* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Modulation of Retinoblastoma and Retinoblastoma
Survey
Document related concepts
Extracellular matrix wikipedia , lookup
Cell culture wikipedia , lookup
Cell growth wikipedia , lookup
Biochemical switches in the cell cycle wikipedia , lookup
Phosphorylation wikipedia , lookup
Magnesium transporter wikipedia , lookup
Cellular differentiation wikipedia , lookup
Protein (nutrient) wikipedia , lookup
Cytokinesis wikipedia , lookup
Signal transduction wikipedia , lookup
Protein moonlighting wikipedia , lookup
Protein phosphorylation wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Transcript
Vol.
6, 1463-1476,
November
1995
Cell
Growth
1463
& Differentiation
Modulation
of Retinoblastoma
and Retinoblastoma-related
Proteins in Regenerating
Rat Liver and Primary
H epatocytes’
Guangsheng
Fan, Ruiling Xu, Martin W. Wessendorf,
Xiaoming Ma, Betsy T. Kren, and Clifford J. Steer
linked to its simultaneous
suppression of cell cycledependent
kinase 4 and cyclin E protein
levels.
Departments
of Medicine
1G. F., R. X., X. M., B. T. K., C. J. 5.1 and Cell
Biology
and Neuroanatomy
IM. W. W., C. J. SI, University
of Minnesota
Medical
School,
Minneapolis,
Minnesota
55455
Introduction
Abstract
Protein expression of the retinoblastoma
(Rb) tumor
suppressor gene product was examined
by immunoblot
analysis of nuclei isolated from regenerating
rat liver
after 70% partial hepatectomy
(PH). Levels were almost
undetectable
in quiescent 0-h livers but increased 1 5- to
60-fold 3 to 24 h post-PH, 1 05-fold at 30 h, and 20- to
50-fold at 60 to 72 h post-PH. Expression returned to
near baseline levels at 1 8, 42, and 48 h post-PH. A
similar pattern of Rb protein expression in the
regenerating
liver was observed by indirect
immunofluorescence
microscopy,
with peak nuclear
expression at 30 h post-PH. Rb-related
proteins with
apparent molecular
masses of 300, 1 56, and 74 kDa
were detected in regenerating
liver using mAbs to the
Rb protein. Their expression increased 6- to 8-fold
during regeneration,
and only p1 56 returned to baseline
levels at 60 h post-PH. Rb and its related proteins were
detected in cultured primary hepatocytes,
and although
total protein levels did not change appreciably,
there
was a dramatic shift from cytosol into nuclei through 96
h. The half-life of the Rb protein was determined
to be
1 .9 h in regenerating
liver and 2.2 h in cultured primary
hepatocytes.
Rb protein abundance
in synchronized
HuH-7 human hepatoma cells was cell cycle dependent
and exhibited peak nuclear expression during S phase.
Rb protein was detected primarily
in its
hyperphosphorylated
state during liver regeneration
and
through the cell cycle of the HuH-7 cells. In vivo
administration
of transforming
growth factor 31 an
inhibitor of DNA synthesis in regenerating
liver, resulted
in reduced expression of Rb as well as its protein
partners, cell cycle-dependent
kinase 4 and cyclin E. The
results suggest that in the regenerating
rat liver and in
synchronized
HuH-7 cells, expression of Rb protein is
modulated
in a cell cycle-dependent
fashion, remains
primarily
in a hyperphosphorylated
state, and exhibits a
relatively short half-life. The inhibition
of Rb protein
expression by transforming
growth factor f31 may be
,
Received
5/19/95;
1 This
study was
Foundation
2 To
whom
revised
supported
)to C. J. S.).
requests
for
7/14/95;
in part
reprints
accepted
9/1/95.
by a grant from
should
Medicine,
UMHC
Box 36, University
Delaware
Street SE, Minneapolis,
MN
(612) 625-5620.
be
the
addressed,
of Minnesota
55455.
Phone:
Minnesota
The Rb3 gene is one of the better characterized
members of
the tumor suppressor
gene family. It was originally
identified and eventually
cloned
by virtue of its absence
in a
number of Rb tumor cell lines (1-3). Subsequent
studies
revealed that inactivation
of the Rb gene was a frequent
event in tumomigenesis.
Rb is known to play a key mole in the
regulation
of cell proliferation
(4), and it now appears to
also be involved
in the induction
of the fully differentiated
state. For example,
it has been suggested
that Rb protein,
in
association
with myogenic
factors such as Myo D, is mequmred to bring about terminal
differentiation
of muscle
cells (5).
The Rb gene encodes a nuclear phosphoprotein
of 1 10
kDa, which
is present in most cell types and which also
exists in phosphorylated
states ranging in size from 1 10 to
1 1 6 kDa (6). It is now well established
that activity ofthe Rb
protein during the cell cycle is regulated
by its level of
phosphomylation.
It is underphosphomylated
in G ; hyperphosphorylated
at the G1-S phase transition;
remains
phomylated
in 5, G2, and most of M; and reverts
undemphosphorylated
Phosphomylation
(7-9).
regulated
Whereby
state at on before
of
the
Rb
the M-G0
protein
appears
phosto an
transition
to
be
at the level of both cell growth and differentiation.
mitogenic
stimulation
activates Rb protein phos-
phomylation,
signals
of differentiation
are associated
with
dephosphomylation
of the protein
(8). In short, unphospho-
rylated Rb suppresses cell proliferation
and promotes
cellulan differentiation
in contrast to phosphorylation,
which
inactivates
Rb activity and allows the cell to enter S phase.
An important
property
in defining
its binding
domains
has been the ability of the functional
Rb protein to form
stable complexes
with the E1A protein of adenovirus
(10),
the large T antigen of polyomavmmus (1 1), and the E7 protein
of papillomavinus
(1 2). Any mutation
that affects the ability
of these oncoproteins
to bind Rb dramatically
reduces the
transformation
potential
of these small DNA tumor viruses
(1 1 , 1 3). Each of the oncoproteins
contains a short homologous, colineam sequence,
which is thought to serve as the
Rb-binding
motif (14). Interestingly,
that binding site maps
to the region of the Rb protein
that is most frequently
mutated
in tumors
(1 5, 1 6). Taken together,
the results
suggest that these DNA tumor viruses may stimulate cellular
proliferation
by binding to and sequestering
Rb protein in a
manner
that mimics its loss in naturally
occurring
tumors.
Medical
at Department
of
Medical
School,
(612) 625-8999;
516
Fax:
3 The
abbreviations
used are: Rb, retinoblastoma;
BndUrd,
5-bromo-2’-deoxyunidine;
CDK,
cell cycle-dependent
kinase;
PH, partial
hepatectomy;
TGF-l
transforming
growth
factor l ; FBS, fetal bovine
serum;
kDa, kilodalton(s).
,
1464
Rb Protein
Expression
Binding
in Regenerating
Rat Liver
to Rb as a necessary
induced
by viral
Rb protein-bound
proliferation
questening
proteins
step
oncoproteins
in the
suggests
displace
cellular proteins required for normal
cell
and/on differentiation.
By binding
to and seRb in its undemphosphorylated
state, the oncoare
thought
to
release
the
growth
posed by Rb and allow for uncontrolled
date, a large group of cellular proteins
that
transformation
that they
bind
Rb,
including
those
restraints
im-
cell growth (4). To
have been identified
involved
in transcriptional
regulation
such as E2F (1 7, 1 8), Myo D (5), Pu.1 (1 9), c-Myc
(20)
and ATF-2 (21), the signal transducing
protein p48 (22),
the structural
protein
lamin
C (23),
and specific
kinases
cellular
proteins
The level
remains
of Rb expression
to be fully
appears
characterized.
to be critical
in de-
tenmining
the status of cell growth.
Overexpression
of Rb by
DNA
transfection
on microinjection
of protein
results
in
inhibition
of cell growth
under
conditions
in which
the
wild-type
cell proliferates
normally
(28). The two populations of cells may represent
the difference
in cell function
associated
with a critical
threshold
of Rb protein.
In fact,
titration
of Rb protein
results first in a gradual decrease in
cell cycle activity and then a sudden absence of activity, as
ifa specific level ofpmotein is required to maintain cell cycle
arrest (29). Interestingly,
the Rb protein
down-regulates
its
own promoter
activity
(30), suggesting
that a threshold
effect may exist in the control
of cell growth
by Rb.
In an effort to further
characterize
its expression
during
the cell cycle, we have investigated
the abundance
and
state of phosphonylation
of Rb protein in regenerating
liven
and primary hepatocytes.
Liven regeneration
after 70% PH
is a well-characterized
in vivo model
of cell replication
(31). Growth ofthe liver is triggered by the decrease in mass
and involves a synchronized
and highly regulated
prolifemation of cells in the remaining
lobes. There is an initial peak
of DNA
synthesis
which
is followed
33). A second,
by hepatocytes
6 to 8 h later
less intense
at 20 to 24 h post-PH,
by a wave of mitosis
(32,
peak
in DNA
approximately
48 h post-PH
and reflects
manly
of nonpanenchymal
cells. However,
ported
recently
that
neously
in
cells
all
the G0-G,
of
the
transition
liven
and
synthesis
occurs
replication,
pniit has been me-
occurs
that
simulta-
G,
of
the
nonparenchymal
cells is simply
prolonged
(34). In the
young adult mat, the original
hepatic mass is usually attained
1 0 to 1 2 days after
70% PH. Once the original
mass is
restored,
the cells of the liver resume a quiescent
state.
Multiple
panacnine and autocnine
factors participate
in the
finely
single
trolling
orchestrated
regulation
of liver regeneration;
and no
factor
has been identified
as the master switch
coninitiation
and termination
of growth.
There
is a
sequential
and regulated
and delayed-early
growth
ing
liven
after
PH (34,
35).
expression
of many immediateresponse genes in the negenematThese
factors
appear
to partici-
pate in the transition
of cells from quiescence
into G.
Because of these characteristics,
the regenerating
liven after
PH provides a unique system to study Rb regulation
of cell
proliferation
and differentiation.
The present study extends the analysis of Rb transcript
modulation
In addition,
protein
in
Immunoblot
tion were
expression
of Rb protein is modulated
in a cell cycle-dependent
fashion and remains primarily
in a hypemphosphorylated
state.
Furthermore,
nuclear expression
of the Rb protein during
liver regeneration
is significantly
uncoupled
from transcript
expression
through
at least 72 h. In addition,
the
inhibition
of DNA
synthesis
in the regenerating
liven by
TGF-31
is associated
with suppression
of Rb abundance
as well
as CDK4
and cyclin
E, two
key modulators
of Rb
phosphonylation.
(24),
phosphatases
(25), and cyclins (26, 27). However,
the biological significance
of the interaction
of Rb protein with
these
phorylated
species.
The results suggest that in the negenerating mat liven and in synchronized
HuH-7 cells, expression
in the regenerating
rat liven after 70% PH (36).
it investigates
the cell cycle expression
ofthe
Rb
synchronized
HuH-7
human
hepatoma
cells.
analysis and immunohistochemical
localizaused to determine
the subcellulan
distribution
and
pattern
of the Rb protein
and its various phos-
Results
Retinoblastoma
Protein Expression
in Regenerating
Rat
Liver. In this study, expression
of the 1 1 0-kDa
Rb tumor
suppressor
gene product
was characterized
in regenerating
mat liven after 70% PH. Nuclei were isolated at varying time
points
from
post-surgery
sham
and regenerating
livers
through
and analyzed
for Rb protein
expression
72 h
using
mAbs to several different
epitopes
of the protein.
XZ1 61
provided
the strongest
signal and was used primarily
throughout
the study. Protein levels were almost undetectable in 0-h liver and remained
so until 3 h post-PH, when
theme was an increase to 50-fold by 1 2 h (Fig. 1 , A and B).
