* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
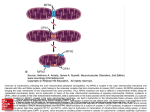
Download Human mitochondrial transfer RNAs: Role of pathogenic
Gene expression programming wikipedia , lookup
Non-coding RNA wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Nutriepigenomics wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
BRCA mutation wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Koinophilia wikipedia , lookup
Genome (book) wikipedia , lookup
Genetic code wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Expanded genetic code wikipedia , lookup
Genome evolution wikipedia , lookup
Population genetics wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Designer baby wikipedia , lookup
Epitranscriptome wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Microevolution wikipedia , lookup
Oncogenomics wikipedia , lookup
Transfer RNA wikipedia , lookup
Frameshift mutation wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Mitochondrial Eve wikipedia , lookup
INVITED REVIEW ABSTRACT: The human mitochondrial genome encodes 13 proteins. All are subunits of the respiratory chain complexes involved in energy metabolism. These proteins are translated by a set of 22 mitochondrial transfer RNAs (tRNAs) that are required for codon reading. Human mitochondrial tRNA genes are hotspots for pathogenic mutations and have attracted interest over the last two decades with the rapid discovery of point mutations associated with a vast array of neuromuscular disorders and diverse clinical phenotypes. In this review, we use a scoring system to determine the pathogenicity of the mutations and summarize the current knowledge of structure–function relationships of these mutant tRNAs. We also provide readers with an overview of a large variety of mechanisms by which mutations may affect the mitochondrial translation machinery and cause disease. Muscle Nerve 37: 150 –171, 2008 HUMAN MITOCHONDRIAL TRANSFER RNAs: ROLE OF PATHOGENIC MUTATION IN DISEASE FERNANDO SCAGLIA, MD, and LEE-JUN C. WONG, PhD Department of Molecular and Human Genetics, Baylor College of Medicine, One Baylor Plaza, Houston, Texas 77030, USA Accepted 17 September 2007 Normal mitochondrial function depends on both the expression of the mitochondrial genome and the expression of at least 1,300 nuclear genes whose products are imported into the mitochondria. Human mitochondrial DNA (mtDNA) is a 16,569-kb circular, double-stranded molecule. It encodes 13 of more than 80 polypeptide subunits of the mitochondrial respiratory chain complexes and contains 24 additional genes required for mitochondrial protein biosynthesis. These include two rRNA genes and 22 tRNA genes, one for each of 20 amino acids, and two each for tRNALeu (UUR and CUN codons) and tRNASer (UCN and AGY codons).140 All partners of the mitochondrial protein synthetic machinery such as aminoacyl-tRNA synthetases, initiation, elongaAvailable for Category 1 CME credit through the AANEM at www.aanem.org. Abbreviations: A, adenine; ATP, adenosine triphosphate; C, cytosine; COX, cytochrome c oxidase; CPEO, chronic progressive external ophthalmoplegia; DHU, dihydrouridine; G, guanine; mtDNA, mitochondrial deoxyribonucleic acid; MELAS, mitochondrial encephalomyopathy, lactic acidosis, and strokelike episodes; MERRF, myoclonic epilepsy with ragged-red fibers; NO, nitric oxide; PEO, progressive external ophthalmoplegia; RC, respiratory chain; ROS, reactive oxygen species; rRNA, ribosomal ribonucleic acid; SNHL, sensorineural hearing loss; T, thymine; tRNA, transfer ribonucleic acid; U, uracil Key words: cardiomyopathy; chronic progressive external ophthalmoplegia; diabetes