* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download What is metabolic engineering?
Biosynthesis wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Gene expression wikipedia , lookup
Genomic library wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Epitranscriptome wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Biochemistry wikipedia , lookup
Expression vector wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Paracrine signalling wikipedia , lookup
Microbial metabolism wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Biochemical cascade wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Basal metabolic rate wikipedia , lookup
Genetic engineering wikipedia , lookup
Citric acid cycle wikipedia , lookup
Gene regulatory network wikipedia , lookup
Pharmacometabolomics wikipedia , lookup
Metabolomics wikipedia , lookup
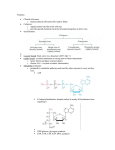
Metabolic Engineering and Systems Biotechnology Ka-Yiu San Departments of Bioengineering Departments of Chemical Engineering Rice University Houston, Texas SOME MILESTONES 1968 Nirenberg, Khorana, and Holley awarded Nobel Prize for elucidating genetic code. 1970 First restriction endonuclease isolated. 1972 DNA ligase joins two DNA fragments, creating first recombinant DNA molecules. 1973 DNA inserted into plasmid vector and transferred to host E. coli cell for propagation; cloning methods established in bacteria. Potential hazards of recombinant DNA technology raise concerns. 1976 National Institutes of Health prepares first guidelines for physical and biological containment; DNA sequencing methods developed. 1977 Genentech, the first biotechnology firm, established. Introns discovered. mRNA Protein Restriction cleavage Recombined plasmid Transcription Restriction sites Restriction cleavage Gene of interest Cloning for rProtein production Cloning vector Host cell Recombinant proteins by microorganisms Some early products Year 1982 Products Humulin (synthetic insulin) Disease Type 1 diabetes Company Genetech, Inc. 1985 Protropin Growth hormone Deficiency Genetech, Inc. Examples of a few biopharmaceutical products in 1994 Biopharmaceutical Disease Annual Sales ($ millions) Erythropoietin (EPO) Anemia 1,650 Factor VIII Hemophilia 250 Human growth Hormones Growth deficiency, renal insufficiency 450 Insulin Diabetes 700 Source: Biotechnology Industry Organization, Pharmaceutical Research and Manufacturers of America, company results, analyst reports What is metabolic engineering? Metabolic engineering is referred to as the directed improvement of cellular properties through the modification of specific biochemical reactions or the introduction of new ones, with the use of recombinant DNA technology Modern biology – central dogma Gene Protein/ enzyme mRNA transcription translation Current metabolic engineering approaches • • • • Amplification of enzyme levels Use enzymes with different properties Addition of new enzymatic pathway Deletion of existing enzymatic pathway Genetic manipulation Gene Protein/ enzyme mRNA transcription translation Current projects 1. Cofactor engineering of Escherichia coli A. Manipulation of NADH availability B. Manipulation of CoA/acetyl-CoA NADH (Reduced) NAD+ (Oxidized) 2. Plant metabolic engineering 3. Quantitative systems biotechnology A. B. C. D. Rational pathway design and optimization Metabolic flux analysis based on dynamic genomic information Design and modeling of artificial genetic networks Metabolite profiling 4. Genetic networks – architectures and physiology Current Projects Pathway and Cofactor Metabolic Engineering I 1 2 II An integrated metabolic engineering study of evolved alcohol acetyl transferase enzymes in flavor compound formation in E. coli (with Dr. Bennett) NSF BES-0118815 USDA 2002-35505-11638 Plant Metabolic Engineering 3 III Collaborative research: Metabolic engineering of hairy roots for alkaloid production (with Dr. Gibson of UM and Dr. Shanks of Iowa State University) NSF BES-0224593 Quantitative Biosystems Engineering 4 Experimental driven computational analysis of E. coli global redox sensing/ regulatory networks and cellular responses (with Drs. Bennett amd Cox) NSF BES-0222691 5 Collaborative research: Metabolic engineering of E. coli sugar-utilization regulatory systems for the consumption of plant biomass sugars (with Drs. Gonzalez