* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Vigneshwaran Mani
Epigenomics wikipedia , lookup
Transposable element wikipedia , lookup
Oncogenomics wikipedia , lookup
Point mutation wikipedia , lookup
Metagenomics wikipedia , lookup
Fetal origins hypothesis wikipedia , lookup
Minimal genome wikipedia , lookup
Genetic engineering wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Ridge (biology) wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Gene desert wikipedia , lookup
Gene therapy wikipedia , lookup
Genomic imprinting wikipedia , lookup
Gene nomenclature wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Genome evolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Helitron (biology) wikipedia , lookup
Public health genomics wikipedia , lookup
Genome (book) wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Gene expression programming wikipedia , lookup
Microevolution wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Designer baby wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
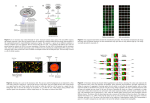
DNA arrays for early disease detection Chem 395 Bioanalytical chemistry Instructor Prof.James F.Rusling Presented by Vigneshwaran Mani Outline What is a Microarray? Types of Microarray Steps involved in Microarray fabrication What happens to the Genes in disease state? Application of Microarray Summary What is a Microarray? A microarray is a spatially ordered, minituarized arrangement of multitude of immobilized reagents Why Microarrays????? Small volume- miniaturization High throughput analysis Large information generated Less time required to analyze Types of Microarrays cDNA and Oligonucleotide Microarrays Probe: ssDNA,Oligonucleotide Target: ssDNA,ssRNA Principle: Hybridization Protein Microarrays Probe: Antibody Target: Antigen Principle: peptide chemistry Others Tissue arrays Gene expression Nucleus RNA POLYMERASE RIBOSOMES Cytoplasm cDNA ARRAYS Marketed By: Agilent technologies Steps Involved in Microarrays Microarray printing Hybridization Detection DNA-Microarray printing 100-10000 spots Glass slide used as substrate DNA is attached covalently to glass slide 96-384 well microtitre plates used Spot Volume 0.25-1nl. Spot size 100-150µm in diameter • • • • • • 1,2,3-X,Y,Z direction 4-print head 5-glass slide 6-Microtitre plate 7-Distilled water 8- Drying Hybridization mRNA Control A C U mRNA Diseased G A Reverse transcriptase T G A C G A U G Reverse transcriptase cDNA T G C A Fluorescent labeling Fluorescent labeling Cy3 T C Cy5 C T G A Hybridize target to microarray C Hybridization Detection: The slide is scanned twice -Once to measure red intensity -Once to measure green intensity The images are overlayed to produce one image Scanning M = logR/G = logR – logG M<0: gene is over-expressed in green-labeled sample compared to red-labeled sample. M=0: gene is equally expressed in both samples. M>0: gene is over-expressed in red-labeled sample compared to green-labeled sample. What happens to Genes in (Cancer) Disease State Certain genes undergo overexpression. No. of copies of particular genes may increase. Gene mutation. Changes in gene expression levels Gene is overexpressed in a certain disease state, More cDNA(target) will hybridizes to probe, as compared to control cDNA, In turn, the spot will fluorescence red with greater intensity than it will be with green. Expression patterns of various genes is characterized involved in many diseases, Compare expression pattern of the gene from the individual with the expression pattern of a known disease. Genomic gains and losses Number of copies of a particular target gene has increased. Large amount of (Disease) sample DNA will hybridize to those spots on the microarray compared to (normal) control DNA hybridizing to those same spots. Those spots containing the sample DNA will fluoresce red with greater intensity than they will fluoresce green, indicating that the number of copies of the gene involved in the disease has gone up. Gene Expression profile analysis in Human hepatocellular carcinoma by cDNA microarray Eun Jung Chung,Young Kwan Sung,MohammadFarooq,Younghee Kim, Sanguk Im,Won Young Tak,Yoon Jin Hwang, Yang Il Kim, Hyung Soo Han, Jung-Chul Kim, and Moon Kyu Kim.,Mol.Cells,Vol.14,pp382-387,2002 HCC (Hepatocellular carcinoma) Primary liver cancer (HCC) Somatic mutations and activation of certain oncogenes. These events Lead to expression changes in genes. cDNA arrays are used to analyze expression patterns cDNA arrays Computer analysis 3063 human cDNA 8 different samples of HCC Up-regulated genes in HCC Galectin-3 Serine/threonine kinase Fibroblast growth factor receptor Ribosomal protein L35A Down-regulated genes in HCC mRNAs of Nip3 Decorin Insulin-like growth factor binding protein-3 Gene expression patterns of 8 hepatocellular carcinomas. The genes were primarily classified into three groups, based on their clustering pattern. Application of Microarrays Application cDNA array. Tumor classification, risk assessment, and prognosis prediction. Expression analysis. Drug development, drug response, and therapy development Summary Data can be generated in a high throughput, parallel fashion. Less time required for analysis. If gene expression data is already known for a certain disease. Then we can compare the gene expression data of the individual with the known, and predict the disease References David J. Duggan, Michael Bittner, Yidong Chen, Paul Meltzer & Jeffrey M. Trent, Nature genetics supplement, volume 21, january 1999. Sunil R. Lakhani, Michael J.O’hare & Alan ashworth, Nature medicine,volume 7,number 4,april 2001. Vivian G. Cheung, Michael Morley, Francisco Aguilar, Aldo Massimi,Raju Kucherlapati & Geoffrey Childs, Nature genetics supplement,volume 21,january 1999. http://www.ncbi.nlm.nih.gov/ Acknowledgement Prof. James F. Rusling Chem 395 students