An abrupt decrease
in expression
was observed
at 1 8 h
post-PH,
followed
by a 55- and 105-fold
increase above
control levels at 24 and 30 h post-PH, respectively.
Expmession decreased
to near baseline
levels at 42 and 48 h
post-PH
and again
significantly
increased
20- and 50-fold
at
60 and 72 h post-PH,
respectively.
By 96 h post-PH,
Rb
expression
returned to near 0-h values. In addition,
protein
levels remained
almost
undetectable
in sham-operated
liv-
ens through 72 h (data not shown). Throughout
the period of
regeneration,
the Rb protein remained
primarily
in the hyperphosphonylated
state (Fig. 1 C, ppRb).
However,
during
G,, theme was a notable increase in the hypophosphorylated
Rb species until 12 h post-PH,
when the ratio returned to
baseline
levels and 90% of Rb was detected
in a hypemphosphorylated
state. A Northern
blot is provided
for cornpanison (Fig. 1A), based on the previous
report that rat liver
expresses
two Rb transcript
species 2.8 and 4.7 kb in length
(36). Again, the switch in transcript
expression
occurred
at
3 to 6 h in which the 2.8-kb species was reduced
approximately 95% and that of the 4.7-kb species increased
ap-
proximately
protein
15-fold
oven baseline
expression
occurred
by immunoblot,
in transcript
expression
levels.
In contrast
no significant
to Rb
changes
at 24 to 30 h post-PH
on
60 to 72 h.
To confirm
the
analyses in tissue,
staining
provided
The
results obtained
from the immunoblot
we used indirect
immunofluomescence
with the same mAbs to Rb (Fig. 2). Again,
XZ161
the strongest
signal and least nonspecific
staining.
results
analysis.
confirmed
In 0-h
liven,
those
determined
theme was
minimal
by Western
cytosolic
blot
staining,
and only 1 to 2% of nuclei showed positive staining for the
Rb protein.
By 1 2 h post-PH, however,
approximately
30%
of nuclei were positive for Rb staining, and by 30 h, 45% of
nuclei were immunoreactive
for Rb antigen.
In a pattern
similar to that observed
by immunoblot
analysis, very few
nuclei
were
moderate
positive
staining
at 1 8, 42, and 48 h post-PH.
was observed
However,
at 60 h, and a relatively
strong signal was detected
at 72 h post-PH,
when
approximately
25%
of nuclei
were
immunoreactive.
Cytosolic
staining
for Rb protein was most significant at 18 and 72 h
Cell
A
Time
0
0
0.5
0.5
1
1
3
3
6
Post-PH
6
12
a
(h)
24
18
24
30
42
--
48
60
0
Growth
& Ditterentiation
ppRb
72
-4.7kb
.#{149}.#{149}
#{149}OOOOO28
B
C
0..-
a
(“C
0.0
ci:
#{174}L
Time Post-PH
Time Post-PH
(h)
(h)
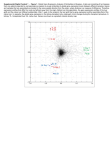
Fig. I.
Rb Protein
expreSsion
in regenerating
rat liver. Rats were subjected
to 70#{176}/
PH and sacrificed
at the indicated
times post-PH.
Nuclear
protein
extracts
‘vere isolated
and resolved
by SDS-PAGE
as described
in “Materials
and Methods.”
A: Top, immunoblot
of Rh expression
determined
using XZ161
mAb. After
transter,
the I)lOt was probed
with antibody
and detected
by the ECL method.
Bottom,
Northern
blot analysis
of Rb transcript
expression.
Poly(A( ‘ -enriched
10-pg
samples
of RNA were isolated
troni livers at the indicated
times post-PH,
processed
for blotting,
and hybridized
with a ‘2P-labeled
9W)-bp
Bgl/ll-()xaNl
fragment
of the niurine
Rb cDNA
clone,
as published
previously
(36).
Equal
lane loading
was determined
with
a 900-bp,
2P-labeled
Pstl fragment
ot the rat
asialoglycoprotein
receptor
cDNA.
Right, transcript
sizes. B, changes
in Rb protein
expression
through
72 h post-PH
relative
to 0-h controls.
Protein
levels we’re
densitometric
ally (luantitated
from the immunoblot
on which
equal amounts
of protein
were added to the lanes. C. relative
changes
in Rh phosphorlation
(luring
72 h of re’gene’ration
determined
from immunoblot
analysis.
The results are representative
of three different
experiments.
pRh, hypophosphorylate’d
Rb; ppRb.
hvpe’rpliosphorvlated
Rb.
post-PH.
niAb
F20.2F
to 3-microglobulin
was used as
control
and showed
no evidence
of nuclear
staining
(data
not shown).
In addition
to characterizing
expression
of the Rb protein
in regenerating
liver, we were interested
in comparing
the
results
to
an
additional
tumor
suppressor
gene product.
Several monoclonal
and polyclonal
antibodies
were used to
detect
p53 through
60 h of liver regeneration
(Fig. 3). The
protein
was detectable
in nuclei
isolated
from 0-h liver and
mildly
fluctuated
until 6 h post-PH,
when
it increased
to
approximately
5-fold
over baseline
levels.
Similar
abundance was noted at 1 2 h, and by 1 8 h post-PH,
expression
had returned
to control
levels.
A 3-fold
increase
was detected
at 24 h, and a dramatic
increase
to greater
than
40-fold
was
noted at 30 h. Levels returned
quickly
to those
of baseline
by 48 to 60 h post-PH.
Cell Cycle-modulated
tein in Synchronized
Expression of Retinoblastoma
HuH-7
Human
Hepatoma
ProCells.
Based on the cyclical
expression
of Rb protein
in the regenerating
liver, we investigated
in greater
detail
the cell
cycle
dependency
of Rb in a synchronized
proliferating
hepatocyte
cell. Because
of the difficulties
involved
in synchronization
of cultured
primary
hepatocytes,
we chose to
examine
expression
of the Rb gene product
in HuH-7
cells,
which
have been characterized
as a well-differentiated
human
hepatoma
cell line (37). Using
a modification
of established
methods,
we were able to synchronize
HuH-7
cells in G,
5, and M phases.
In fact, cells arrested
in G,
with hydroxyurea
exhibited
less than 8% BrdUrd
labeling,
in contrast
to those blocked
in S phase, where
97% were
labeled
with BrdUrd.
Rb protein
abundance
was examined
by immunoprecipitation
analysis
of total cell lysate during
each phase of the cell cycle,
as well
as 2 and 6 h after
S-phase release and 4 h after M-phase
release (Fig. 4A). The
results
indicated
that Rb protein
levels steadily
decreased
from S phase through
6 h after S-phase
release,
when
Rb
expression
was approximately
15% of G1 levels (Fig. 48).
Protein abundance
increased
in M phase and reached
maximum
expression
4 h after M-phase
release,
when
it was
consistently
10 to 15% greater than levels in G1. Although
maximum
expression
of hyperphosphorylated
Rh protein
was detected
during
S-phase
release,
its abundance
was 2
to 4 times greaten than the un/hypophosphorylated
protein
species
throughout
the cycle
(Fig. 4C). It is unclear
as to
whether
the molecular
weight
bands below
pRb represent
differentially
processed
Rb protein
species
found
in the cell
lystate or Rb-related proteins.
It was apparent
that Rb protein
levels
in the total cell
lysate
fluctuated
significantly
during
the cell
cycle
of
HuH-7
cells. It was important
then to examine
the nuclear
abundance
of the Rb protein
species
in the synchronized
cells to establish
whether
redistribution
of the protein
was
occurring
during the various
phases ofthe cell cycle. In fact,
Rb abundance
in nuclear
extracts
displayed
a somewhat
different
pattern
of expression
than observed
in whole
cell
lysates (Fig. 5A). Rb expression was almost 3-fold greater in
1465
1466
3
:-‘
‘A.
I
,,;
<..#{149}
...,
_‘,
-.‘
.
-..,.
. .
I
fr.
‘
.4’
r
p..
i
[B’k
S
‘p
4i
,
ib’
S
.-.I.
!
a
.
S
(I
,.
a
-.
S
I
S
.5
I
I
,L’
S
#{149}1#{149}
.b
,‘
S
S
S
I’ll’.’.
#{149}.
S
I.
‘
.
.
t
,,_4
-S
I
F
‘
Lv:
S
,
S
.
,
._#{149}sS
.4..
.-
S’-
#{149},
‘.
b-
‘
SI
I
11.1*.
1
41:?
I
#{149}41P
.
I
.
,,
.1
S
,
#{149}s ‘
“1
#{149}
.‘
-,,
1,55s
S
#{149},‘4.I
I
,
.A
‘I
I.
t
a
#{149}
..
‘
:
,‘
‘#{149}f’.
*
I.
I
P”#{231},,
‘S
,.
:
.
.
I
S
p.
-.5
t,
.-.--.
,.*..
.‘-4_
.
.,
u:wr;
-r#
S
‘:;t,
S
‘t
.b
bF:;
.\r:?:.
,
.
>“ii”I’#{149}
-
h._.(
I
,
:
S’
-
1
/
-4
..-sO
#{149}_#{149}I’
‘‘
#{149}S
I
J.
(elI
Time
0.5
0
1
3
C
0
(n
12
,.
p53
B
6
Post-PH
18
A
(h)
24
30
42
48
Cell
60
Gi
S
Cycle
S2
Growth
& Diffe’rentiation
1467
Phase
S6
M
M4
‘,
:LtIIJL
B
G)ce
LO
.0(l)C
C
0
0.
08
0.
.
o:j1
0.25
0
0.5
1
3
Time
6
12
18
Post-PH
24
30
42
48
60
0
(h)
Fig. I.
))5 1 protein
(‘xpression
in regenerating
rat liver. Rats were subjected
to 70’
PH
sat ritk ed at the indicated
times
post-PH
through
60 h.
Nu lear Prot(’irl
e’xtracts
ss-ere’ isolated
and resolve(I
by SDS-PAGE
as descrilwd
in ‘Pslaterials
and Methods.”
A, immunoblot
of p53 expression
det’rmined
using
mAh Pab 240. The blot was prolwd
and processed
as
(Ies( ribed
in Fig. I . B. the relative
amounts
of p5
were
quantitated
by
densitonletrk
analysis
and plotted
on an arbitrary
scale using 0-h expression
as unity.
TIic imn#{236}unoblot is repre’sentative
ot tour experiments,
using three
different
niAbs
to p5 1.
.111(1
than G and reached
a nadir
6 h after S-phase
(Fig. 58). Interestingly, there was a reproducible
increase
in M phase
that dissipated
after release.
Phosphorylation analysis of the Rb protein in nuclei more closely
resembled
that observed in total cell lysate in that maximal
expression
of the hyperphosphorylated
species
occurred
2
h after S-phase release relative to G1 and returned to lower
levels by 4 h after M-phase
release (Fig. SC). In contrast
to
total
cell lysate,
molecular
hands
below
pRb were
not
detected in protein extracts from the isolated nuclei.
As with the regenerating
liver, we confirmed
the results
from
the immunoblot
analyses
in the synchronized
HuH-7
cells with
indirect
immunofluorescence
of Rh protein
expression.
Again,
XZ161
provided
the strongest
signal
and
least amount
of nonspecific
staining.
The results confirmed
those obtained by Western blot analysis (Fig. 6). In G1, the
majority
of immunofluorescence
staining
was detected
in
the cytosol
of the nonconfluent
HuH-7
cells. S phase was
associated
with a significant
increase
in nuclear
signaling
with some cytosolic
staining
still detectable.