mellitus; encephalomyopathy; focal segmental glomerulosclerosis; human mitochondrial tRNA genes; mitochondrial cytopathies; mitochondrial myopathy; myoclonic epilepsy; pathogenic mutations; ragged-red fibers; retinitis pigmentosa; sensorineural hearing loss; stroke-like episodes Correspondence to: L. J. C. Wong; e-mail: [email protected] © 2007 Wiley Periodicals, Inc. Published online 9 November 2007 in Wiley InterScience (www.interscience. wiley.com). DOI 10.1002/mus.20917 150 Human Mitochondrial tRNAs tion and termination factors, ribosomal proteins, and enzymes involved in maturation and posttranscriptional modification of tRNAs are nuclear encoded. However, the complete set of 22 tRNAs and two major rRNAs are of mitochondrial origin. The adaptation of the bacterial ancestors of mitochondria to the new environment of eukaryotic cells brought the advantage of ATP production via the process of oxidative phosphorylation. However, the presence in mitochondria of an electron transport chain that continuously reduces oxygen to build up an electrochemical gradient required for ATP synthesis has a potentially deleterious side effect for these eukaryotic cells. This side effect consists of the constant generation of reactive oxygen species (ROS). Due to the continuous function of the electron transport chain, the minimal electron leakage is sufficient to make mitochondrial O2.- generation the major cellular source of ROS in most tissues. MtDNA is located in the mitochondrial matrix and its close proximity to the respiratory chain complexes embedded in the inner membrane renders it susceptible to oxidative stress, causing its high mutation rate (10 –17-fold higher rate of mutations than the nuclear genome).1,122 Since the first description of pathogenic mutations in the mitochondrial genome, over 200 diseasecorrelated point mutations and rearrangements have been found in association with a variety of mitochondrial cytopathies.88 More than half of these mutations have been located in tRNA genes that MUSCLE & NERVE February 2008 constitute ⬃9% of the entire mitochondrial genome.91 Thus, mitochondrial tRNA genes are hotspots for mitochondrial pathogenesis and contribute in a disproportionate way to the etiology of disorders caused by mitochondrial DNA mutations, which is conceivable due to their central role in mitochondrial protein synthesis. In comparison, a little less than half of the mitochondrial mutations affect protein coding genes, which comprises 68% of the entire mitochondrial genome.42 To achieve their goal of carrying amino acids for protein synthesis, these tRNA molecules have to undergo different steps of processing and modification, and interact with many protein factors.128 Disease-related mutations could potentially affect the primary, secondary, and tertiary structures of a given mitochondrial tRNA. Misfolding and instability of the mutant tRNA result in increased degradation and ultimately lead to decreased steady-state levels of a mitochondrial tRNA. In addition, the efficiency of the aminoacylation process will be particularly sensitive to point mutations because the enzymes involved recognize a specific structure of a tRNA.196 There are still many unanswered questions regarding the phenotypic effect of these mutations. Perhaps one of the most intriguing is the incredible diversity of clinical phenotypes associated with mitochondrial tRNA mutations. This is a phenomenon that has been observed even among individuals harboring the same mutation. Several factors including the percentage of heteroplasmy, the different tissues affected with the mutation, the position of the mutation within the structure of