* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Document
Transposable element wikipedia , lookup
History of genetic engineering wikipedia , lookup
Non-coding DNA wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
X-inactivation wikipedia , lookup
Gene therapy wikipedia , lookup
Genome evolution wikipedia , lookup
Point mutation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Microevolution wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
Designer baby wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Helitron (biology) wikipedia , lookup
Polyadenylation wikipedia , lookup
Messenger RNA wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Epigenetics of human development wikipedia , lookup
History of RNA biology wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
RNA-binding protein wikipedia , lookup
Epitranscriptome wikipedia , lookup
Non-coding RNA wikipedia , lookup
Primary transcript wikipedia , lookup
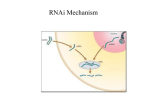
REVIEW RNA INTERFERENCE G. Russev Institute of Molecular Biology, Bulgarian Academy of Sciences, Sofia, Bulgaria Correspondence to: George Russev Email: [email protected] ABSTRACT RNA interference (RNAi) refers to the ability of double-stranded RNA molecules to cause sequence-specific degradation of single stranded RNAs such as messenger RNAs and viral RNAs in vivo. RNAi is an ancient mechanism of gene regulation, genome maintenance and antiviral defense found in all eukaryotes. The effector molecules are 21 to 23 nucleotides small interfering RNAs (siRNAs) that are anticipated to serve as novel therapeutic agents in the battle against cancer, AIDS and neurodegenerative diseases. In the laboratory, RNAi is used to investigate complex biological phenomena such as development, cell signaling and infection, and thanks to its application many important breakthroughs have been made in these and related areas during the last years. On the other hand its therapeutic application in the clinic may be few years away. The main obstacle in this field is not the RNAi itself, but rather problems connected with the targeting and delivery of the therapeutic siRNAs. Keywords: RNA interference, RNAi, small interfering RNA, gene regulation, gene silencing, antiviral treatment; Introduction RNA interference (RNAi) is a highly evolutionally conserved process of post-transcriptional gene silencing (PTGS) by which double stranded RNA (dsRNA) causes sequencespecific degradation of homologous mRNA sequences, when introduced into cells. There is general agreement that it is very strange indeed that such a fundamental cellular regulatory and defense pathway has remained unnoticed for so long. First R. Jorgensen and colleagues (4), while trying to improve the coloration of petunias by introducing extra copies of genes responsible for pigmentation, noticed that in some cases instead of intensification, colors were entirely or partially lost. They correctly concluded that the treatment has switched the respective genes off. Later on the same results were obtained with other organisms (2, 6, 8) and it soon became evident that the phenomenon represents a major and universal regulatory pathway in the eukaryotic cells. However, the mechanism of this process remained elusive until 1998 when Andrew Fire, Craig Mello and colleagues published their fundamental paper ”Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans” in Nature (1). In this paper they described the mechanism of the so called gene silencing and used for the first time the popular name by which it is known since then – RNA interference. In the following years RNAi became a major tool in the life science experiments and a promising therapeutic opportunity. Its impact on the development of the biological sciences was so deep and profound that it came as no surprise when in 2006 the Nobel committee awarded the Nobel prize for physiology or medicine to Fire and Mello for discovering the RNA interference. Mechanism of RNAi The first step in protein synthesis, transcription, takes place in the cell’s nucleus, where the DNA template is used to make single strand mRNA. The RNA then exits the nucleus and enters BIOTECHNOL. & BIOTECHNOL. EQ. 21/2007/3 the cytoplasm, where it, in its turn serves as template in the translation, the process of protein synthesis on the ribosomes. RNA interference acts between the steps of transcription and translation. It has long been known that introduction of RNA into cells interferes with the function of the genes (3, 5). These effects have been proposed to result from the so called “antisense” mechanism that depends on hybridization between the exogenous RNA and endogenous messenger RNA transcripts thus blocking the translation of the latter into proteins. However, antisense RNA alone was marginally effective in silencing the targeted genes and the reason for this poor performance was not clear. Fire and Mello set themselves to study the requirements for structure and delivery of the interfering RNA (1). As a model organism they used the nematode C. elegance and as a model gene - the so called ung-22 gene, which encodes a non-essential myofilament protein. The expression of this gene is easily detected and monitored since the decrease, or the absence of the respective protein produced obvious and specific phenotypic characteristic – twitching of the worms. After injection of purified single strand or double strand RNAs, complementary to regions of the unc-22 gene, Fire and Mello found out that dsRNA was much more effective in the silencing of the gene than either of the strands. In another series of experiments they used specific reporter gene – the gene for the green fluorescent protein (GFP) and in this case again the dsRNA targeted to the gene was much more effective in blocking the expression of the GFP than either of the strands alone. Thus they came to the conclusion that dsRNA is much more