and Shanks of Iowa State University) 6 Modeling and design of gene switching networks for optimal control of PHA nanostructures (with Drs. Mantzaris and Bennett,) BES0331324 From Genetic Architecture to Adaptation Dynamics (with Drs. Mantzaris – PI, Bennett, and Zygourakis). NIH R01GM071888 7 IV EPA RD-83144101 NSF Instrumentation 8 MRI: Acquisition of Multiple Instruments for Research and Education 9 Shimadzu Instrumentation Grant NSF BES-0420840 Cofactor engineering Motivations and hypothesis Motivations • Existing metabolic engineering methodologies include – pathway deletion – pathway addition – pathway modification: amplification, modulation or use of isozymes (or enzyme from directed evolution study) with different enzymatic properties • Cofactors play an essential role in a large number of biochemical reactions Hypothesis Cofactor manipulation can be used as an additional tool to achieve desired metabolic engineering goals Importance of cofactor manipulation Enzymes + Cofactors Substrate Products Cofactor engineering • NAD+/NADH • CoA/acetyl-CoA NADH/NAD+ Cofactor Pair • Important in metabolism – Cofactor in > 300 red-ox reactions – Regulates genes and enzymes • Donor or acceptor of reducing equivalents • Reversible transformation NADH (Reduced) NAD+ (Oxidized) • Recycle of cofactors necessary for cell growth Coenzyme A (CoA) • Essential intermediates in many biosynthetic and energy yielding metabolic pathways • CoA is a carrier of acyl group • Important role in enzymatic production of industrially useful compounds like esters, biopolymers, polyketides etc. Acetyl-CoA • Entry point to Energy yielding TCA cycle • Important component in fatty acid metabolism • Precursor of malonyl-CoA, acetoacetyl-CoA • Allosteric activator of certain enzymes Example: Lactic acid formation Lactic acid Polylactic acid (PLA) LDH Pyruvate NADH Lactate NAD+ Biopolymer production Poly(3-hydroxybutyrate- co-3(PHB/PHV block copolymer) hydroxyvalerate) Glycerol Propionate Acetyl-CoA Propionyl-CoA Acetyl-CoA 3-Ketothiolase (PhaA) HSCoA Acetoacetyl-CoA 3-Ketovaleryl-CoA NADPH Acetoacetyl-CoA Reductase (PhaB) NADP+ 3-Hydroxybutyryl-CoA 3-Hydroxyvalery-CoA PHA Synthase (PhaC) HSCoA P(HB-co-HV) HSCoA Polyketide production • Complex natural products • > 10,000 polyketides identified • Broad range of therapeutic applications • Cancer (adriamycin) • Infection disease (tetracyclines, erythromycin) • Cardiovascular (mevacor, lovastatin) • Immunosuppression (rapamycin, tacrolimus) 6-deoxyerythronolide B Polyketide production Precursor supply - example Ref: Precursor Supply for Polyketide Biosynthesis: The Role of Crotonyl-CoA Reductase, Metabolic Engineering 3, 40-48 (2001) Approach Systematic manipulation of cofactor levels by genetic engineering means Model systems Simple model systems, such as biosynthesis of succinate and ester, to illustrate the concept Results • increased NADH availability to the cell • increased levels of CoA and acetyl CoA • significantly change metabolite redistribution Manipulation of NADH availability Fermentation Pathway of E. coli Glucose NAD+ NADH Succinate 2NAD+ 2NADH Pyruvate Lactate NADH Formate NAD+ Acetyl-CoA 2NADH 2NAD+ Ethanol Acetate NADH Regeneration Pyruvate NADH NAD+ Formate CO2 FDH1 PFL Acetyl-CoA FDHF CO2 H2 original NAD independent pathway (FDHF: formate dehydrogenase, NAD independent) Newly added NAD+ dependent pathway (FDH1: NAD+ dependent formate dehydrogenase FDH1 encoded by fdh1 from Candida boidinii) Construction of pSBF2 Overexpressing FDH pFDH1 PCR fdh fdh pSBF2 XbaI pSBF2 fdh fdh EcoRI/XbaI pUC18 pUCFDH XbaI pDHK30 pDHK30 Assay of FDH activity Strain FDH activity (units/mg protein) GJT001(pSBF2) 0.42 BS1(pSBF2) 0.28 GJT001(pDHK29) Not detected BS1(pDHK30) Not detected Characterization of NADH-dependent FDH PanK NADH-dependent FDH PanK NADH-dependent FDH XbaI lacZ' lacZ MCS KmR pDHK29 pSBF2 fdh KmR Ori Ori GJT (pDHK29) (Control strain) GJT (pSBF2) (New strain) Anaerobic Tubes : Experimental Method • Strains : Escherichia coli (MC4100 derivative) – GJT001 (pDHK29): wild type (control plasmid) – GJT001 (pSBF2): wild type (new FDH plasmid) • Media: – LB + 1g/L NaHCO3 – 100mg/L Kanamycin – 20g/L Glucose • • • • Temperature: 37 ºC Agitation: 250 rpm Samples: 72 hrs after inoculation HPLC Effect of Increasing NADH Availability % of Increase/Decrease for GJT001 (pSBF2) relative to GJT001 (pDHK29) 3-fold Glucose Consumed Succinate 55% NAD+ NADH 2NAD+ 2NADH Pyruvate NAD H Formate Converted 8-fold NADH CO2 NAD+ Lactate 91% Acetyl-CoA 2NADH 2NAD+ Acetate 43% NAD+ Formate FDH1 FDHF CO2 H2 Ethanol 15-fold O.D.600nm: 59% Et/Ac: 27-fold mol NADH/mol glucose NADH Availability 5.0 4.0 3.0 2.0 1.0 0.0 GJT(pDHK29) GJT(pSBF2) Ethanol Concentration (mM) Ethanol Concentration (reduced product) 200 180 160 140 120 100 80 60 40 20 0 GJT001(pDHK29) GJT001(pSBF2) Summary of results Effect of NADH regeneration (overexpressing NAD+-dependent FDH): – Increases intracellular NADH availability – Provide a more reduced environment – Increase reduced product (such as ethanol and succinate) productivity significantly Quantitative systems biotechnology Projects 1. Metabolic flux analysis based on dynamic genomic information 2. Rational pathway design and optimization - feasible and realizable new network design 3. Design and modeling of artificial genetic networks Motivations Observations Traditional reductionist approach • Knowledge at the basic and fundamental level – but mostly isolated Information overflow • Genome database, gene expression database (functional genomic), proteomic, metabolomics, metabolic pathway database Most of the existing data base – static • Genome database, metabolic pathway database Motivations and objectives: How can one utilize the static genomic and metabolic databases (especially when genetic/regulatory network structures are available) to describe and predict cellular functions, such as metabolic patterns? Traditional flux balance analysis (FBA) Genome Database Pathway Database A priori Knowledge Metabolic Network FBA Metabolic Pattern Metabolic Network (From http://www.genome.ad.jp/kegg/pathway/map/map00020.html) Metabolic Pattern (Illustration) 1.0 0.8 0.2 0.8: Metabolic rates (From http://www.genome.ad.jp/kegg/pathway/map/map00020.html) genotype phenotype genetic environmental perturbations perturbations (mutant strains) Transcription Translation Metabolic Flux Analysis Gene mRNA Protein/ enzyme Stimuli traditional metabolic engineering study Cellular Responses OR Metabolite Patterns Proposed New Approach Genome Database Pathway Database A priori Knowledge Genetic Structure Metabolic Network FBA Metabolic Patterns ? Expression Patterns Genetic Network Environmental Conditions Gene Regulation Knowledge Gene Chip (Array) Data Model System • Oxygen and redox sensing/regulation system • Sugar utilization regulatory network Simplified schematic of E. coli central metabolic pathways Glucose PEP Pyruvate ppc CoA NADH, CO2 Formate [4.1.1.31] pdh [1.1.1.28] H2 + CO2 pfl [1.2.4.1] CO2 Lactate ldhA NAD+,CoA [2.3.1.54] Acetyl- CoA Ethanol gltA aspC [4.1.3.7] Oxaloacetate NADH [2.6.1.1] NAD+ [1.1.1.37] NAD+ Aspartate acnB mdh NADH Acetate Citrate [4.2.1.3] Isocitrate Malate NADP+ aspA fumB fumA icd [4.3.1.1] [4.2.1.2] [4.2.1.2] [1.2.4.2] NADPH Fumarate frdABCD [1.3.1.6] sdhCDAB NADH NAD+ CO2 [1.3.99.1] Succinate sucCD [6.2.1.5] sucAB 2-ketoglutarate [1.2.4.2] NAD+ NADH Succinyl-CoA CO2 Schematic showing selected oxygen and redox sensing pathways in E. coli (adopted from Sawers, 1999) Cytoplasmic membrane FNR FNR e- transport Redox, metabolites ArcB P Redox? Aer Dos ArcA O2 ArcA-P CheW,A,Y Transcription O2 unknown Energy taxis Transcription Some example of available pathway information Recommended Name EC number Reactions pyruvate dehydrogenase complex 1.2.4.1 Acetyl-CoA + CO2 +NADH = CoA + pyruvate + NAD aceEF ArcA(-) FNR(-) 1,3 4 2.3.1.54 CoA + pyruvate = acetyl-CoA + formate pfl ArcA(+) FNR(+) 2 1 citrate synthase 4.1.3.7 Acetyl-CoA + H2O + oxaloacetate = citrate + CoA gltA ArcA(-) 1,3 fumarate hydratase (fumarase) 4.2.1.2 fumarate + H2O = (S)-malate fumA FNR(0) 1 fumarate hydratase (fumerase) 4.2.1.2 (S)-malate = fumarate + H2O fumB FNR(+) 1,2 pyruvate formate-lyase Encoded by Effect Ref succinate dehydrogenase 1.3.99.1 Succinate + acceptor = fumarate + reduced acceptor sdhCDAB ArcA(-) FNR(-) 1,2,3 2 fumarate reductase 1.3.1.6 Fumarate + NADH = succinate + NAD+ frdABCD ArcA(+) FNR(+) 1 1,2,4 FNR active in the absence of oxygen; ArcA is activated in the absence of oxygen Ref 1: “Reg of gene expression in fermentative and respiratory systems in Escherichia coli and related bacteria”, E.C.E. Lin and S. Iuchi, . Annual Rev. Genet, 1991, 