the tRNA gene, the efficiency in the utilization of different codons for the same amino acid, and the different amino acid composition among the mitochondrial DNAencoded protein subunits may account for this variability. In addition, nuclear genes, different mitochondrial genetic backgrounds, and environmental factors may also exert a modifier effect. The aim of this review is to focus on the assessment of pathogenic mitochondrial tRNA mutations, the pathogenic mechanism of disease of known and well-established mutations, the description of new pathogenic mutations, and the elucidation of their clinical significance. CANONICAL CRITERIA FOR ASSESSING PATHOGENICITY OF MITOCHONDRIAL tRNA MUTATIONS MtDNA alterations can be divided into three classes: pathogenic, adaptive (advantageous in certain environments), and neutral (accumulated by chance). Human Mitochondrial tRNAs Most of the observed mtDNA changes represent neutral polymorphisms and have been used to track human migrations.70 The large prevalence of variations in tRNA genes calls for the elucidation of their pathogenicity. In addition, clinical misattribution of pathogenicity is an important issue due to the consequences for genetic counseling that is provided to individuals and families with mitochondrial disease. Benign variants can be mistaken as bona fide pathogenic mutations when found in one or a limited number of individuals, especially if present in the heteroplasmic state, which has been regarded as direct evidence for pathogenicity. A set of rules had to be established to confirm the pathogenic nature of novel mtDNA mutations. Di Mauro and Schon35 have previously described the canonical pathogenic criteria for mtDNA point mutations. These rules specified that the mutation should (1) be absent in healthy subjects, (2) alter an evolutionary conserved residue, (3) be heteroplasmic and segregate with disease, (4) be such that the degree of mutant heteroplasmy correlates with clinical severity and cell pathology as documented by single-fiber polymerase chain reaction (PCR), and (5) cause defects of mitochondrial protein synthesis and respiratory chain deficiencies in single or multiple affected tissues of patients and demonstrable in cybrid cell lines. Cybrid cell lines are generated by transferring mitochondria isolated from patients harboring pathogenic mutations into cultured human cells treated with ethidium bromide to deplete their mitochondrial DNA.81 However, some pathogenic mtDNA mutations fail to meet these established canonical criteria. In particular, the homoplasmic mutations often do not demonstrate the required segregation with either biochemical deficiency or clinical disease. It is now known that homoplasmic mitochondrial tRNA mutations can be pathogenic, as demonstrated with several mutations in the tRNASer(UCN) gene,72 the 1624 C⬎T mutation in the tRNAVal that caused multiple neonatal deaths and Leigh syndrome in the offspring of a homoplasmic and mildly affected woman,100 the 4291 T⬎C in the tRNAIle gene associated with hypertension and dyslipidemia,195 the 4300 A⬎G in the tRNAIle gene responsible for maternally inherited cardiomyopathy in two families,173 and the 5693 T⬎C change in the mitochondrial tRNAAsn gene that caused encephalomyopathy.27 In addition, true pathogenic mutations may escape identification because of poor evolutionary conservation in the region or because segregation with disease cannot be demonstrated in small families. MUSCLE & NERVE February 2008 151 In order to take into consideration these known exceptions to the canonical criteria, newer classification systems have been proposed by different groups. Mitchell