* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Chapter 10 Intracellular Compartments and Transport
Survey
Document related concepts
Model lipid bilayer wikipedia , lookup
Cytokinesis wikipedia , lookup
Protein phosphorylation wikipedia , lookup
Magnesium transporter wikipedia , lookup
Protein moonlighting wikipedia , lookup
Cell nucleus wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
SNARE (protein) wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Cell membrane wikipedia , lookup
Signal transduction wikipedia , lookup
Proteolysis wikipedia , lookup
Western blot wikipedia , lookup
Transcript
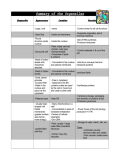
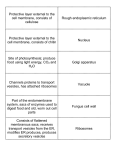
Chapter 10 Intracellular Compartments and Transport In eucaryotic cells, internal membrane create enclosed compartments and organelles in which different metabolic processes are segregated Membrane-enclosed organelles Aggregates of glycogen in liver cell A cell from the lining of the intestine contains the basic set of organelles found in most animal cells (nuclear envelope) glycosylation steroid smooth ER rough ER ; Ca2+ store Nuclear membranes and the ER may have evolved through investigation of the plasma membrane Endomembrane system Nuclear membrane ER Golgi apparatus Endosomes Lysosomes Mitochondria are though to have originated when a procaryote was engulfed by a large eucaryotic cell symbiosis a. b. c. d. Two membranes Do not participate in the vesicle traffic Have their own small genomes Can make some of their own proteins similar to bacteria Chloroplasts almost certainly evolved from engulfed bacteria Membrane-enclosed organelles import proteins 1. Require energy by one of three mechanisms 2. Sorting signal: amino acid sequence nuclear pores (folded proteins) protein translocators few proteins are synthesized on ribosomes inside these organelles (unfolded proteins) Proteins sorting Cytosol Protein ER Mitochondria Chloroplasts Interior of the nucleus Golgi apparatus Lysosomes Endosomes Nuclear membranes Plasma membrane (folded proteins) 15-60 amino acids KDEL 1. Physical properties (hydrophobicity) 2. Charged amino acids Signal sequences direct proteins to the correct organelle The outer nuclear membrane is continuous with the ER A finely woven meshwork of protein filaments (lamin) Molecules in the cell membrane About 30 different proteins (1) Small water soluble molecules → pass freely and non-selectively (2) Large molecules (RNAs, proteins) & macromolecular complexes (ribosomal subunits) → sorting signal Proteins bound for the nucleus are actively transported through nuclear pores Several positively charged lysines or arginines In their fully folded conformation The energy supplied by GTP hydrolysis drives nuclear transport Proteins are imported into mitochondria in an unfolded form Unfolded form Chaperone proteins Other sorting signals The endoplasmic reticulum is the most extensive membrane network in eucaryotic cells ER Nucleus Ribosome A common pool of ribosomes is used to synthesize both the proteins that stay in the cytosol and those that are transported into membrane-enclosed organelles, including the ER An ER signal sequence and an SRP direct a ribosome to the ER membrane A soluble protein crosses the ER membrane and enters the lumen A single-pass transmembrane protein is integrated into the ER membrane A double-pass transmembrane protein uses an internal star-transfer sequence to integrate into the ER membrane Membrane-based vesicle transport Vesicles bud from one membrane and fuse with another, carrying membrane components and soluble proteins between cell compartments (digestion) (1) endosome: the main sorting station in the inward endocytotic pathway (2) trans Golgi: the main sorting station in the outward secretory pathway Clathrin molecules form basketlike cages that help shape membrane into vesicles Clathrin-coated pit Clathrin-coated vesicle Golgi Plasma membrane Clathrin-coated pits and vesicles Clathrin-coated vesicles transport selected cargo molecules COP (coat protein)-coated vesicles Golgi ER Rab proteins and SNAREs help direct transport vesicles to their target membranes Vesicle-specific SNAP receptor or Vesicle-soluble NSF attachment protein receptor Target membrane-specific SNAP receptor SNARE proteins play a central role in membrane fusion H2O out 1.5 nm Many proteins are glycosylated in the ER Asparagine-X-Serine Asparagine-X-Threonine Oligosaccharide protein transferase (oligosaccharyl transferase) Asparagine 14 sugars N (NH2)-linked glycosylation Chaperones prevent misfolded or partially assembled proteins from leaving the ER ER retention signal Misfolded proteins in the ER lumen trigger the production of chaperones and the expansion of the ER Unfolded protein response (UPR) ER stress and UPR The Golgi apparatus is made of a stack of flatted, membrane-enclosed sacs ER entry 3-20 cisterna exit Plasma membrane Several methods can be used to determine whether a protein bearing a particle signal sequence is transported into a preparation of isolated organelles Temperature-sensitive mutants have been used to dissect the protein secretory pathway in yeast GFP fusion allows proteins to be tracked GFP-viral coat protein throughout the cell ER ER Golgi Golgi → Plasma membrane Exocytosis In secretory cells, the regulated and constitutive pathways of exocytosis diverge in the trans Golgi network Exocytosis Default pathway Regulated exocytosis pathway Hormones Mucus Digestive enzymes Secretory vesicles package and discharge concentrated aggregates of protein A pancreatic cell Endocytosis Phagocytic cells (cellular eating) All eucaryocytic cells (cellular drinking) < 150 nm > 250 nm Molecules in the cell membrane Phagocytosis pseudopod Red blood cell macrophage neutrophil Pinocytosis by clathrin-coated pits and vesicles LDL enters cells via receptor-mediated endocytosis e.g. B12, iron, virus Cholesterol + proteins Acidic environment PH = 5-6 The fate of the receptor proteins involved in endocytosis depends on the type of receptor Materials destined for degradation follow different pathways to the lysosome amino acids sugars nucleotides H+ ATPase Materials destined for degradation follow different pathways to the lysosome