* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download An example of HDLSS: Microarray data
Transposable element wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Genomic library wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Metagenomics wikipedia , lookup
DNA vaccination wikipedia , lookup
Non-coding DNA wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Genomic imprinting wikipedia , lookup
Primary transcript wikipedia , lookup
Minimal genome wikipedia , lookup
Gene therapy wikipedia , lookup
Point mutation wikipedia , lookup
Epigenomics wikipedia , lookup
Genetic engineering wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Genome (book) wikipedia , lookup
Genome evolution wikipedia , lookup
Oncogenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Ridge (biology) wikipedia , lookup
Genome editing wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Gene expression programming wikipedia , lookup
Helitron (biology) wikipedia , lookup
History of genetic engineering wikipedia , lookup
Microevolution wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
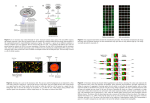
Statistical Analysis of DNA
Microarray.
An Example of HDLSS in Genetics.
The Data
Expression Matrix
• Rows represent genes
= feature vectors.
• Columns represent
different cell samples.
Ex: cancer cells from
different patients.
• Each element (i,j) of
the array represents
the expression level of
gene i in cell sample
j.
Goal of Analysis of Expression
Matrix
• Some statistical methods applied to:
1. “Group” similar genes together => groups
of functionally similar genes.
2. “Extract” representative gene in each
group.
3. ”Group” similar cell samples together.
Overview DNA Microarray
Technology
• One cell sample.
• Level of expression.
• Microarray technique.
Getting the Data... One Cell Sample at a Time
Getting the Data…measuring the Level of
Expression Gene by Gene.
• Each spot in this DNA
microarray represents the
level of expression of a
single gene in the tumor
cell compared to a
reference cell.
• Standardize the level of
expression of this cell to
make it comparable to
other cells.
Expressed in reference cell.
Expressed in reference and tumor cell.
Expressed in tumor cell
Nor expressed.
Level of Expression … mRNA
Level of Expression …mRNA
• All the cells contain the same DNA = same
genes, but in one cell not all genes are
active.
• What differentiate the cells is what genes
are active or expressed.
• To measure the cell expression we measure
the genetic molecule “RNA messenger”
denoted by mRNA.
Measuring The Level of Expression
… Complementary Strands
RNAm … DNA
• RNAm is one strand
copy of a piece of
DNA.
• Highly unstable.
• DNA is double
stranded, one strand
complementary to the
other.
• Stable.
Getting One Sample … Microarray Technique
Microarray Technique (Cont.)…The Microarray
Microarrays are made from a collection of purified DNA's. A
drop of each type of DNA in solution is placed onto a
specially-prepared glass microscope slide by an arraying
machine. The arraying machine can quickly produce a regular
grid of thousands of spots in a square about 2 cm on a side,
small enough to fit under a standard slide coverslip. The DNA
in the spots is bonded to the glass to keep it from washing off
during the hybridization reaction
Microarray Technique (Cont.) …Description of
the Method
• Definition of Microarray from the National Human
Genome Research Institute :
“…The method uses a robot to precisely apply droplets
containing functional DNA to glass slides. Researchers
then attach fluorescent labels to DNA from the cell they
are studying. The labeled probes are allowed to bind to
complementary DNA strands on the slides. The slides are
put into a scanning microscope that can measure the
brightness of each fluorescent dot; brightness reveals how
much of a specific DNA fragment is present, an indicator
of how active it is.”
Microarray Technique (Cont.) …The Method
Step by Step
• First step : to measure the gene expression level of a cell,
collect RNAm from the cell of interest, usually cancer
cell. Have the same quantity of RNAm from a “reference
cell”.
• Second step: RNAm to cDNA.
The RNAm is highly unstable, to stabilize it we
complement the strand and create cDNA(complementary
DNA) .
• Third step: creates cDNA probes.
Label cDNA from each cell by fluorescent dyes. A
differently colored fluor is used for each sample.
Microarray Technique …The Method Step by
Step (Contd.)
•
Fourth step: hybridize the cDNA probes from the two
samples to the Microarray. Once the cDNA probes have
been hybridized to the array and any loose probe has been
washed off, the array must be scanned to determine how
much of each probe is bound to each spot.
Statistical Methods
• Clustering.
• Gene shaving
algorithm: use of
PCA for clustering.
Clustering Overview
- Kmean clustering.
- Hierarchical clustering.
- Validation method.
What Is Clustering?
For a sample of size n
described by a ddimensional feature space,
Clustering is a procedure
that:
1. Divide the d-dimensional
feature space in k disjoint
groups.
Illustration for n = 45, d = 2
and k = 3.
2. Data points within each
group are more similar to
each other than to any data
point in other groups.
Similarity Between Feature Vectors
• Choice of the similarity function depends on the data. For
example: if data is invariant by linear transformation or
rotation than the similarity function has to be invariant too.
Similarity function could be a distance or an inner product.
• Examples of similarity functions:
1 Euclidean distance, used to illustrate for d = 2.
2 Correlation is used for microarray data.
K-means Clustering
• Divide the d dimensional
feature space on “k” parts
described by Voronoi
partition of the k mean
vectors.
• Algorithm finds the vector
of means of clusters.
Illustration for d =2 and k = 3,
red points represent means of clusters
and red lines represent Voronoi
partition.
Algorithm for K-means Clustering
•
Algorithm
1.