et al.108 designed a scoring system to assess the pathogenicity of a given sequence variant in the mitochondrial encoded complex I (MTND) subunit genes. Another scoring system specific for the mitochondrial tRNA gene mutations listed on MITOMAP was devised by McFarland et al.101 with specific scoring criteria that gave higher scores to transmitochondrial cybrid experiments, measurements of steady-state levels, and segregation of the mutation with a biochemical defect at the level of single cell. CRITERIA FOR ASSESSMENT OF PATHOGENICITY OF mtDNA ALTERATIONS AND CLASSIFICATION OF MUTATIONS We evaluated the mitochondrial tRNA mutations listed on MITOMAP, which is the largest publicly available compendium of mtDNA mutations and polymorphisms (MITOMAP: A Human Mitochondrial Genome Database; http://www.mitomap.org accessed on January 31, 2007). In addition, we searched PubMed (http://www.ncbi.nlm.nih.gov/ entrez accessed January 31, 2007) in order to analyze a thorough and complete list of mitochondrial tRNA gene mutations. Since the determination of the pathological significance of these mitochondrial tRNA variants is required in order to provide accurate genetic services, we evaluated the mutations in the mitochondrial tRNA genes using a modified scoring system by taking earlier scoring systems101,108 into consideration. Functional studies including steady-state levels of mutated mitochondrial tRNA, evidence of pathogenicity from cybrid cells or single fiber studies, were given the highest scores. We did not score the presence of positive single fibers additively since the levels of the mutant tRNA would be higher in COXnegative fibers than in adjacent COX-positive fibers if there is a decline in steady-state mRNA level or evidence of pathogenicity from transmitochondrial cybrid studies. Positive evidence from any of these experiments was scored a maximum of 9 points. The presence of strong histochemical evidence including COX-negative ragged-red fibers or cristae abnormalities by electron microscopy was assigned 5 points. The respiratory chain (RC) defects were classified according to the modified Walker criteria8 and a score of 5 or 3 points was given for fitting major or minor criteria, respectively. Evolutionary conservation weighed second to functional evidence, with a 152 Human Mitochondrial tRNAs maximum of 8 points. The nucleotide sequences from yeast (Saccharomyces cerevisiae), sea urchin (Strongylocentrotus purpuratus), fruit fly (Drosophila melanogaster), frog (Xenopus laevis), mouse (Mus musculus), cattle (Bos taurus), and human (Homo sapiens) were compared. One point was subtracted for each nucleotide variation occurring in the species listed above. One additional point was subtracted for variations that occurred in each of the four adjacent nucleotides. For nucleotides in the stem region, their complementary nucleotides rather than the adjacent nucleotides were considered. The existence of more than one independent report and the presence of heteroplasmy were given 5 points each. Three points were given for segregation of the mutation with the disease (Table 1). For each mutation, references were evaluated according to the criteria described above and a score was given. If an experiment was not performed or the data were not provided for any reason, a score of 0 was given to that criterion. Based on our scoring system, each mutation could receive a maximum score of 40. Overall, the variations in the mitochondrial tRNA genes that were classified according to these new criteria were assigned to four distinct categories: (1) definite pathogenicity (ⱖ30 points), (2) probable pathogenicity (20 –29 