lookup
Designer baby wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Genetic code wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
History of genetic engineering wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Microsatellite wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Genome evolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genome editing wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Helitron (biology) wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Oncogenomics wikipedia , lookup
Transfer RNA wikipedia , lookup
Microevolution wikipedia , lookup
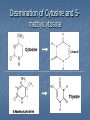
Deamination of Cytosine and 5methylcytosine ------------------------------------------------------------------------------- Chemical Mutagens Intercalating Agents EX, Ethidium bromide, acridine orange Can induce frameshift mutations Uv Induced Dimers Thymine dimers and T-C dimers Replication problems Interferes with the ability of the T’s to base pair to the opposite strand, and blocks DNA replication Other Mutagens Transposable elements—”jumping genes”. Major frameshift mutations Factors in evolution Mutator genes—mutations increase mutation rate. Four potent mutator genes Mutant Mutant Mutant Mutant DNA pol III 3’5’ exonuclease activity methylation enzymes(ex dam) enzymes in excision repair system enzymes in SOS system Reversions Mutations in an mutant can restore the wild type function (reversion, back mutation, or reverse mutation) Spontaneous or induced If mutation occurs at the site of the original=True reversion Wild type restored by mutation at another site= second site mutation Second site in same gene= intragenic suppression Second site in another gene=intergenic suppression Intragenic Revertant Types Same site reversion Second site revertant NOTE: shape of R-groups can also be a factor. EX Reversion of Frameshift Mutations For reversion to be successful Reversion must be near original site to reduce # of aa altered Section of polypeptide must be able to withstand alteration without eliminating function Intergenic Suppression Refers to a chnge in another gene which suppresses or eliminates the mutant phenotype. EX Multisubunit proteins—Mutation in one subunit may be masked by mutation in another subunit (ex. restoring hydrophobic patches) Suppression via suppressor tRNAs Suppressor tRNA Nonsense mutation- aa codon “Stop” Some bacteria can “read through” these mutations (though protein function may be altered). HOW? Mutant tRNA that has an anticodon that recognizes “Stop” as a reading codon. Ex. AAGUAG aa encoded depends on which tRNA is mutated Not every suppressor restores normal function Suppressor Mutants (cont’d) EX. UUG (Leu)UAG (Stop) (AUC anticodon) A mutation in a tRNA resulting in “AUC” allows that tRNA to recognize “Stop”. Can get suppression or partial suppression NOTE: must be 2 copies of tRNA mutated. Why? In any cell containing mutator, must also be a wild type Suppressors allow survival, even if suboptimal Termination of Translation in Suppressor Strains Problem: Must be a means of terminating translation. HOW? Release factors still present, will compete for the “Stop “ site Many genes are double-terminated EX. UAG-UAA Repair by Direct Reversal Photolyase O6-methylguanine methyl transferase Fig. 20.39 Fig. 20.40 Excision Repair Most common repair mechanism EX. Uvr system NOTE: preferentially repairs dimers in essential regions of genome UvRA recognizes damage and binds w/UvrB UvRA released, UvrC binds UvrC nicks on both sides of damageUvrD unwinds region Damaged strand released DAN pol I Ligase Recombinational Repair Sister Strand Exchange Recombinational Repair, aka, Sister Strand Exchange SOS Response Is an inducible system of last resort Also called error prone replication because it inactivates the proofreading function of DNA pol III. Turned on only when DNA damage is extreme Main players: recA and lexA and a battery of inducible enzymes SOS Response