potent and specific inhibitor of the gene expression then single strand RNA. Secondly they noticed that even if only a few copies of the respective dsRNA were present in the cell, they can completely prevent translation of highly abundant RNAs, which indicates that the process of RNA interference occurs by the involvement of a complex, but at that time totally unknown cellular catalytic mechanism and not by a conventional antisense mechanism in which sense and antisense molecules hybridize in 1:1 ratio. A major step in 283 the deciphering of this mechanism was made in 2001 when G. Hannon (5) reported the discovery of 2 specific enzyme complexes with ribonuclease activity. The first one called Dicer cuts dsRNA into small uniform double-stranded pieces 21-23nt in length called small interfering RNAs (siRNA). Recent works have shown that Dicer acts as a molecular ruler measuring pieces of dsRNA in the proper 23nt range and cutting them off. Dicer contains a conserved dsRNA binding domain called PAZ and 65 angstrom (which corresponds to 25 nt) apart a RNA cutting domain containing 2 RNAse III sites that cut across the dsRNA. The second enzyme, called RISC (RNA Induced Silencing Complex) cuts the target mRNA. RISC is the conglomeration of several proteins including certain RNA unwinding proteins, and a protein called Argonaute which is central for the RNA cutting endonuclease activity of RISC. First Argonaute degrades one of the strands of the siRNA called “passenger” strand. The strand selection is carried out on the basis of the thermodynamic stability of the siRNA duplex termini, the strand whose 5’-terminus has higher base-pairing stability being degraded. Thus in its final form RISC is loaded with only one of the siRNA strands, called “guide strand” that forms a temporary complex with the target mRNA on the basis of sense-antisense complementation, and cuts it. The RISC complex with the guide RNA strand is quite stable and can successively degrade many mRNA molecules thus acting as a catalyst (7). The whole process of RNA interference is shown in (Fig. 1). Role of RNAi in eukaryotic cells The finding that dsRNA introduced into eukaryotic cells executes a gene-silencing effect using cell’s own enzymatic activities and protein complexes, shows that the process of RNA interference is a normal cellular pathway evolved to perform specific tasks. These include antiviral defense, regulation of the gene expression and protection of the genome. • Antiviral defense mechanism Prokaryotic cells have developed the system of restriction enzymes, which cut any foreign DNA that eventually appears in the cells. The higher organisms also possess effective ways to fight foreign DNA, the most common being the virus induced apoptosis. To avoid these defense mechanisms many eukaryotic viruses have evolved into RNA viruses and many plant, animal and human viruses such as HIV for instance, are dsRNA viruses. Eukaryotic cells have reacted to this challenge by developing the RNA interference. By this pathway any viral dsRNA is chopped by the cellular Dicers and then incorporated into the cellular RISCs to degrade the viral mRNAs. Thus RNAi seems to serve as an antiviral immune system evolved to protect eukaryotic cells from invasion of foreign genetic (RNA) material. • Gene expression regulation Since RNAi degrades mRNAs and is conveniently positioned between transcription and translation, it can regulate the gene expression, i.e. the rate of the respective gene product synthesis, without changing the gene activity. Cells are using this way to achieve “fine tuning” of the expression without interfering with transcription. It has long been known that a class of short dsRNA molecules called micro RNA (miRNA) 284 Fig. 1. RNA interference. (A) On entering the cell, long ds RNA act as a trigger of RNAi process. (B) It is first processed by the RNAse III enzyme Dicer in an ATP-dependent reaction. (C) Dicer processed dsRNA into 21-23 nt short interfering RNA (siRNA) with 2 nt 3’-overhangs. SiRNA can also be synthesized outside the cell and then introduced into a cell. (D) The siRNAs are incorporated into the RNA-inducing silencing complex (RISC) which consists of the RNAse Argonaute as one of its main components. At this point one of the RNA strands of the dsRNA (passenger strand) is degraded. (E) RISC’s guide siRNA strand is complexed with the target mRNA on the basis of senseantisense hybridization. (F) mRNA is cut by RISC and RISC itself is released for interaction with another molecule mRNA exists in eukaryotic cells. However, only lately it has been discovered that they play the role to control the amount of different mRNAs. They are transcribed from regions of the genome with specific sequence symmetry and after transcription the resulting RNAs fold back to form a hairpin structures with double-strand stem and a loop. These RNAs are called hairpin RNAs. They are recognized by the Dicers and chopped to give small hairpin RNAs (shRNA). They have homology to different mRNAs and could degrade the latter thus maintaining the mRNA populations within certain limits. Over 30% of the genes are fine regulated by this mechanism. • Genome stability maintenance The RNAi is also used to control the spread of transposons. Transposons are like the viruses that are inserted into the genomes. A good example are the Alu elements which represent a few hundred bp DNA sequence repeated over 105 times in the human genome. Transposons can arrange their own transcription to produce RNA copies. These RNA copies are then used as templates to synthesize complementary DNA by a BIOTECHNOL. & BIOTECHNOL. EQ. 21/2007/3 process known as reverse transcription. Then the DNA strand is replicated by the normal cellular replication machinery to produce dsDNA copies of the original Alu element, which are inserted randomly in the genome. If left unchecked this process would lead to ever increasing