The most dramatic
change
occurred
at 6 h after S-phase
release
when
both nuclear
and cytosolic
abundance
of Rb staining
was
significantly
decreased
to approximately
1 5% of
levels.
At M phase and 4 h after M-phase
release,
there was reappearance of significant staining in both nuclei and cytosol.
In fact, approximately
20% of the cells had significantly
elevated
levels of nuclear
staining,
although
the intensity
was less than that observed
in S phase. A number
of cells
showed
unique
distributions
of Rb protein,
suggesting
dynamic
changes
in suhcellular
localization
during
and after
release
from
M phase.
mAh F20.2F to f3,-microglohulin
was
used as control
and showed
no evidence
of nuclear
staining
(data not shown).
Gi
S
S2h
S6h
M
Cell Cycle
Phase
Cell Cycle
Phase
M4h
C
a)
U)
C
S phase
release
Fig. 4.
Total RI) protein
expression
in synchronized
HuH-7
human
liepatonia cells. Subconfluent
cells were synchronized
in G,, S. and M phases
with hydroxyurea,
aphidicolin,
and noco(IaZOle,
respectively,
as described
in “Materials
and Methods.”
Cells treated
with aphidkolin
were harvested
in
S phase (5) or released
harvested
at 2 h (S2(,
in media containing
and 6 h (S6(. Cells
10/ FBS and no aphidicolin
incubated
with
nocodazole
and
and
Colcemid
were harvested
in M phase (M( or released
in media
containing
1 0’%, FBS and harvested
at 4 h (M4(.
Inimunoprecipitation
and Western
blotting
were
arried out as described
in “Materials
and Methods
.“ Cell
cycle
status was determined
using
BrdUrcl
incorporation
at the’ indicated
time
points as reomnie’nded
I)y the Boehringer
Mannhe’ini
( elI
proliferation
kit.
A. immunoblot
of pRI) trom synchronized
HuH-7
( ells
after inimunopre
ipitation
of total
cell lysate.
B, changes
in
total
RI) protein
expression
in
synchronized
HuH-7
cells relative
to G , . Rb protein
levels
were densitometrically
quantitated
from Western
blots in whkh
equal amounts
of protein
were added
to each lane. (, relative
changes
in Rb phosphorylation
during
different
phases of Iii’ cell cycle
determined
froni immunOl)lot
and clensitometric
analyses.
The sum of the phosphorylated
state’s equals
the total
amount
of Rb expressed
at each cell cycle phase. The’ results are representative
of tour different
experiments.
pRh, hypophosphorylated
RI); ppRb.
hyperphosphorylated
RI).
Expression of Retinoblastoma-related
erating Liver and Primary Hepatocytes.
Proteins
in Regen-
In characterizing
Rb expression
in regenerating
rat liver with the different
mAbs,
we detected
a previously
reported
300-kDa
Rbrelated
protein
as well
as two
novel,
immunologically
cross-reactive
polypeptides
(Fig. 7A). They exhibited
apparent molecular
masses of 1 56 and 74 kDa by SDS-gel
elec-
1468
Rb Protein
Expression
in Regenerating
A
Cell
Gi
S
Rat Liver
Phase
Cycle
S2
S6
M
M4
B
C
.
0
2
0.0
1
0
31
S
52h
S6h
M
Cell Cycle
Phase
Cell Cycle
Phase
M4h
C
a)
C/)
C
21
Fig. 5.
Nuclear
Rb protein
expression
in synchronized
HuH-7
cells. HuH-7
cells were synchronized
in the different
phases of the cell cycle as described
in Fig. 4. Nuclei
and nuclear
protein
extracts
were isolated
and immunoprecipitated
for immunoblot
analysis
as described
in “Materials
and Methods .“ A, immunoblot
of Rb from synchronized
HuH-7
cells after immunoprecipitation
ot nuclear
extract.
B. changes
in nuclear
RI) protein
expression
in synchronized
HuH-7
cells relative
to G,. Rh protein
levels were densitometrically
quantitated
from irnmunoblots
containing
equal amounts
of protein/lane.
C, relative
changes
in Rb phosphorylation
during
the different
analyses.
protein
four
of the cell
The’ sum
cycle
deterniined
the phosphorylated
d)f
expressed
at each
different
experiments.
phorylated
from
immunoblot
states equals
cell cycle phase. The
pRh, hypophosphorylated
results
and
the total
densitometric
amount
of Rb
are representative
of
RI); ppRh,
hyperphos-
Rb.
trophoresis.
Nuclei
were isolated
from various
time points
through
72 h of regeneration
and were processed
for immunoblot
analysis
using different
mAbs to Rb. Each of the
proteins
showed
significant
increases
in nuclear
abundance
after the livers underwent
70% PH. p300 showed
increases
at 0.5 and 3 h, returned to baseline levels at 6 h post-PH,
and then increased
3.5-fold
over 0-h levels at 1 8 h post-PH
and remained
elevated through 72 h of growth (Fig. 78).
p156
levels fluctuated during the first 3 h, increased approximately
6-fold
at 6 h post-PH,
and remained
approximately
3-fold
increased
until
60 h post-PH,
when
they
returned to near baseline levels. The p7’4 Rb-related protein
showed
the same dramatic
modulation
exhibited
by p300
and p1 56 during
the regenerative
period.
It decreased
im-
mediately
after PH until
6 h, when
it increased
6.5-fold
above
0-h levels.
Its abundance
was 2- to 3-fold
above
baseline
until 60 h post-PH,
when
it peaked
7-fold
above
baseline
levels and then returned
to near 0-h levels at 72 h
post-PH.
To further
characterize
the antigenic
similarities
between
Rb and its related
proteins,
a series of mAbs raised against
different
epitopes
of native
Rb protein
were used to map
p300,
p156,
and
p74.
Monoclonal
antibodies
XZ161,
xz1 21 , and XZ77, which
were used in the immunoprecipitation and immunoblot
analyses
ofthe
1 1 0-kDa
Rb protein,
recognize
epitopes
393-621
and 715-802,
444-621,
and
444-535
and 620-665,
respectively,
in the protein
(38).
p300, p1 56, and p7’4 were detected
by XZ1 61 in rat liver,
but p74 displayed
very weak antigenicity.
XZ1 21 exhibited
very strong affinity
for p1 56, as did XZ77 for p74, but only
weak affinity
for the p1 1 0 Rb protein.
Taken together,
p1 56
expressed
the putative
Rb protein
epitopes
corresponding
to
aminoacids
536-621
and possibly
715-802
and 393-443
in the Rb protein;
p74 appeared
to share an epitope
at
620-665,
and p300 expressed
epitopes
corresponding
to
amino
acids
715-802
and possibly
393-443
in the Rb
protein.
Based on the results of Rb and Rb-related
protein
levels in
the regenerating
liver, we investigated
their expression
in
isolated hepatocytes that were maintained
in culture for 96
h. We were particularly
interested
in comparing
Rb expression to that of the novel related proteins
p1 56 and p74 (Fig.
8). Under
culture
conditions
in which
the cells were incubated in 1 0% fetal bovine
serum and allowed
to replicate,
both the pl 10 Rb protein and itsrelated proteins p1 56 and
p74
exhibited
very similar
patterns
of expression
in cultured
primary
hepatocytes.
Although
the total abundance
of protein did not change
dramatically
during
96 h, there was a
reproducible
redistibution
of protein
from the cytosol
into
the nucleus.
Nuclear
expression
reached
almost
maximum
levels
by 24, 48, and 12 h for p156,
p110,
and p7’4,
respectively.
In the case of the p1 1 0 Rb protein,
the major
species
was, in fact, in the hyperphosphorylated
state. In
contrast,
hepatocytes
cultured
in serum-free
media
expressed
primarily
the hypophosphorylated
species
in their
nuclei,
which
decreased
to minimally
detectable
levels by
42 h in culture (Fig. 9). Interestingly, from 3 h on, the lower
molecular
weight
species
below
pRb were again
detectable, as noted previously
in the synchronized
HuH-7
cells
(Fig. 4A).
Half-Life
in Cultured
Determination
Hepatocytes
of the Retinoblastoma
Protein
and Regenerating
Liver. It was
apparent
that expression
of the Rb protein
fluctuated
significantly
over relatively
short periods
during
liver regeneration.
It has been reported
previously
that in the regenerating rat liven, the half-life
of both the canonical
4.7-kb
transcript
as well as the 2.8-kb
transcript
exhibited
mRNA
half-lives
of approximately
40 mm, both at 0 time and 6 h
post-PH
(36). In addition,
transcriptional
activity
of the Rb
mRNA
increased
approximately
6-fold
within
the first 30 to
60 mm after PH and then returned
to near baseline
levels.
The half-life
of the Rb protein
was obviously
an important
factor
involved
in its modulation
in the regenerating
rat
liver.
To determine
the half-life
of the protein
in vivo,
cycloheximide
was used to block protein
synthesis
between
3 and 6 h post-PH.
This time point was chosen
because
the
abundance
of Rb protein
in 0-h liver was not sufficient
to
perform
adequate
Western
blot and densitometric
analysis.
Based on the dramatic
changes
in Rb protein
abundance
C(’ll
A
Time
0.5
0
A.
1
3
Post-PH
6
18
(;r’th
5I tjiffer(ntiation
1469
(h)
24
30
42
48
60
72
p300
p1 56
-
-
-
-
p74
B
7
#{149}p300
0p156
Dp74
6
w
#{182})14k
B
,
-.-.
‘
*‘Pkt
.
:
a)
0)5
.
C
:
.C4
0
D
:
I
I
#{149}*:*
‘4fr
?::;:.
;
,-
*4
-
.
0
.
0.51
3
Time
.-
‘
C
:1
.
HinLrn?dhh
t
,.
I
3
0
-,
I
6
1824304248
Post-PH
6072
(h)
Fig. 7.
Expression
of Rb-related
proteins
in regenerating
rat liver. Rats were
subjected
to 70/
PH, and livers
were
harvested
at the incli ated times
post-PH.
Nuclei
and nuclear
protein
extracts
were isolated
and resolved
by
SDS-PAGE
as described
in “Materials
and Methods
.“
After transfer,
the blots
were probed
with mAbs recognizing
different
epitopes
of human
RI) protein.
A, immunoblots
of Rh-related
proteins
expressed
through
72 h post-PH
and
visualized
by the ECL method.
1)300 was hybridized
with XZ1 61 , p1 56 was
hybridized
with XZ121,
and p74 was hybridized
with XL77. B. (hanges
in
0300,
01 56, and p74 expression
through
72 h P05t4’H
relative
to
controls.
The Rb-related
proteins
were densitometrically
quantitated
from
Western
I)lOts containing
equal
quantities
d)f
protein/lane.
The results
representative
of at least three different
experiments.
D
-
during
the regenerative
period,
it was not surprising
to
determine
that the half-life
of the Rb protein
in regenerating
liver was 1 .9 h (data not shown).
This was similar
to the
half-life
of the protein
in cultured
primary
hepatocytes,
which
was determined
to be 2.2 h using an S-labeling
technique and no cycloheximide
(Fig. 1OA). As a comparison, the protein half-lifeofthe p1 56 Rb-related protein was
4.25
h (Fig. 108).
.
.‘
, ,.
i_’
*
-
.
!i’:*
,.s’.
,
:
?‘:
-
t
E
.
.,-
,‘,.
;
...-
,‘
‘
I
‘i
‘-
Effect of TGF-1
on Retinoblastoma,
E Protein Expression in Regenerating
CDK4, and Cyclin
Liver. It is well es-
tablished
that TGFinhibits
the growth
of certain
cell
types,
including
hepatocytes,
by modulating
progression
through
the late G, phase
of the cell
cycle
(39).