Begin initialize n, k, m1,
m2,..., mk
Do classify n samples
according to nearest mi
recompute mi
until no change in mi
return m1, m2,..., mk
end
2.
3.
4.
5.
6.
•
For d = 2, illustration of the trajectories of
the 3 means.
Computational Complexity
O(ndkT) T is the number
of iterations
K-mean Clustering for Microarray Data
• Cf picture k.mean.
• K-means clustering of lymphoma data. Lymphoma
profiles were clustered using the expression of 148
germinal-center-specific genes and Euclidean distance
metric.(a) represents the germinal-cell subtype; and (b)
represents the activated subtype. Each column represents a
specific gene and each row a specific cancer profile.
Hierarchical Clustering
Dendrogram
Venn Diagram of
Clustered Data
Hierarchical Clustering (Cont.)
• Multilevel clustering, at level 1 we have n clusters
and at level n we have one cluster.
• Agglomerative HC: starts with singleton and
merge clusters.
• Divisive HC :starts with one sample and split
clusters.
Hierarchical Clustering …Nearest Neighbor
Algorithm
• Nearest Neighbor Algorithm is an agglomerative
HC (bottom-up).
• The algorithm starts with n nodes (n is the size of
our sample). At every level the 2 most similar
nodes are merged together into one node. The
algorithm stops when we get the desired number
of clusters.
Nearest Neighbor, data to cluster.
Nearest Neighbor, Level 2, k = 7 clusters.
Nearest Neighbor, Level 3, k = 6 clusters.
Nearest Neighbor, Level 4, k = 5 clusters.
Nearest Neighbor, Level 5, k = 4 clusters.
Nearest Neighbor, Level 6, k = 3 clusters.
Nearest Neighbor, Level 7, k = 2 clusters.
Nearest Neighbor, Level 8, k = 1 cluster.
Results of Hierarchical Clustering on Microarray
Data
• Grouping similar functional genes.
• Grouping similar cell samples.
•
Cf picture Perou.trend.review2001.pdf file page6.
Criterion Function for Clustering
•
Criterion Functions depend on grouping and
number of clusters. Examples are:
1. Sum of squared errors || x - mi || 2.
2. Scatter Criteria |SW| / |ST| ; where
ST=SW+SB .
i.e. decompose the total scatter matrix into betweencluster scatter matrix and within-cluster scatter
matrix.
• Best cluster minimizes the criterion.
Gene Shaving
• The “gene shaving” method is also a
method of clustering genes and sample
cells. But unlike classic clustering, in this
method one gene could belong to more than
one cluster.
Gene Shaving Iteration
Gene Shaving…iteration
1. Start with the entire expression
matrix X, each row centered to
have zero mean.
2. Compute the leading PC of the
rows of X.
3. Shave off the proportion alpha
(10%) of the genes having
smallest absolute inner-product
with the leading PC.
4. repeats steps 2 and 3 until only
one gene remains.
5. This produces a nested sequence
of gene clusters Sn... Sk … S 1
where Sk denotes a cluster of k
genes. Estimates the optimal
cluster size k using the gap
statistic.
6. Orthogonalize each row of X
with respect to Sk , the
average gene in Sk , optimal
from step5.
7. Repeat steps 1-5 with
orthogonalized data, to find the
second optimal cluster. This
process continued until a max
of M clusters are found.
To Estimate Cluster Size : Gap Estimate
For cluster Sk let Dk be the scatter estimate. i.e
Dk = 100 SB/ST.
• For b in {1,…,B}, let
1. X * (b) permuted data matrix ( permuting the
elements within each row of X ).
2. Dk* (b) is the scatter estimate for cluster Sk *(b).
• Dk* is the mean of Dk* (b)’s.
• Gap(k) = Dk - Dk* .
• Choose k that produces the largest gap.
•
Gene Shaving (Cont.)
The first three gene clusters
found for the DLCL data
Gene Shaving (Cont.)
Percent of gene variance explained by
first j gene shaving column averages (j
= 1,2,... 10) (solid curve), and by first
j principal components (broken
curve). For the shaving results, the
total number of genes in the first j
clusters is also indicated.
Gene Shaving ( Cont.)
a) Variance plots for real and
randomized data. The percent
variance explained by each
cluster, both for the original
data, and for an average over
three randomized versions. (b)
Gap estimates of cluster size.
The gap curve, which highlights
the difference between the pair
of curves, is shown.
References
• Pattern Classification Richard O.Duda, Peter E.Hart and
David G.Stork Chapter 10.
• ‘Gene Shaving’ as a method for identifying distinct sets of
genes with similar expression patterns T. Hastie, R.
Tibshirani, M.B. Eisen, A Alizadeh, R. Levy,L Staudt, W.C
Chan, D.Botstein and P. Brown. Genome Biology 2000.
http://genomebiology.com/2000/1/2/research/0003/#B14.
• Cluster analysis and display of genome-wide expression
patterns, PNAS (1998).
References
• Basic microarray analysis: grouping and feature reduction.
S. Raychaudhuri, P.Sutphin, J.T. Chang and Russ B. Trends
in Biotechnology 2001.
• Tumor classification using gene expression patterns from
DNA microarrays.Charles M. Perou, Patrick O.Brown and
David Botstein. Trends in Molecular medicine ,December
2000.
• Pictures and definition of microarray technology from
National Human Genome Research Institute