25:361-87Ref 2: Ref 2 “O2-Sensing and o2 dependent gene regulation in facultatively anaerobic bacteria”, G. Unden, S. Becker, J. Bongaerts, G.Holighaus, J. Schirawski, and S. Six, Arch Microbi. (1995) 164:81-90 Ref 3: “Regualtion of gene expression in E. coli” E.C.C. Lin and A.S. Lynch eds. (1996) Chapman & Hall, New York (p370) Ref 4: “Regualtion of gene expression in E. coli” E.C.C. Lin and A.S. Lynch eds. (1996) Chapman & Hall, New York (p322) ldhA aceB mqo aspA fumB frdABCD pfl cyd cyo ArcB ArcA FNR fumC aceEF acnB sdhCDAB fumA mdh gltA icd sucAB sucCD We have 3 sensing/regulatory components whose activity evolves according to the Boolean mapping coded in the figure. Here green red denotes repress and denotes activate. When two components regulate a third we suppose their action to be an “and”. These regulatory components determine the O2 ArcA Stimulus FNR Sensors/regulators aceEF pfl genes PDH PFL enzymes CO2 CoA NADH NAD+ Acetyl-CoA formate Metabolites pyruvate activation repression Work in progress To develop a model that can provide dynamic and automatic adaptation of pathway map to environmental conditions Biosystems • Systems biology is the study of living organisms at the systems level rather than simply their individual components • High-throughput, quantitative technologies are essential to provide the necessary data to understand the interactions among the components • Computation tools are also required to handle and interpret the volumes of data necessary to understand complex biological systems genotype phenotype genetic environmental perturbations perturbations (mutant strains) Gene mRNA Protein/ enzyme Stimuli Functional Genomics Genomics Cellular Responses OR Metabolite Patterns Metabolomics Proteomics Functional Genomics Proteinomics • 2D gel electrophoresis • Mass spectrometry • Bioinformatics • Protein "chips" 2D gel electrophoresis • IEF • Size Protein Chips • The basic construction of such protein chips has some similarities to DNA chips, such as the use of a glass or plastic surface dotted with an array of molecules. • Known proteins are analyzed using functional assays that are on the chip. For example, chip surfaces can contain enzymes, receptor proteins, or antibodies that enable researchers to conduct protein-protein interaction studies, ligand binding studies, or immunoassays • High-end quadruple TOF tandem mass spectrometers enable high-performance protein identification, epitope and phosphorylation mapping, and protein-interaction analyses. Metabolomics • Metabolomics is a relatively new discipline and techniques for high-throughput metabolic profiling are still under development. • No single technique is suitable for the analysis of all different types of molecule, so a mixture of techniques is used. • Methods such as gas chromatography, high-pressure liquid chromatography and capillary electrophoresis are used to separate metabolites according to various chemical and physical properties. The molecules are then identified using methods such as mass spectrometry. Shimadzu LCMS 2010A Shimadzu QP-2010 Collaborators Dr. George N. Bennett Department of Biochemistry and Cell Biology Rice University Dr. Steve Cox Department of Computational & Applied Math Dr. Nikos Mantzaris Department of Chemical Engineering Dr. Kyriacos Zygourakis Department of Chemical Engineering Dr. Jacqueline V. Shanks Department of Chemical Engineering Dr. Ramon Gonzalez Department of Chemical Engineering Dr. Sue Gibson Department of Plant Biology Recent Graduates Aristos Aristidou, Ph.D. Cargill Dow Chih-Hsiung Chou, Ph.D. University of Waterloo, Canada Peng Yu, Ph.D. BMS Derek Sykes, M.S. Life Technology Irena Ying Chen, M.S. Kellog Yea-Tyng Yang, Ph.D. M.I.T. Susana Joanne Berrios Ortiz, Ph.D Shell Development Erik Hughes, Ph.D Wyeth Ravi Vadali Eli Lilly Valentis, Inc. Current Lab Members Name Project Christie Peebles Plant Metabolic Engineering Sagit Shalel-Levanon Quantitative Systems Biotechnology Randeep Singh Quantitative Systems Biotechnology Ailen Sanchez Cofactor Metabolic Engineering – NAD+/NADH Cheryl Dittrich Cofactor Metabolic Engineering Henry Lin Pathway design and analysis Stephanie Portle Genetic networks Metabolic Engineering and Systems Biotechnology Laboratory Ka-Yiu San ([email protected]) Office: Lab: GRB E200K GRB E201, E202, E210, E128 Questions ? ???