In
addition,
it has also been shown
that i.v.-administered
TGF-f31
inhibited
DNA synthesis
in the regenerating
liver
after PH (40). A candidate
target for the growth-inhibitory
5...
0’:
0-h
the
are’
,
S
‘p
Fig. 6.
Immunofluorescence
localization
of Rb antigen
in sync Iironiied
HuH-7
cells. Cells were synchronized
into the different
phases
of the cell
cycle as described
in Fig. 4 and “Materials
and Methods,”
fixed in paratormaldehyde,
and subjected
to indirect
immunofluorescence
with affinity-punfied mAb XZ1 61 . RI) is distributed
primarily
in the cytosol
(luring
G , (A) and
located
almost
entirely
in the nucleus
during
S phase (B). C, 6 h after S-phase
release,
Rb is almost
undetectable
in the nucleus
and only slightly
(Iete’( table
in the cytosol.
In M phase )D) and 4 h (F) after M-phase
release,
RI) antigen
is distributed
in both the’ nuclear
and cytosolic
compartments,
with parti
ularly
strong staining
in 5OfllC
nuc lei. The immunofluores
e’nce distribution
of Rb antigen
during
the difterent
cell cycle phases agrees with the distnil)ution of Rb protein
dlet(’rmine(I
by immunol)lot
analysis
0)
HuH-7
cells. B,ir.
pm.
25
1470
RI) Protein
Expression
Regenerating
in
Time
0
3
6
12
Rat Liver
in Culture
18
24
36
significantly
reduced
(P< 0.001 ; Fig. 1 1). In addition,
hypophosphorylated
species
as a percentage
of total
protein
decreased
more rapidly
than the hyperphospho-
(h)
48
72 84
96
___________
60
uclei
rylated
animals
p156
--
-
whole
.-.
cells
-wwwww
______________
p110
#{149},a. #{149}
0 0
nuclei
whole
cells
______
nuclei
-
w
p74
i#{149}i.i#{149}iii
whole
cells
Fig. 8.
Expression
of RI) arid Rb-related
proteins
in cultured
primary
hepatocytes.
Hepatocytes
were isolated
and maintained
in culture
with 1 0% FCS
through
96 h. At the indicated
times, cells were harvested
and processed
for
total cell lysate and nuclear
proteins
as described
in “Materials
and MethO(IS.”
Equal amounts
of protein
from each time point were loaded
onto gels
and processed
for immunOl)lot
analysis
and detection
of p156,
pRb(110(,
and p74 using mAbs XZ121,
XZ161,
and xZ77, respectively.
Nuclear
and
whole
lysate
proteins
were
subjected
to ECL Western
blot analysis.
The
results are representative
of three different
experiments.
A
Time
0
0.5
1
3
6
in Culture
12
18
24
6 h in the treated
expression
at 6 h
post-PH
was
not significantly
different,
a faint
above
ppRb and perhaps
representing
an additional
perphosphonylated
species
consistently
disappeared
band
hyin
the
b b
0
Rb protein between
1 and
(P < 0.05).
Although
Rb
the
Rb
TGF-j31-treated
group
(Fig. 1 1A).
It has been reported
that during
the cell cycle,
the Rb
protein
is phosphorylated
under the control
of certain
cyclin-CDK
complexes,
including
cyclin
D-CDK4
and cyclin
E-CDK2
(41 ). Based
on these putative
interactions,
we examined
the effect of TGF-31
on expression
of CDK4
and
cyclin
F in regenerating
rat liver.
TGF-f31
administration
dramatically
inhibited
the expression
of CDK4 at 1 and 6 h
post-PH
(P < 0.001 ) and reproducibly
increased
levels of
the protein
absence
of
creased
in
between
1
at 24 h post-PH
(Fig. 1 2A). Interestingly,
in the
the cytokine,
the CDK4
protein
steadily
deabundance
from
1 to 24 h post-PH
(P < 0.01
and 6 h; P < 0.001
between
6 and 24 h).
However,
the levels of CDK4
associated
with TGF-1
inhibition
at 1 , 6, and 24 h post-PH
were almost
identical.
Cyclin
E protein
expression
in control
regenerating
liver
increased
approximately
3-fold
between
1 and 6 h (P <
(h)
30
42
48
ia#{225}f#{149}
60
72
A
-pRb
100
B
C
0
Cl)
C
0
.o
15
cog
-
Ui-
0.0
.
Ui0
-cD
.._4
.
.
.
.
Time
Time
in Culture
hyperphosphorylated
(h)
100
C
0
Cl)
Cl)
each time point were loaded
on gels and processed
for inimunoblot
analysis
using XZ161
mAb.
The immunoblot
reactions
were visualized
by the ECL
method.
B, changes
in total Rh protein
expression
through
72 Ii in culture
relative
to 0-h controls.
RI) protein
levels were densitometrically
quantitated
from imniunoblots
in which
equal amounts
of protein
were added
to each
lane.
The results
are representative
of three
different
experiments.
pRb,
Rb; ppRb,
Post-Chase
B
(h)
Fig. ‘).
Rb protein
expression
in nonreplicating
cultured
hepatocytes.
A,
hepatocytes
were isolated
and cultured
in media
without
serum through
72
h as described
in “Materials
and Methods.”
Equal amounts
of protein
from
hypophosphorylated
.
0.
)(
10
LO
Rb.
effect
of TGF-1
in replicating
hepatocytes
was the Rb
protein.
To determine
whether
administration
of the cytokine
was associated
with changes
in abundance
and/or
phosphorylation
of Rb expression
in the regenerating
liver,
TGF-j31
was iv. administered
45 mm prior to sungery and 1 2 h after PH at a dose sufficient
to inhibit
DNA
synthesis.
Although
it has been
shown
previously
that
TGF-pl
administration
had little effect
on transcript
expression,
Rb protein
expression
at 24 h post-PH
was
o
(0
Time
1
0
-
0.
Post-Chase
2
4
6
-
2
Fig.
10.
Rb protein
and
p156
4
(h)
8
10
-
6
Time
Post-Chase
half-life
determinations
8
10
(h)
in cultured
primary
hepatocytes.
Isolated
primary
hepatocytes
were
pulse-chase
labeled
with
ltrans-35Slmethionine.
At the indicated
times,
the cells
were
harvested,
lysed, and incubated
with XZ161
for pRb and XZ121
for p156.
The immunoprecipitated
Rb protein
(A) and p1 56 (B) were analyzed
by SDS-PAGE
and
subjected
to densitometnic
analysis
as described
in “Materials
and Methods.”
The apparent
half-lives
were determined
by linear
regression
analysis.
Cell
A
Time Post-PH
1
24
:..
I
B
staining
tocyte
C
0Cl)’-
.0
1-
+11-
l
1
24
6
Time
Post-PH
& Differentiation
1471
and corresponds
to the peak of delayed-early
gene expression (34). After a dramatic
decrease
just prior to peak S
phase, Rb reaches its highest levels at 24 to 30 h post-PH,
corresponding
to peak DNA synthesis and initiation
of the
first wave of mitosis.
A similar but less abundant
cycle of Rb
expression
also occurs during
the second mound of cell
proliferation
in the regenerating
liven. Very similar findings
were determined
by immunocytochemistry
in which
Rb
protein expression
was identified
by immunofluorescence
(h)
6
Growth
(h)
Fig. 1 1. Effects oITGF-f31
on Rb protein
expression
in regenerating
rat liver.
Animals
were injected
iv. with vehicle
or 1 0 pg ofTGF-f31
45 mm before PH
and 1 2 h post-PH.
A, immunoblot
analysis
of Rb protein
expression
in nuclei
isolated
from 1 -, 6-, and 24-h regenerating
liver treated
with vehicle
(-) or
TGF-f31
(+) as described
in “Materials
and Methods.”
B, densitometric
quantitation
of changes
in total Rb protein
expression
and phosphorylation
status
from
1 -, 6-, and 24-h
post-PH
regenerating
livers
relative
to 0-h
controls.
Equal amounts
of protein
were added
to each lane of the immunoblots.
The results are representative
of three different
experiments.
pRb,
hypophosphorylated
Rb; ppRb,
hyperphosphorylated
Rb.
of the regenerating
nuclei
that stained
liven.
The
percentage
of hepa-
positively
for Rb at 24 to 30 h
post-PH is almost identical
to the percentage
of cells undengoing DNA synthesis and mitosis (32) and supports the
observation
by Western
blot analysis that Rb protein expression and cellular
localization
are cell cycle regulated.
The results are consistent with the notion that the Rb gene
plays a key role in the regulation
of the cell cycle (28).
However,
the data also suggest a significant
uncoupling
of
transcript
and protein expression
for the tumor suppressor
gene. It has been shown previously
that major changes in
Rb transcript
expression
occur only through the first 1 2 h in
the regenerating
liven (36). Although
the total abundance
of
mRNA does not change during that period, theme is a significant
shift in expression
between
the 4.7- and 2.8-kb
species. No additional
changes in transcript
expression
or
transcriptional
mate occur at times when peak protein expression is observed.
Interestingly,
the p53 tumor suppresson gene exhibits
similar
patterns of both transcript
and
protein expression
following
PH as does Rb. Peak transcript
expression
occurs
during
G1 at 6 h post-PH
(data not
shown),
and
similar
to Rb, the
major
peak
in protein
ex-
pression
and then decreased 60% (P< 0.001)
at 24 h post-PH
(Fig. 1 28). TGF-1
administration
was associated
with significant
decreases,
as great as 90%, in protein
expression
at
each time point (P< 0.001).
0.01)
Discussion
The liven is unique in its ability to regenerate.
It represents
a remarkable
in vivo model for the study of gene expression
and growth regulation.
Within minutes after PH, numerous
cellular
changes take place that prime hepatocytes
to replicate. The process involves
a complex
pattern of gene
expression
and a modulation
of numerous
transcripts
duning
the
growth
period
(33,
34).
Although
the
immediate-
and delayed-early
genes are probably
responsible
for
hepatocytes
to transition
from G0 to G1, many other cell
cycle- and growth-regulated
genes are responsible
for the
progression
examining
through
expression
mitosis.
This is the first detailed
of the Rb tumor
suppressor
product
in regenerating
hepatocytes.
In addition,
hepatoma
cell
line,
Rb protein
cell cycle dependent
Rb
expression
Hepatocytes
liven as well as primary
using a well-differentiated
abundance
and exhibited
similar
in an essentially
maintain
capacity
a remarkable
presented
in this study indicate
liver expresses low levels of Rb
increase in Rb protein expression
erating state after 70% PH. The
1 2 h post-PH
occurs within the
gene
cultured
human
was shown
to be
characteristics
in the in vivo liven regeneration
of the adult
liven are long-lived
ated cells that remain
report
quiescent
to
model.
diffenenti-
state but
to proliferate (33). The data
that the normal adult mat
protein and that a marked
occurs during the megenfirst significant
increase at
G1 phase of the cell cycle
is at 30 h post-PH. This parallel pattern of expression for two very different
tumor suppressor
genes appears
to be more than coincidental
and may reflect their involvement in the first rnitotic wave after PH as a checkpoint
for
further proliferation.
In this regard, a recent report demonstrates a direct link between the two proteins in controlling
cell growth and apoptosis
(42).
In support of the apparent
uncoupling
between
protein
and mRNA levels, cytosolic
levels of p53 and Rb protein in
the regenerating
liver, although
less abundant,
show a similan pattern of modulation
to that exhibited
in nuclei (data
not shown).
Theme is a growing
list of genes, in fact, in
which steady-state
transcript
expression
is uncoupled
from
protein expression
(43). In this regard, it has been shown
that selective
translational
control
of mibosomal
protein
mRNAs constitute
an important
regulatory
mechanism
opemating in vivo in the course of liven regeneration
(44).