points), (3) possible pathogenicity (10 –19 points), and (4) unlikely pathogenicity (ⱕ9 points). The scores listed in Table 2 represent the minimal scores since not all the relevant experiments to confirm pathogenicity were performed in each case. Table 1. Scoring criteria applied to mitochondrial tRNA mutations listed on MITOMAP and PubMed. Score (points) Pathogenic criteria Evolutionary conservation of the base More than one independent report Presence of heteroplasmy Histochemical evidence RC defects (in tissues) Segregation of mutation with the disease Single fiber studies, or steady-state levels, or cybrid studies No change Each change Yes No Yes No Strong evidence Weak evidence No evidence Strong Moderate No Yes No Yes No 8 ⫺1 5 0 5 0 5 3 0 5 3 0 3 0 9 0 RC, respiratory chain. MUSCLE & NERVE February 2008 Table 2. Structural distribution of tRNA mutations and clinical phenotype. tRNA gene Nucleotide change Region MTTF MTTF MTTF MTTF MTTF MTTF MTTV 582 T⬎C 583 G⬎A 608 A⬎G 611 G⬎A 618 T⬎C 622 G⬎A 1606 G⬎A Acceptor stem Acceptor stem AC stem AC wobble base AC stem Variable loop Acceptor stem MTTV MTTV MTTV MTTV 1624 C⬎T 1642 G⬎A 1644 G⬎T 1659 T⬎C DHU stem AC stem Variable region TC stem MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTL (UUR) MTTI MTTI MTTI MTTI MTTI MTTI MTTI MTTI MTTI MTTI MTTI MTTI MTTQ MTTQ 3243 A⬎G 3243 A⬎T 3249 G⬎A 3250 T⬎C 3251 A⬎G 3252 A⬎G 3254 C⬎G 3255 G⬎A 3256 C⬎T 3258 T⬎C 3260 A⬎G 3264 T⬎C 3271 T⬎C 3271 delT 3273 T⬎C 3274 A⬎G 3280 A⬎G 3287C⬎A 3288 A⬎G 3291 T⬎C 3302 A⬎G 3303 C⬎T 4267 A⬎G 4269 A⬎G 4274 T⬎C 4284 G⬎A 4285 T⬎C 4291 T⬎C 4295 A⬎G 4298 G⬎A 4300 A⬎G 4309 G⬎A 4317 A⬎G 4320 C⬎T 4332G⬎A 4336 T⬎C DHU loop DHU loop DHU loop DHU loop DHU loop DHU loop DHU stem DHU stem DHU stem AC stem AC stem AC loop AC stem AC stem AC stem AC stem TC stem TC loop TC loop TC loop AC stem Acceptor stem Acceptor stem Acceptor stem DHU stem DHU/AC stem AC stem AC loop AC stem AC stem AC stem TC stem TC loop TC stem Acceptor stem Acceptor stem MM MTTM Het 4409 T⬎C 30 DHU stem MTTM MTTW MTTW MTTW MTTW 4450 G⬎A 5521 G⬎A 5532 G⬎A 5537 InsT 5540 G⬎A TC stem DHU stem DHU stem AC stem AC stem Human Mitochondrial tRNAs Clinical features of related pathologies M.M., EI, tachycardia MELAS, EI, RP TIN MERRF MM EI Ataxia, PEM, SNHL, Sz, RP, cataracts, hypotonia Adult LS MELAS LS Movement disorder, hemiplegia MELAS, DM, SNHL PEM, MM KSS MM MM MELAS MM MERRF/KSS overlap MERRF, MELAS MM MM, HCM DM MELAS, DM PEM PEO, EI Depression, cataracts MM PEM, RP, SNHL MM MELAS MM HCM MM, ataxia, SNHL DCM, Sz, SNHL, FSGS CPEO, ALS CPEO, HCM PEO HTN, dsylipidemia HCM CPEO, MS, MM HCM CPEO DCM PEM, HCM Atypical MELAS, DM Alzheimer disease/Parkinson disease S (male) EI, muscle atrophy, hypotonia, short stature Splenic lymphoma MM MNGIE-like syndrome MILS PEM Het/Hom Score Inheritance References Het Het Homo Het Het Het Het 35 38 11 35 26 31 27 S (female) D I D D D/I I, D 112 30,57 184 94 85 33 130,178 Homo Het Het Het 27 26 21 16 I D I I 100 31 21 12 Het Het Het Het Het Het Het Het Het Het Het Het Het Het Het Het Het Het Het Het Het Het/homo Het Het Het Het Het Homo Het Het Homo Het Het Het Het Homo 40 21 26 18 25 28 13 35 40 39 40 10 27 21 35 33 40 33 21 21 40 29 30 27 37 22 26 21 21 38 34 27 33 13 30 15 I ND I I I I I I I, D I I I I D I D I ND I D I I D I ND I ND I I S (female) I ND ND ND D ND 25 150 147 49 168 111 78 115 75,110 18,163 98 166 132 152 17 73 18,163 3 54 50 11,187 48,155 172 170 13,23 26 156 195 105 29,171 20,173 43 169 134 6 39 34 Het 30 D 190 Het Het Het Het Het 27 35 34 38 32 ND I I I ND 92 157 96 137, 182 158 MUSCLE & NERVE February 2008 153 Table 2. Continued tRNA gene Nucleotide change Region Clinical