* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Finding Genes
Epigenetics in learning and memory wikipedia , lookup
Oncogenomics wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Primary transcript wikipedia , lookup
Public health genomics wikipedia , lookup
Transposable element wikipedia , lookup
Gene desert wikipedia , lookup
Metagenomics wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Expanded genetic code wikipedia , lookup
Human genome wikipedia , lookup
Gene expression programming wikipedia , lookup
Frameshift mutation wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Point mutation wikipedia , lookup
Essential gene wikipedia , lookup
Genetic engineering wikipedia , lookup
Pathogenomics wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Non-coding DNA wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genome editing wikipedia , lookup
Genomic imprinting wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Ridge (biology) wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Minimal genome wikipedia , lookup
Genome (book) wikipedia , lookup
Gene expression profiling wikipedia , lookup
Helitron (biology) wikipedia , lookup
Designer baby wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genome evolution wikipedia , lookup
Microevolution wikipedia , lookup
An Introduction to Bioinformatics
Finding genes in prokaryotes
AIMS
To establish the concept of ORFs and their relationship to genes
To describe the features used by software to find ORFs/genes
To become familiar with Web-based programmes used to find
ORFs/genes
OBJECTIVES
To be able to distinguish between the concepts of ORF and gene
Use ORF Finder to find ORFs in prokaryotic nucleotide sequences
Usually the primary challenge that follows the sequencing of
anything from a small segment of DNA to a complete genome
is to establish where the location functional elements such as:
genes (intron/exon boundaries)
promoters,
terminators etc
DNA sequences that may potentially encode proteins are called
Open Reading Frames (ORFs)
The situation in prokaryotes is relatively straightforward since
scarcely any eubacterial and archaeal genes contain introns
FINDING ORFs
The simplest method in prokaryotes is to scan the DNA for
start and stop codons
The DNA is double stranded and each strand has three
potential reading frames (codons are groups of 3 bases)
THE CAT ATE THE RAT
Frame 1
T HEC ATA TET HER AT Frame 2
TH ECA TAT ETH ERA T Frame 3
The scan must look at all 6 reading frames
Any region of DNA between a start codon and a stop codon in
the same reading frame could potentially code for a polypeptide
and is therefore an ORF
Start AUG (methionine)
Stop UAA UAG UGA
small potential coding sequences like this will occur frequently
by chance, and therefore the longer they are the more likely
they are to represent real coding regions, genes
Problems
Small genes may be missed
The actual start codon may be internal to the ORF
There may be overlapping genes
The simplest tool for finding ORFs is ORF Finder at NCBI
It simply scans all 6 reading frames and shows the position of
the ORFs which are greater than a user defined minimum size
The genetic code used for the analysis can be altered by the
user
This would be important if e.g. mitochondrial or ciliate nuclear
DNA were being analysed
To overcome the limitations of ORF finder, more sophisticated
programmes detect compositional biases and increase the
reliability of gene detection
These compositional biases are regular, though very diffuse,
And arise for a variety of reasons:
many organisms there is a detectable preference for G or C
over A and T in the third ("wobble") position in a codon
all organisms do not utilize synonymous codons with the same
frequency - consequently there is a codon bias
there is an unequal usage of amino acids in proteins sufficient to
cause a bias in all three positions of codons and increase the
overall codon bias
the %GC content of the first two codon positions of the
universal genetic code is approximately 50%, therefore,
organisms which have a low or high %GC content will exhibit
a marked bias at the third position of codons to achieve their
overall %GC content
The most recent approaches to using compositional features
to distinguish coding from non-coding regions employ ‘Markov
models’
such approaches include the popular GENEMARK and
GLIMMER programs
An Introduction to Bioinformatics
Finding Genes in Eukaryotes
AIMS
To establish the concept of ORFs and their relationship to genes
To describe the features used by software to find ORFs/genes
To become familiar with Web-based programmes used to find
ORFs/genes
To describe the complications of the eukaryote “signals”
To be aware of the Web-based programmes
OBJECTIVES
To be able to distinguish between the concepts of ORF and gene
Use ORF Finder to find ORFs in prokaryotic nucleotide sequences
To be able to use the eukaryote programmes for a number of
organisms
Organisms whose cells have a membrane-bound
nucleus and many specialised structures located within
their cell boundary.
In these organisms, genetic material is organized into
chromosomes that reside in the nucleus.
Principles
• Content - codon usage
– often species or class specific
• Signals - PWMs
– principle is the same, signals are different
– Complication of introns/exons
Eukaryotic promoter
-110
5’
-40
CAAT box
GC box
-25
+1mRNA
TATA box
In addition - transcription factor binding sites
Genes can be enormous!
Controlled by “distant” enhancers
3’
Signals on the mRNA
Polyadenylation sequence
AUG
AAUAA
STOP
AAAAA…...
~ 12bp polyA
Kozak sequence
At translational start
Introns and Exons
Chicken 12 collagen gene
has - 38 kb > 50 Introns
Muscular Dystrophy gene is 2.5 Mb and has
? Exons!
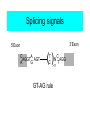
Splicing signals
3’Exon
5’Exon
C
A
AGGT AGT
A
G
()
T
C
C
N AGG
T
>11
GT-AG rule
Exon finding
• Initial exons, from the initiation codon to the first
splice site;
• Internal exons from splice site to splice site;
• Terminal exons from splice site to stop codon;
• Single introns corresponding to uninterrupted,
intronless genes, i.e., running from initiation codon to
stop codon.
Intergrated Gene Parsing
• Search for signals
• Perform a content analysis
• Define the intron/exon boundaries
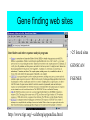
Gene finding web sites
>25 listed sites
GENSCAN
FGENES
http://www.tigr.org/~salzberg/appendixa.html