Theme has been some controversy
as to whether
abundance of the Rb protein is modulated
during the cell cycle.
It has been reported
that no significant
differences
were
detectable
in the staining pattern on distribution
of the Rb
protein in the G1, S, and G2 phases of the cell cycle in a
collection
of Rb-expressing
cell lines (45). In contrast, the
apparent lack of nuclear staining in a group of Rb-positive
tumor cells resulted
from a significant
decrease
in total
cellular Rb protein during G0 on middle G1 (46). Progression
of embryonic
stern cells towards
the G1-S transition
was
similarly
accompanied
by a marked
decrease
in total
abun-
dance of Rb protein (47). In addition,
differentiation
of the
embryonic
stem cells was associated
with a marked
increase in total amounts of Rb protein as observed previously
when embryonal
carcinoma
cells were induced to differentiate into neumoectodemmal
cells (48). The results of the
present study indicate
that in regenerating
mat liver and
1472
Rb Protein
Expression
in Regenerating
Rat Liver
B
A
Time
Post-PH
1
-
Time
(h)
6
+
-
I
24
+
-
Post-PH
(h)
6
Fig. 12.
Effects
of TGF-(31
on CDK4 and cyclin
E protein
expression
in
regenerating
liver.
Immunoblot
analysis
and
densitometric
quantitation ofCDK4
(A) and cyclin
E
(B) expression
in regenerating liver treated
with TGF-j31
were performed
as described
in Fig. 1 1 and “Materials
and
Methods.”
Densitometric
quantitation
of changes
in
24
+
Cyclin E
CDK4
C
C
0
(I)
0
Co.
nuclear
protein
expression
from 1 -, 6-, and 24-h post-PH
regenerating
livers
was expressed
relative
to 0-h controls. Equal quantities
of protein were loaded onto each of
the lanes. Representative
immunoblots
from three
different experiments
are shown.
0.
0
0
>
0
Time Post-PH
Time Post-PH
(h)
HuH-7
human
hepatoma
cells, the total amount
of Rb
protein changes dramatically
during the various phases of
the cell cycle. This may be facilitated,
in part, by the nelatively short half-life ofthe Rb protein in both systems. In the
hepatoma
cell line, maximal
decrease in total cellular
Rb
protein occurred subsequent
to S-phase release and prior to
M phase. In contrast,
the quiescent
liven representing
a
unique
in vivo example
of the G0 state of the cell cycle
exhibited
almost
undetectable
levels of Rb protein.
It is well documented
that the state of phosphonylation
of
the Rb protein fluctuates
during the various phases of the
cell cycle (7, 49). Generally,
in G0 and early G1, Rb protein
exists
primarily
in the
hypophosphorylated
form.
As the
cells transit into mid/late
G), the protein undergoes
additional phosphorylation
and then remains in this hypemphosphorylated
form throughout
S phase, G2, and most of M
phase (50). In short, underphosphorylated
Rb protein appears to block passage through the G1-S boundary
of the
cell cycle; hypemphosphorylation
relieves
the block and
allows cell replication
to occur. It was, therefore,
somewhat
surprising
that the Rb protein
remained
primarily
in the
hypemphosphonylated
state in regenerating
liven and cultuned primary hepatocytes.
In contrast, primary hepatocytes
cultured
in serum-free
media expressed
primarily
the hypophosphonylated
form of the Rb protein. It was also interesting that a significant
portion of the Rb protein during the
G1 block in HuH-7
cells was hypenphosphonylated.
Howeven, it has been reported
previously
that cells growtharrested in G1 with hydroxyunea
exhibited
hypemphosphomylation
of Rb protein
(51 ). Our
results support the recent
observation
that substantial
phosphorylation
of Rb exists in
G1 even prior to the hypemphosphorylation
point, suggesting the existence
of distinct patterns of phosphorylation
that
are associated
with different
subsets of Rb protein molecules (52). In these hepatocyte
models of cell growth, the
phosphonylation
of Rb is not coordinated
with the G1-S
transition
and may not directly regulate it. The fact that most
of the Rb protein was phosphorylated
in regenerating
liver
and highly proliferating
HuH-7 cells implies that in these
replicating
models, the hyperphosphonylated
Rb has lost its
ability to interact with a variety
of nuclear
proteins.
How-
(h)
even, although
the loss of binding
to proteins
such as transcmiption
factor E2F is assumed
to reflect a functional
mactivation
of Rb it may, in fact, permit
additional
functions
of
the Rb protein.
For example,
a recent
report demonstrated
that the COOH-temminal
domain
of Rb, outside
of the NB
pocket,
complexes
and regulates
the activity
ofc-abl,
which
has been shown
to phosphorylate
the catalytic
subunit
of
RNA
polymemase
II (53). Furthermore,
the c-AbI-Rb
cornplex
is disrupted
by phosphorylation
of the Rb protein
during the cell cycle.
Theme is a growing family of proteins that shame structural
similarity
with the Rb protein.
Two of those proteins,
p107
(54)
and p1 30 (55), were isolated
by their interaction
with
the region
of adenovinus
E1A that binds
Rb. In addition,
p300 also shows immunological
cross-reactivity
to the vanious subsets of Rb protein
(38), although
its binding
site to
adenovirus
El A is distinct
from that of Rb, p1 07, and p130
(56). In the present study, two additional
and novel immunologically
cross-reactive
proteins,
p1 56 and p74,
were
identified
in regenerating
liven by a series of mAbs against
human
Rb protein
(38). In subsequent
experiments,
both
proteins
could
be immunoprecipitated
from three human
hepatoma
cell lines (HepG2,
HuH-7,
and Hep3B),
Ads
transformed
primary
human
embryonal
kidney
cells 293,
and human
osteogenic
sarcoma
cells (Saos-2)
but not from
African
green monkey
kidney
cells (CV-1 ), or an immortal-
ized mouse
hepatocyte
cell line (AML-l
Both proteins
exhibited
significant
pression
during
liven regeneration
primary
hepatocytes.
p74 exhibited
bution
as p1 56 except
2; data not shown).
induction
as well
a similar
that it was additionally
in nuclear
exas in cultured
cellular
distni-
detected
in
cells (data not presented).
However,
tryptic
digests
of the isolated proteins indicated that they are different and
that p1 56 is not simply
a dimem of p74 (data not shown).
The
precise moles of p156 and p74 in cell growth remain to be
determined.
However,
it is interesting
that p1 56 and p74
are present
in the Rb protein-deficient
Saos-2 and Hep3B
cells. The results suggest that both Rb-related
proteins
may
substitute
certain
functions
of Rb protein
in the control
of
cell proliferation
and differentiation.
COS-7
Cell
The mechanism
by which
TGF-l
inhibits
cell prolifenation is poorly
understood
and probably
involves
the interplay of a number
of gene products.
For example,
it has been
reported
that TGF-l
suppresses
c-myc
gene transcription
by modulating
the binding of cellular factors, including Rb,
to the 5’ regulatory region of the gene (57, 58). More
recently,
the mechanism
of TGF-l
inhibition
has been
related
to its ability
to prevent
hyperphosphorylation
of Rb
protein
through
its effects on expression
of Gl cyclmns and
their associated
cyclmn-dependent
kinases
(59). It has been
shown
previously
that TGF-31
induces
transient
inhibition
of liver regeneration
in rats and mice (40, 60) in the absence
of changes
in Rb transcript
expression
(36). However,
the
present
study indicates
that TGF-l
not only
inhibits
Rb
protein
phosphonylation
in cultured
primary
hepatocytes
(data not shown)
but also inhibits
Rb protein
expression
in
the regenerating
liver. The results also indicate
that TGF-l
induces
significant
decreases
in both CDK4 and cyclin
E as
early as 1 h post-PH.
In this regard,
it has been reported
recently
in Mvl Lu mink
lung epithelial
cells that TGF-3l
induced
suppression
of CDK4 synthesis
during
G1 (61 ) and
inhibited
cyclin
E-associated
kinase activity
(62). Moreover,
in Mvl Lu mink
lung epithelial
cells, TGF-f31
functions
in
another
manner
by raising
the threshold
level of cyclin
E
necessary
to activate
CDK2 through
an inhibitor
that binds
cyclin
E-CDK2
complexes
(63). Inhibition
of CDK4 synthesis by TGF-f31
is linked
to G1 arrest and probably
involves
a collaboration
of Gi cyclins
in the functional
inactivation
of the Rb protein
(64, 65). These data suggest that TGF-f31
inhibits
Rb protein
phosphorylation,
at least seemingly
by
suppressing
expression
of CDK4 and cyclin
E, resulting
in a
decrease
of cyclin
D-CDK4
and cyclin
E-CDK2
complexes.
Furthermore,
it was reported
recently
that the cell cycledependent
expression
of cyclin
Dl
is dependent
on the
presence
of functional
Rb protein
(66). Our results suggest
that the effect of TGF-l
on the phosphorylation
status of
Rb in the regenerating
liven may involve
numerous
factors
as well as the total abundance
of the protein.
In conclusion,
the results of the present
study
indicate
that the regenerating
mat liven represents
a remarkably
unique
in vivo system for studying
cell cycle expression
of
the Rb tumor
suppressor
gene product.
It provides
an opportunity
for examining
the function
of the Rb protein
in
normal
cell growth
and differentiation
of cells in the whole
organism.
It is attractive
to consider,
for example,
that p1S6
and p74 are similar
enough
to Rb that the three proteins
shame similar
functions
in vivo. Future studies will undoubtedly provide
us with the necessary
information
to establish
their own role as potential
tumor
suppressors.
Factors controlling
the regeneration
of an entire
organ
are obviously
complex.
For example,
the pattern
of Rb protein
levels
during
the first round of cell replication
indicates
a significant uncoupling
of transcript
expression
and translation.
Interestingly,
a similar
uncoupling
of mRNA
and protein
expression
was also observed
for the p53 tumor
suppressor
gene (data not shown).
Future studies
will provide
impomtant information
regarding
the role of translational-dependent expression
of these tumor
suppressor
genes and their
role in modulating
hepatocyte
growth
and differentiation.
Materials
Materials.
and Methods
mAbs XZ1 61 , XZ1 21 , and XZ77 to human
Rb
protein
and Pab 242 and 421 to human
p53 were genenously
provided
by Dr. Ed Hanlow
(MGH
Cancer
Center,
Growth
& Differentiation
Chanlestown,
MA). mAb F20.2F
to j32-microglobulin
was
kindly
provided
by Dr. Ronald
P. Messnen
(University
of
Minnesota
Medical
School,
Minneapolis,
MN).
Goat antimouse
and anti-rabbit
IgG horseradish
penoxidase
were
purchased
from Bio-Rad
Laboratories
(Hercules,
CA). Nonmal goat serum
and goat anti-mouse
IgG Cy3 conjugates
were purchased
from Jackson
ImmunoReseanch
Labonatonies, Inc. (West Grove,
PA). Protein
A-Sephanose
6 MB was
obtained
from
Pharmacia
Biotech,
Inc. (Piscataway,
NJ).
mAbs against cyclin
E (HE-i 2) and p53 (Pab 240) and rabbit
polyclonal
antibody
to CDK4
were purchased
from Santa
Cmuz Biotechnology,
Inc. (Santa Cnuz, CA). Cell cycle synchronization
reagents
and Hoechst
dye were
purchased
from Sigma Chemical
Company
(St. Louis, MO). All other
standard
reagents
were purchased
from Aldrich
Chemical
Co. (Milwaukee,
WI),
Curtin
Matheson
Scientific
(Eden
Praine, MN), on Fisher Scientific
(Itasca,
IL).
Animals
and
Surgical
Procedures.