number of Alu elements. Apart from increasing the so called “junk” DNA this process represents a treat to the integrity of the genome since the Alu elements could be inserted in exons or other important regions and to interfere with their normal function. To avoid this cells recruit the RNAi mechanism. Certain DNA regions contain sequences which after transcription give rise to shRNAs with homology to the transposon sequences. They are cut by the Dicers and used by the RISCs to degrade the initial transposone RNA transcripts thus maintaining the genome stability. RNAi application RNAi holds many promises as antiviral treatment and for controlling gene expression in eukaryotic cells. However, for the time being it is only used as experimental tool. There is hardly any molecular biology or molecular genetics lab in the world that is not using RNAi to knock down different genes to study their functioning. The procedure is simple, and the “knock down” is reversible which represent clear advantages over the so called “knock out” procedures in which the gene is deleted or irreversibly damaged. As for its application in the clinics – it still seems several years in the future, although a few trials on patients have been reported. In one of them the company Acuity Pharmaceuticals in Philadelphia has completed safety tests on RNAi treatment for macular degeneration, the leading cause for blindness in the elderly. This disease is triggered by the loss of control on the expression of a protein called VEGF (Vascular Endothelial Growth Factor) which is responsible for the formation of the blood vessels. Normally this protein is not expressed in the retina which stays transparent and clear. However, when it is expressed, extensive blood vessel formation occurs there and the retina losses its transparency thus causing blindness. In the trial a group of about 24 people have undergone direct injection of dsRNA against VEGF mRNA in the eye. Two months after being injected with the siRNA, a quarter of the patients had significantly clearer vision and the other patients’ vision has stabilized. Another clinical trial of siRNA based drug was carried out in Cambridge, MA by the pharmaceutical company Alnylam. In this case patients were treated with siRNA against RSV gene by inhalation. RSV (Respiratorial Syncythial Virus) infects almost all children by the age of two. Infection typically leads to cold-like symptoms, but in many cases have far serious consequences such as pneumonia and respiratory failure. The treatment led to reduction of the viral loads. Prospects and problems Although in its infancy, RNAi therapy is expected to cure almost everything from common flu to cancer. However, the hopes are especially high in two fields - those of RNA virus caused diseases such as AIDS and Hepatitis B and C, and of neurological disorders such as Alzheimer’s, Jacob-Croizfeld’s and Hutchington’s diseases. The antiviral strategies aim to use dsRNA to target mRNA for an important viral gene, whose BIOTECHNOL. & BIOTECHNOL. EQ. 21/2007/3 knock down would inhibit viral replication or its ability to infect cells. The same result could also be achieved if host cells are treated with dsRNA against mRNA for cell surfice receptors specific for the virus. In this case the virus will not infect the cells since it will not find the receptors on the cell surface. Most of the neurodegenerative diseases are result of minor nonessential mutations in some of the genes coding for specific class of proteins called amyloids. These mutations occur only in one of the alleles and produce altered proteins which tend to aggregate. The peculiar thing is that this aggregation can involve the normal protein as well thus causing the aggregation of essentially all protein molecules, both mutated and normal, to form extracellular deposits of amyloid filaments. The obvious approach to treat such a condition is to silence the mutated gene thus allowing the normal gene to express the normal protein that would not aggregate alone. In both cases preliminary experiments on animal models with siRNAs have shown very promising results. However, in these and all other cases the problem is not the siRNA itself, but its delivery into the target cells. To achieve the desired therapeutic effect it is not enough siRNA to reach and enter few cells. For successful therapy the siRNA has to be introduced in all cells of the affected tissue which makes the things difficult. Relatively simple are the situations when these tissues are easily accessible and siRNA can be applied directly. Such are the cases with macular degeneration treated by direct injection of siRNA in the eye, the RSV infection, when the drug is administered by inhalation, and of certain vaginal and anal viral infections when the drug is directly applied as ungventum. In all these cases to help siRNA to get into the cells it was complexed with lipids or chemically modified to minimize its charge. On the other hand, for most of the diseases expected to treat with siRNA, the direct application is not an option. To this end specific and at the same time harmless for the host organism vectors have to be constructed. They should be able to infect cells with high efficiency and to integrate in their genomes and should carry cloned DNA sequences that after transcription would give the therapeutic shRNAs. REFERENCES 1. Fire A., Xu S., Montgomery M., Kostas S., Driver S., Mello C. (1998) Nature 391, 806-811. 2. Guo S., Kemphues K. (1995). Cell 81, 611-20. 3. Izant J., Weintraub H. (1984) Cell 36, 1007-1015. 4. Napoli C., Lemieux C., Jorgensen R. (1990) Plant Cell. 2, 279-289. 5. Nellen W., Lichtenstain C. (1993) Trends Biochem. Sci. 18, 419-423. 6. Pal-Bhadra M, Bhadra U, Birchler J. (1997) Cell 90, 479-90. 7. Preall J.B., He Z., Gorra J.M., Sontheimer E.S. (2006) Curr. Biol. 16, 530-535. 8. Romano N, Macino G. (1992). Mol Microbiol 6, 334-353. 285