In
brief,
male
Sprague-Dawley
mats (250 to 275 g) were purchased
from
Harlan
Sprague-Dawley,
Inc. (Indianapolis,
IN), maintained
on a standard
12-h light/dark
cycle,
and fed commercial
laboratory
chow
ad libitum.
They were subjected
to midventral
lapanotomy
and 70% PH under
ether anesthesia
between
9 a.m. and 1 1 a.m., as described
previously
(31).
At various
times after PH, the animals
were sacrificed,
and
the remnant
livers
were
removed
and rinsed
in normal
saline
solution;
then a 0.5-cm
cube from the night lower
lobe was excised
and embedded
in OCT (Baxter
Scientific,
Minneapolis,
MN)
for immunohistology.
The remaining
liver was flash-frozen
in liquid
nitrogen.
Sham control
livers
were obtained
under similar
conditions
but without
PH. To
inhibit
protein
synthesis,
a dose of 5 mg/i 00 g body weight
of cycloheximide
(Sigma) was administered
i.p. 1 h before
surgery
(36). Ten pg of TGF-(31
(R & D Systems,
Inc.,
Minneapolis,
MN) were administered
as an i.v. bolus 45
mm prior to surgery
and 1 2 h after PH.
All animals
received
humane
came in compliance
with
the institute’s
guidelines
as outlined
in “Guide
for the Cane
and Use of Laboratory
Animals”
prepared
by the National
Academy
of Sciences and published by the NIH (NIH publication
number
86-23,
revised
1985).
Isolation
of Hepatocytes
and Cell Culture.
Rat hepato-
cytes were isolated
by standard
collagenase
perfusion
as
described
previously
(67).
The hepatocyte
suspensions
were obtained
by filtering
the digested
livers through
5evemal layers of gauze
to remove
undigested
material.
The
cells were then centrifuged
at 50 x g for 2 mm, and the
supennatant
was removed
by gentle
aspiration.
After filtration through
a single layer of gauze, the cells were washed
in modified
Eagle’s medium;
cell viability
was 85 to 90%, as
determined
by trypan
blue exclusion.
The freshly
isolated
hepatocytes
were plated on 1 00-mm
Pnimania tissue culture
dishes (Becton
Dickinson
Labware,
Lincoln
Park, NJ) at 7.5
x i0
cells/cm2
in Williams’
E medium
(GIBCO,
Grand
Island, NY) supplemented
with 26 mi sodium
bicarbonate,
23 mi HEPES, 0.01 uniVml
insulin,
2 mi i-glutammne,
10
nM dexamethasone,
5.5 mrvi glucose,
1 00 units/mI
penicillin, and 100 units/mI streptomycin.
Cultures were maintamed at 37#{176}C
in a humidified
atmosphere
of 5% CO2. After
2 h, the cultures
were washed
with fresh media to remove
dead cells and loosely
attached
aggregates.
The medium
was changed
every 24 h by gentle
pipetting.
To stimulate
growth,
1 0% heat-inactivated
FCS (Atlanta
Biologicals,
Inc.,
Noncross,
GA) was added to the medium.
Hepatocytes
were
harvested
at times
indicated
in the figure
legends,
flash-
1473
1474
Rb Protein
Expression
in Regenerating
Rat Liver
frozen
in liquid
nitrogen,
hepatocellular carcinoma
in DMEM
supplemented
icillin,
and 100 units/mI
and stoned at -70#{176}C.The human
cell line HuH-7
was maintained
with 1 0% FCS, 1 00 units/mI
penstreptomycin.
Cell Cycle Synchronization.
HuH-7
cells were synchronized
as described
previously
(47) with modifications.
In
brief,
subconfluent
cells were
starved
for 24 h in media
containing
0.i%
FBS.
The cells were
then washed
and
incubated
for 1 8 h in media containing
1 0% FBS and either
2.5
pg/mI aphidicolin
to establish
S phase on 0.1 mri hydnoxyunea
to synchronize
in G1 . Cells treated
with aphidicohn were washed
twice and incubated
in fresh media with
1 0%
FBS for 2 on 6 h. For M-phase
synchronization,
cells
were incubated
for 1 6 h in media containing
1 0% FBS and
80
np/mI nocodazole;
0.06 igJml
Colcemid
was added
to
the media
for an additional
2-h incubation.
Cells
were
washed
twice
in media
with
1 0% FBS and harvested
immediately
(M) or after 4 h of incubation
(M4). The cells were
pelleted, flash-frozen in liquid nitrogen, and stored for
Western
blot analysis
at -70#{176}Cfor no longer
than 3 days.
Cell cycle
phases were determined
by BndUnd
inconponation at various
times using a cell proliferation
kit and the
manufacturer’s
recommendations
(Boehninger
Mannheim
Corp.,
Indianapolis,
IN). The labeled
cells were washed
in
PBS and fixed in 75% ethanol
at -20#{176}Cfor 30 mm. The
dishes were washed
twice
in PBS containing
0.1% Tween
20 and incubated
in PBS containing
5% normal
goat serum
to reduce
nonspecific
binding.
The BndUnd-Iabeled
cells
were
stained
with
anti-BmdUnd
mAb
supplemented
with
DNase
and sequentially
incubated
with anti-mouse-immunoglobulmn-fluonescemn.
Total DNA
was stained with Hoechst dye 33258
(Sigma).
Immunolabeling
was analyzed
using a Zeiss standard
fluorescence
microscope
(Carl Zeiss,
Inc., Thonnwood,
NY). Photographs
were taken with Kodak
Ektan-l 000 film (Eastman
Kodak
Co., Rochester,
NY). At
least 300 cells/time point were counted
for evaluation of
BmdUnd incorporation.
Gel
Electrophoresis
and
Immunoblotting.
Rb protein
was
cells
immunoprecipitated
from total
lysate
by incubating
in 1 ml lysis buffer
containing
250 msi NaCI,
0.1%
NP4O,
50 mM HEPES (pH 7.0), 5 mM EDTA,
50 mr’i NaF, 0.1
mM sodium
orthovanadate,
50 pg phenylmethylsulfonyl
fluonide, 1 jg leupeptin,
1 pg aprotinin,
and 1 mi DII
for 30
mm on ice. Cell debris
was removed
by centnifugation
at
1 0,000
x g for 1 mm. The lysate was precleaned
with 40 p1
of normal
goat serum and 1 00 p1 of fixed,
killed
Staph ybcoccus
aureus
cells (Zymed
Laboratories,
Inc., South San
Francisco, CA) and then incubated with 1 00 p1 of cultured
hybmidoma
supennatant
overnight
at 4#{176}C.
The lysate was
then incubated
with Protein
A-Sepharose
for 2 h at room
temperature.
The Sephanose
beads were washed
three times
with lysis buffer and then incubated
in denaturing
buffer at
95#{176}C
for 3 mm for immunoblot
analysis.
Nuclei
were isolated from liver tissue and cultured
cells as described
previously
(68, 69). Nuclear
proteins
were isolated
and immediately
flash-frozen
as small aliquots
in liquid
nitrogen
or
electrophonesed
immediately
on 6% SDS-polyacrylamide
gels. After electrophonetic
transfer
to nitmocellulose
membranes,
the blots were incubated
with i5%
hydrogen
peroxide
for 1 5 mm. The blots
were
probed
with
specific
primary
mAb
after
residual
protein
binding
sites were
blocked
with 5% milk in Tnis-buffemed
saline
(TBS). After
several
washes
in TBS, the blots were incubated
with goat
anti-mouse
antibody
for 1 h, and protein
was visualized
by
ECL detection
(Amensham
Corp.,
Arlington
Heights,
cord i ng to the manufacturer’s
recommendations.
RNA
Isolation
and
Northern
Blot
Analysis.
IL) ac-
Total
RNA
was isolated
from liver tissue as described
previously
(70).
PoIy(A)-enmiched
RNA,
obtained
by oligo-dT
(New
England
Biolabs,
Beverly,
MA) chromatography
was electrophomesed
on 1 % agamose, 2.2 M formaldehyde,
1 x 4-mompholinpmopanesulfonic
acid (pH 7.0) denaturing
gels and
transferred
by passive
capillary
diffusion
to MagnaGmaph
nylon
membrane
(Micron
Separations,
Inc.,
Westbomo,
MA). The large molecular
weight
RNA ladder
from
BRL
(Bethesda
Research
Labs, Gaithersbung,
MD) was used for
nucleic
acid standards.
A 960-bp
BgblI-OxaNI
fragment
of
the mouse
Rb gene (71) and a 900-bp
Pstl fragment
of the
mat asialoglycoprotemn
receptor
gene (72) were labeled
with
[a-32P]dCTP
(3000 Ci/mmol;
Amersham
Corp.)
by random
priming
(73) using a commercial
kit (United
States Biochemical
Corp.,
Cleveland,
OH). Hybridizations
were penformed
for 1 8 to 24 h at 42#{176}C,
as described
previously
(36).
Membranes
were washed
twice
for 1 5 mm at room ternpematune in 1 X SSPE-0.l%
SDS and twice
at 42#{176}Cin 1 X
SSPE-0.5%
SDS. Autoradiography
was done
with
Kodak
XAR film (Eastman
Kodak
Co.) at -70#{176}Cusing an intensifying screen.
Densitometric
Analysis. Video densitometny
was penformed
using a Macintosh
II (Apple
Computer,
Cupertino,
CA) coupled
to a Data Translation
DT2255
video
digitizer
(Data Translation,
Marlboro,
MA) and a JVC GX-N8
video
camera
(JVC Corporation
of America,
Elmwood
Park, NJ),
as described
previously
(74). Quantitation
of autonadiograms and fluonograms
used the NIH Image
1 .4 Densitometric
Analysis
Program.
Statistical
analysis
was performed
using
InStat version
2.01 to calculate
ANOVA
and Bonfenmoni
multiple
compamisons test P values.
Immunocytochemistry.
from
control
and
OCT
(Baxter
Scientific).
Sample
regenerating
Cryostat
tissues
mat livers
sections
were
and
collected
embedded
5 pm-thick
in
were
cut, mounted
on slides, fixed for 5 mm in 4% buffered
(pH
6.9)
pamafonmaldehyde
containing
15%
(v/v)
saturated
picnic acid, and incubated
with 5% normal
goat serum
in
TBS for 2 h. Sections
were then incubated
with the Rb mAb
XZ161
or mAb
F20.2F
against
32-rnicroglobuImn
in 0.3%
Triton
X-l 00/PBS
overnight.
After three washes,
the slides
were incubated
with a secondary
antibody
(cyanine
3.18conjugated
goat anti-mouse
IgG; Jackson
ImmunoReseanch)
in 0.3% Triton
X-iOO/PBS
(1 :200) for 1 h and then countenstained
for cell nuclei
in a solution
of 1 pg/mI of Hoechst
dye 33258
for S mm. Sections
were examined
using
an
Olympus
BH2 microscope
equipped
for vertical
dankfield
illumination
using a mercury
lamp (Olympus,
Lake Success,
NY). Hoechst
33258
was visualized
using a Schott
UG1
exciter filter and a 420-nm
Iongpass
emission
filter; cyanine
3.18 was visualized
using a 541-551-nm
exciter
filter and
a 573-607-nm
emission
filter.
Colon
photographs
were
made using Kodak
Ektam-l000
films (Eastman
Kodak
Co.).
Synchronized
HuH-7
cells were fixed and processed
in a
similar fashion.
Protein Half-Life
Determinations.
The in vitro Rb protein half-life
was determined
in hepatocytes
that were isolated from control
livers and grown
on 3.5-cm
Pmimania
culture
dishes (Falcon
#3801)
as described
above.
After 24
h in culture,
the cells were washed
in serum-free
media and
then incubated
in 2 ml of methionine-free
DMEM
containing 0.2 mCi/mI
of trans-355
label (ICN Biomedicals,
Inc.,
Cell
Costa Mesa, CA) for 4 h at 37#{176}C.
The radioactive
medium
was removed,
and the cells were washed
several times with
Williams’
E medium
containing
10% FBS and then with
chase medium
containing
1 5 mg/mI
of methionine
and 10
mg/mI
of cysteine
and harvested
at the indicated
time
points. Proteins were normalized
to total cell protein content and analyzed
by immunoprecipitation
and electmophonesis.
Gels were dried and subjected
to automadiogmaphy
and quantitated
by densitometry
as described
previously
(36). The in vivo half-life
of the Rb protein
was assessed
in
the regenerating liven 3 to 6 h post-PH by inhibiting protein
synthesis
with cycloheximide
at the 3-h time point post-PH.
The decay rate of the Rb protein
was determined
on immunoblots
using video
densitometry
as described
above.
Because Rb was almost undetectable
in 0-h liven, the 3-h time
point after PH was designated
as the initial decay point from
which
to determine
the half-life.
Remnant
Rb protein
at
3+n h after decay was calculated
by standard
regression
analysis
using the formula
[fts3+nhPH
ft”3+nhPH]
[fts6hPH
_
ft6hPi-i],
where
t represents
time, s represents
protein
synthesis,
d represents
protein
degradation,
and PH
is post-partial
hepatectomy.
Acknowledgments
We are especially
grateful
to Dr. Ed Harlow
and Chidi
Ngwu (Massachusetts
General
Hospital,
Harvard
Medical
School,
Charlestown,
MA) for mAbs to
the Rb and Rb-related
proteins.
We also thank
Drs. Cary
Mariash
and
Yuichiro
Sudo for isolation
of the primary
hepatocytes,
Dr. Richard
Stockert
(Albert
Einstein
College
of Medicine,
Bronx,
NY) for the hepatoma
cell line
HuH-7,
Dr. Wen-Hwa
Lee (University
of Texas Health
Science
Center,
San
Antonio,
TX) for helpful
discussions,
and Janeen Trembley
for critical
reading
of the manuscript.
References
1 . Friend,
S. H., Bernards,
R., Rogelj,
S., Weinberg,
R. A., Rapaport,
Albert,
D. M., and Dryja, T. P. A human
DNA segment
with properties
gene that predisposes
to retinoblastoma
and osteosarcoma.
Nature
323: 643-646,
1986.
J. M.,
of the
(Lond.),
1 1 . DeCaprio,
W-H.,
Marsilio,
J. A., Ludlow,
E., Paucha,
antigen
forms
a specific
susceptibility
gene. Cell,
1 2. Dyson,
N.,
papilloma
virus-i
product.
Science
Growth
I. W., Figge, I., Shew, J-Y.,
E., and Livingston,
D. M.
complex
with
the product
54: 275-283,
1988.
Howley,
P. M., MOnger,
6 E7 oncoprotein
is able
(Washington
DC), 243:
1 3. Horowitz,
J. M., Yandell,
D. W.,
Buchkovich,
K., Harlow,
E., Weinberg,
tional
inactivation
ofthe
retinoblastoma
DC), 243: 937-940,
1989.
14. Hensey,
C. E., Hong,
F., Durfee,
T., Qian, Y-W.,
W-H.
Identification
of discrete
structural
domains
Lee, E. Y-H. P., and Lee,
in the retinoblastoma
protein.
I.
Biol.
Chem.,
269:
1380-1387,
1994.
1 5. Hu, Q., Dyson,
N., and Harlow,
E. The regions
of the retinoblastoma
protein
needed
for binding
to adenovirus
E1A or SV4O large T antigen
common
sites for mutations.
EMBO J., 9: 1 1 47-1 1 55, 1990.
17. Chittenden,
binding
domain
a sequence-specific
T., Livingston,
D. M., and Kaelin,
W. G., Jr. The T/E1Aof the retinoblastoma
product
can interact
selectively
with
DNA-binding
protein.
Cell, 65: 1073-1082,
1991.
1 8. Kaelin,
W. G., Jr., Krek, W., Sellers, W. R., DeCaprio,
J. A., Ajchenbaum,
F., Fuchs, C. S., Chittenden,
T., Li, Y., Farnham,
P. J., Blanar,
M. A., Livingston, D. M., and Flemington,
E. K. Expression
cloning
of a cDNA
encoding
a retinoblastoma-binding
protein
with
E2F-like
properties.
Cell,
70: 351364, 1992.
19. Hagemeier,
C., Bannister,
A. J., Cook,
A., and Kouzarides,
T. The
activation
domain
of transcription
factor PU.1 binds the retinoblastoma
(RB)
protein
and the transcription
factor
TFIID
in vitro:
RB shows
sequence
similarity
to TFIID and TFIIB. Proc. NatI. Acad.
Sci. USA, 90: 1580-1
584,
1993.
20. Rustgi,
A. K., Dyson,
N., and Bernards,
R. Amino-terminal
domains
of
c-myc
and N-myc
proteins
mediate
binding
to the retinoblastoma
gene
product.
Nature
(Lond.),
352: 541-544,
1991.
21 . Kim, S. J., Wagner,
M. R. Retinoblastoma
TGF-f32
gene through
334, 1992.
S., Liu, F., O’Reilly,
M. A., Robbins,
P. D., and Green,
gene
product
activates
expression
of the human
transcription
factor ATF-2.
Nature
(Lond.),
358: 331-
22. Qian,
Y-W., Wang,
Y-C. J., Hollingsworth,
and Lee, E. Y-H. P. A retinoblastoma-binding
regulator
of Ras in yeast. Nature
(Lond.),
364:
R. E., Jr., Jones, D., Ling, N.,
protein
related
to a negative
648-652,
1993.
USA,
24. Hu, Q., Lees, J. A., Buchkovich,
protein
physically
associates
with
12:971-980,
1992.
genes.
Science
(Washington
DC),
254:
5. Gu, W., Schneider,
J. W., Condorelli,
Nadal-Ginard,
B. Interaction
of myogenic
protein
mediates
muscle
cell commitment
309-324,
1993.
G.,
Kaushal,
S., Mahdavi,
V., and
factors
and the retinoblastoma
and differentiation.
Cell,
72:
6. Xu, H-i., Hu, S-X., Hashimoto,
T., Takahashi,
R., and Benedict,
W. F. The
retinoblastoma
susceptibility
gene product:
a characteristic
pattern
in normal
cells and abnormal
expression
in malignant
cells. Oncogene,
4: 807-81
2,
1989.
7. Buchkovich,
is phosphorylated
1105, 1989.
K., Duffy,
during
L. A., and Harlow,
E. The
specific
phases of the cell
8. Chen,
P-L., Scully,
P., Shew,
phorylation
of the retinoblastoma
cycle and cellular
differentiation.
retinoblastoma
cycle.
Cell,
58:
protein
1097-
J-Y., Wang,
I. Y. J., and Lee, W-H.
Phosgene product
is modulated
during
the cell
Cell, 58: 1193-1198,
1989.
9. Furukawa,
Y., DeCaprio,
I. A., Freedman,
A., Kanakura,
Y., Nakamura,
M., Ernst, T. I., Livingston,
D. M., and Griffin,
J. D. Expression
and state of
phosphorylation
of the retinoblastoma
susceptibility
gene product
in cycling
and noncycling
human
hematopoietic
cells. Proc. NatI. Acad. Sci. USA, 87:
2770-2774,
1990.
10. Whyte,
P., Buchkovich,
K. J., Horowitz,
I. M., Friend,
S. H., Raybuck,
M., Weinberg,
R. A., and Harlow,
E. Association
between
an oncogene
and
an anti-oncogene:
the adenovirus
E1A proteins
bind to the retinoblastoma
gene product.
Nature
(Lond.),
334: 124-129,
1988.
are
16. Huang,
S., Wang,
N-P., Tseng,
B. Y., Lee, W-H.,
Lee, E. H-H.
P. Two
distinct
and frequently
mutated
regions
of retinoblastoma
protein
are required
for binding
to SV4O T antigen.
EMBO
J., 9: 1815-1822,
1990.
associated
suppressor
human
gene
Canning,
S., Whyte,
P.,
Dryja,
T. P. Point mutaScience
(Washington
3. Lee, W-H.,
Shew, J-Y., Hong,
F. D., Sery, T. W., Donoso,
L. A., Young,
L-J., Bookstein,
R., and Lee, E. Y-H. P. The retinoblastoma
susceptibility
gene
encodes
a nuclear
phosphoprotein
associated
with
DNA
binding
activity.
Nature
(Lond.),
329: 642-645,
1987.
R. A. Tumor
retinoblastoma
Park, S-H.,
R. A., and
antioncogene.
J. A., Penman,
cycle-dependent,
1991.
1475
C-M.,
Lee,
large tumor
K., and Harlow,
E. The
to bind to the retinoblastoma
934-937,
1989.
23. Mancini,
M. A., Shan, B., Nickerson,
The retinoblastoma
gene product
is a cell
1138-1146,
Huang,
SV4O
of the
2. Lee, W-H.,
Bookstein,
R., Hong,
F. D., Young,
L-J., Shew, J-Y., and Lee,
E. Y-H. P. Human
retinoblastoma
susceptibility
gene: cloning,
identification,
and sequence.
Science
(Washington
DC), 235: 1394-1399,
1987.
4. Weinberg,
& Differentiation
protein.
Proc.
NatI.
Acad.
Sd.
K. J., and
the human
S., and Lee, W-H.
nuclear
matrix-
91:418-422,
1994.
Harlow,
E. The retinoblastoma
cdc2 kinase.
Mol. Cell. Biol.,
25. Durfee,
T., Becherer,
K., Chen,
P-L., Yeh,
Lee, W-H.,
and Elledge,
S. J. The retinoblastoma
protein
phosphatase
type 1 catalytic
subunit.
1993.
S-H., Yang, Y., Kilburn,
A. E.,
protein
associates
with the
Genes
& Dev.,
7: 555-569,
26. Dowdy,
S. F., Hinds,
P. W., Louie,
K.,
Weinberg,
R. A. Physical
interaction
of the
human
D cyclins.
Cell, 73:499-511,
1993.
Reed,
S. I., Arnold,
A., and
retinoblastoma
protein
with
27. Ewen,
Livingston,
mammalian
M. E., Sluss, H. K., Sherr, C. J., Matsushime,
H.,
D. M. Functional
interactions
of the retinoblastoma
D-type
cyclins.
Cell, 73: 487-497,
1993.
Kato, J-y., and
protein
with
28. Goodrich,
D. W., Wang,
N. P., Qian,
Y-W.,
Lee, E. Y-H. P., and Lee,
W-H. The retinoblastoma
gene product
regulates
progression
through
the G1
phase of the cell cycle.
Cell, 67: 293-302,
1991.
29. Goodrich,
retinoblastoma
1993.
D. W., and
susceptibility
Lee, W-H.
Molecular
characterization
gene.
Biochim.
Biophys.
Acta,
1 155:
30. Shan, B., Chang,
C-Y., Jones,
E2F-i
mediates
the autoregulation
14:299-309,
1994.
D., and Lee, W-H.
The transcription
of RB gene expression.
Mol. Cell.
31 . Higgins,
G. M., and Anderson,
R. M. Experimental
I. Restoration
of the liver of the white
rat following
Arch.
Pathol.,
12: 186-202,
1931.
32. Grisham,
J. W. A morphologic
and cell proliferation
in regenerating
dine-H3.
Cancer
Res., 22: 842-849,
of the
43-61,
factor
Biol.,
pathology
ofthe
liver.
partial
surgical
removal.
study of deoxyribonucleic
rat liver: autoradiography
1962.
acid synthesis
with thymi-
1476
Rb Protein
Expression
in Regenerating
33. Fausto,
N. Hepatic
Hepatology:
A Textbook
Saunders,
1990.
Rat Liver
regeneration.
In: D. Zakim
and T. D. Boyer
of Liver Disease,
pp. 49-65.
Philadelphia:
34. Haber,
B. A., Mohn,
patterns
of 70 genes during
course
of liver regeneration.
K. L., Diamond,
R. H., and Taub,
nine days after hepatectomy
define
J. Clin. Invest.,
91: 1319-1326,
R. Induction
the temporal
1993.
35. Thompson,
N. L., Mead,
J. E., Braun,
L., Goyette,
M., Shank,
Fausto,
N. Sequential
protooncogene
expression
during
rat liver
tion. Cancer
Res., 46:3111-3117,
1986.
36. Kren, B. T., Teel, A. L., and Steer,
state changes
of retinoblastoma
mRNA
19:1214-1222,
1994.
(eds.),
W. B.
38. Hu,
Harlow,
a family
J.
Bautista,
C., Edwards,
G. M., Defeo-Jones,
D., Jones, R. E., and
E. Antibodies
specific
for the human
retinoblastoma
protein
identify
of related
polypeptides.
Mol. Cell. Biol.,
1 1: 5792-5799,
1991.
42. Haupt,
Y., Rowan,
cells can be overcome
J.
Cyclins
and cancer
79: 573-582,
1994.
II: cyclin
D and
CDK
R., Peleg, D., and Meyuhas,
0.
posttranscriptional
regulation
development
and regeneration
2203-2212,
J. A., Holt,
suppression
proliferation.
Beach,
CDK2
D. Isolation
and
E. DNA-binding
Mol. Cell. Biol.,
cyclins.
of the Rb-related
Genes
properties
of the E1A-associ12: 2826-2836,
1992.
J. T.,
f31
Stein, R. W., and Moses,
H. L. Transforming
of c-myc
gene transcription:
role in inhibition
Proc. NatI. Acad.
Sci. USA, 87: 3758-3762,
Y., and Weinberg,
R. A. Transforming
of G1 cyclins
and cyclin-dependent
USA, 90: 10315-10319,
1993.
growth
protein
factor
kinases.
61 . Ewen, M. E., Sluss, H. K., Whitehouse,
L. L., and
inhibition
of Cdk4 synthesis
is linked
to cell cycle
1020,
1993.
62. Koff, A., Ohtsuki,
M., Polyak,
K., Roberts,
J. M., and Massagu#{233}, J.
Negative
regulation
of G1 in mammalian
cells: inhibition
of cyclin
E-dependent kinase by TGF-)3.
Science
(Washington
DC), 260: 536-538,
1993.
Polyak,
K., Kato, J-y., Solomon,
M. 1., Sherr, C. J., Massagu#{233}, J., Roberts,
and Koff, A. p27K,
a cyclin-Cdk
inhibitor,
links transforming
growth
factor-f3
and contact
inhibition
to cell cycle
arrest. Genes
& Dev.,
8: 9-22,
1994.
,
64. Hatakeyama,
M., Brill, J. A., Fink,
ration
of G1 cyclins
in the functional
protein.
Genes & Dev.,
8: 1759-1771,
G. R., and
inactivation
1994.
Weinberg,
of the
66. Muller,
H., Lukas, J., Schneider,
A., Warthoe,
and Strauss,
M. Cyclin
Dl expression
is regulated
protein.
Proc. NatI. Acad.
Sci. USA, 91: 2945-2949,
F. Lack of
to changes
K. G., Klein,
protein.
Cell
nuclear
in total
RB protein
RB protein
67. Mariash,
C. N., Seelig,
Rapid synergistic
interaction
mRNA514
induction.
J. Biol.
P., Bartek,
J., Eilers, M.,
by the retinoblastoma
1994.
S., Schwartz,
H. L., and
between
thyroid
hormone
Chem.,
261:9583-9586,
Oppenheimer,
and carbohydrate
1986.
48. Slack,
R. S., Hamel,
P. A., Bladon,
T. S., Gill,
M. W. Regulated
expression
of the retinoblastoma
embryonal
carcinoma
cells. Oncogene,
8: 1585-1591,
69. Bates,
F. Apoptosis
403-415,
R. M., and McBurney,
gene in differentiating
1993.
of purified
retinoblastoma
phosphorylation.
Mol.
gene
Cell.
prodBiol.,
Mittnacht,
Wang,
J. Y. I. Cell cycle
Ciba Found.
Symp.,
S., Lees,
J.
A.,
Desai,
R. A. Distinct
sub-populations
pattern
of phosphorylation.
D.,
Harlow,
E., Morgan,
D.
protein
1994.
53. Welch,
P. J., and Wang,
J. Y. J. A C-terminal
protein-binding
the retinoblastoma
protein
regulates
nuclear
c-Abl tyrosine
kinase
cycle.
Cell, 75: 779-790,
1993.
0.,
method
of RNA
extraction.
72. Holland,
E. C., Leung,
J. Q., and Drickamer,
K. Rat liver
protein
receptor
lacks a cleavable
NH2-terminal
signal sequence.
Acad Sd. USA, 81: 7338-7342,
1984.
and
show
domain
in
in the cell
54. Ewen, M. E., Xing, Y., Lawrence,
J. B., and Livingston,
D. M. Molecular
cloning,
chromosomal
mapping,
and expression
of the cDNA
for plO7,
retinoblastoma
gene product-related
protein.
Cell, 66: 1 1 55-1 1 64, 1 991.
Brenner,
D. A.
active
liver
isolation
Anal.
by
Bio-
71 . Bernards,
R., Schackleford,
G. M., Gerber,
M. R., Horowitz,
J. M.,
Friend,
S. H., Schartl,
M., Bogenmann,
E., Rapaport,
J. M., McGee,
T., Dryja,
T. P., and Weinberg,
R. A. Structure
and expression
of the murine
retinoblastoma
gene and characterization
of its encoded
protein.
Proc. NatI. Acad
Sci. USA, 86:6474-6478,
1989.
regulation
of retinoblastoma
170: 227-243,
1992.
ofthe
retinoblastoma
EMBO
J., 13: 118-127,
J. H.
on
R. C., Buret, A., van Helden,
D. F., Horton,
M. A., and Burns, G.
induced
by inhibition
of intercellular
contact.
I. Cell Biol.,
125:
1994.
70. Chomczynski,
P., and Sacchi,
N. Single-step
acid
guanidinium
thiocyanate-phenol-chloroform
chem.,
162: 156-159,
1987.
1992.
50. Cobrinik,
D., Dowdy,
S. F., Hinds,
P. W., Mittnacht,
S., and Weinberg,
R. A. The retinoblastoma
protein
and the regulation
of cell cycling.
Trends
Biochem.Sci.,
17:312-315,
1992.
52.
R. A. Collaboretinoblastoma
Durko,
M., Wiman,
ofthe
retinoblastoma
68. Hattori,
M., Tugores,
A., Veloz,
L., Karin,
M., and
A simplified
method
for the preparation
of transcriptionally
nuclear
extracts.
DNA
Cell Biol.,
9: 777-781,
1990.
Weinberg,
a distinct
of liver
in lipo-
Livingston,
D. M. TGFf3
arrest.
Cell,
74: 1009-
47. Savatier,
P., Huang,
S., Szekely,
L., Wiman,
K. G., and Samarut,
J.
Contrasting
patterns
of retinoblastoma
protein
expression
in mouse
embryonic stem cells and embryonic
fibroblasts.
Oncogene,
9: 809-818,
1994.
51 . Lin, B. T. Y., and
protein
phosphorylation.
j3 effects on
Proc. NatI.
65. Kato, J-y., Matsushime,
H., Hiebert,
S. W., Ewen, M. E., and Sherr, C. J.
Direct
binding
of cyclin
D to the retinoblastoma
gene product
(pRb) and pRb
phosphorylation
by the cyclin
D-dependent
kinase CDK4.
Genes & Dev., 7:
331-342,
1993.
46. Xu, H-i.,
Hu, S-X., and Benedict,
W.
staining
in G1Imiddle
G1 cells: correlation
level. Oncogene,
6: 1 139-1
146, 1991.
12:435-443,
7:
I. M.,
in HeLa
1995.
1992.
D. J. Nuclear
binding
by cell cycle-regulated
Dev.,
Selective
translational
control
and
of ribosomal
protein
gene expresof rat liver. Mol. Cell. Biol.,
12:
45. Szekely,
L., Uzvolgyi,
E., jiang, W-Q.,
G., and Sumegi,
I. Subcellular
localization
Growth
& Differ.,
2:287-295,
1991.
49. Templeton,
uct is determined
&
63.
S., and Oren,
M. p53-mediated
apoptosis
by excess pRB. Oncogene,
10: 1 563-1 571
43. Sheets, M. D., Fox, C. A., Hunt, T. Vande Woude,
G., and Wickens,
M.
The 3-untranslated
regions
ofc-mos
and cyclin
mRNAs
stimulate
translation
by regulating
cytoplasmic
polyadenylation.
Genes
& Dev.,
8: 926-938,
1994.
44. Aloni,
nonspecific
sion during
with
60. Schackert,
H. K., Fan, D., and Fidler,
I. J. Transient
inhibition
regeneration
in mice by transforming
growth
factor-f31
encapsulated
somes.
Cancer
Commun.,
2: 165-171,
1990.
M., and
of retino-
40. Russell,
W. E., Coffey,
R. I., Jr., Ouellette,
A. J., and Moses,
H. L. Type
j3 transforming
growth
factor
reversibly
inhibits
the early
proliferative
response
to partial
hepatectomy
in the rat. Proc. Nat). Acad.
Sci. USA, 85:
5126-5130,
1988.
41. Hunter,
T., and Pines,
inhibitors
come of age. Cell,
D., and
its interaction
1993.
58. Pietenpol,
J. A., Munger,
K., Howley,
P. M., Stein, R. W., and Moses,
H. L. Factor-binding
element
in the human
c-myc
promoter
involved
in
transcriptional
regulation
by transforming
growth
factor 3i and by the retinoblastoma
gene product.
Proc. NatI. Acad.
Sci. USA, 88: 10227-10231,
1991.
59. Geng,
expression
Acad.
Sd.
Q.,
D.
G. J., Demetrick,
57. Pietenpol,
growth
factor
of keratinocyte
1990.
C. J. Transcriptional
rate and steadyin regenerating
rat liver. Hepatology,
39. Laiho,
M., DeCaprio,
j. A., Ludlow,
J. W.,
Livingston,
Massagu#{233}, I. Growth
inhibition
by TGF-f3
linked
to suppression
blastoma
protein
phosphorylation.
Cell, 62: 1 75-1 85, 1990.
Hannon,
p1 30 through
2378-2391,
56. Rikitake,
Y., and Moran,
ated 300-kilodalton
protein.
P. R., and
regenera-
37. Nakabayashi,
H., Taketa,
K., Miyano,
K., Yamane,
T., and Sato,
Growth
of human
hepatoma
cell lines with differentiated
functions
in chemically
defined
medium.
Cancer
Res., 42: 3858-3863,
1982.
55.
a
73. Feinberg,
A. P., and
restriction
endonuclease
132:6-13,
1983.
74. Correa-Rotter,
transfer
control
1992.
for
Vogelstein,
fragments
B. A technique
to high specific
R., Mariash,
C. N., and
northern
hybridization.
asialoglycoProc. NatI.
for radiolabelling
DNA
activity.
Anal. Biochem.,
Rosenberg,
M.
BioTechniques,
E. Loading
and
12: 1 54-1 58,