features of related pathologies Het/Hom Score Inheritance References Het Het Het Het 35 23 34 33 D ND I I 3 113 167 160 Het 32 I 40,66 Het Homo 36 28 ND ND 143 27 Het Het Het Het Het 35 32 34 26 39 D ND D I I 161,163 61,192 106 38 76,95,136 Homo Het Het Het Homo 16 35 26 35 20 I S (female) I S (female) I 138 123 131 126 119 MTTW MTTW MTTA MTTA 5543 T⬎C 5549 G⬎A 5591 G⬎A 5628 T⬎C AC loop AC stem Acceptor stem AC stem MTTA 5650 G⬎A Acceptor stem MTTN MTTN 5692 T⬎C 5693 T⬎C AC loop AC loop MTTN MTTN MTTN MTTC MTTC 5698 G⬎A 5703 G⬎A 5728 T⬎C 5783 G⬎A 5814 T⬎C AC loop AC stem Acceptor stem TC stem DHU stem MTTY MTTY MTTY MTTY MTSS1 (UCN) MTSS1 (UCN) MTSS1 (UCN) MTTS1(UCN) 5843 A⬎G 5874 T⬎C 5877 C⬎T 5885 DelT 7443 A⬎G TC loop DHU stem DHU loop Acceptor stem Acceptor stem MM Dementia, chorea, DM MM, myalgia, high CK CPEO, weakness, dysphagia MM, myotonic dystrophylike phenotype CPEO PEM, hypotonia, liver dysfunction CPEO, MM CPEO, MM FSGS, GH deficiency, PEM MM, HCM, TIN, SNHL MELAS, Sz, PEO, ataxia, HCM FSGS EI, weakness CPEO CPEO SNHL 7444 G⬎A Acceptor stem SNHL Homo 20 I 119 7445 A⬎C Acceptor stem SNHL Homo 15 I 119 7445 A⬎G Acceptor stem Het/homo 32 I 127, 149 MTTS1(UCN) MTSS1 (UCN) MTSS1 (UCN) MTTS1(UCN) MTTS1(UCN) MTTS1(UCN) MTTS1 (UCN) MTTD 7472insA 7472insC Variable region Variable region SNHL, palmoplantar keratoderma Sz SNHL, MERRF, EI Homo Het 40 40 I I 72 124,177 7480A⬎G AC loop Het 35 D 9 7497G⬎A 7510 T⬎C 7511 T⬎C 7512 T⬎C DHU stem Acceptor stem Acceptor stem Acceptor stem MM, SNHL, dementia, ataxia MM, PEM, EI SNHL SNHL PEM Het/homo Het Homo Homo 29 15 40 23 I I I I 51,72 68 22,164 72 7526 A⬎G MM Het 26 S (female) 148 Sz, DD DM, SNHL, MERRF, MELAS EI MNGIE-like syndrome Movement disorder EM PEO, movement disorder MERRF HCM CPEO, MM, hypoventilation MERRF, MELAS, SNHL MERRF MM MERRF HCM CIP, CPEO Het Het 22 21 I ND 154 133 Het Het Het Het Het Het Het Het Het Het Het Het Het Nd 22 27 16 35 27 38 8 32 38 33 32 35 21 5 I D I D S (female) I I ND I I ND I I ND 46 189 198 67 179 7 175 163 99 129 163 118 104 89 MTTD MTTK 7543 A⬎G 8296 A⬎G connector region (between acceptor/D stem) AC stem Acceptor stem MTTK MTTK MTTK MTTK MTTK MTTK MTTK MTTK MTTK MTTK MTTK MTTK MTTG MTTG 8300 T⬎C 8313 G⬎A 8326 A⬎G 8328 G⬎A 8342 G⬎A 8344 A⬎G 8348 A⬎G 8355 T⬎C 8356 T⬎C 8361 G⬎A 8362 T⬎G 8363 G⬎A 9997 T⬎C 10006 A⬎G Acceptor stem DHU stem AC stem AC stem TC stem TC loop TC loop TC stem TC stem Acceptor stem Acceptor stem Acceptor stem Acceptor stem DHU loop 154 Human Mitochondrial tRNAs MUSCLE & NERVE February 2008 Table 2. Continued tRNA gene Nucleotide change Region MTTG 10010 T⬎C DHU stem MTTG MTTR MTTH 10044 A⬎G 10438 A⬎G 12147 G⬎A TC loop AC loop DHU stem MTTH 12183 G⬎A TC stem MTTH MTTS2 (AGY) MTTS2 (AGY) MTTL (CUN) MTTL(CUN) MTTL (CUN) MTTL (CUN) MTTL (CUN) MTTL (CUN) MTTL (CUN) MTTE MTTE MTTE 12192 G⬎A 12246 C⬎G Clinical features of related pathologies Het/Hom Score Inheritance References Het 40 ND 10,29,114 Het Het Het 13 21 40 I I D 135 185 103, 174 Het 30 I 28 TC stem TC loop EM, MM, polyendocrine dysfunction EM EM MELAS, MERRF, cerebral edema RP, SNHL, muscle atrophy, ataxia HCM, DCM CIP, CPEO Homo Homo 11 5 ND ND 151 89 12258 C⬎A Acceptor stem DM, SNHL Het 35 I 93, 97 12276 G⬎A 12297 T⬎C 12301 G⬎A 12311 T⬎C 12315 G⬎A 12320 A⬎G 12334 G⬎A 14687 T⬎C 14696 A⬎G 14709 T⬎C DHU stem AC loop AC loop Variable loop TC stem TC loop Acceptor stem TC loop DHU stem AC loop Het Het Het Het Het Het Het Het Het Homo 16 26 15 18 38 35 35 31 21 40 ND I S (male) ND D ND S (female) S (male) I I MTTE 14710G⬎A AC (E⬎K) Het 35 D 19 53,176 47 63 44, 77 194 191 16 185 56, 59, 102, 107 3 MTTT 15915 G⬎A AC stem Het 21 D 116,144 MTTT MTTT MTTT 15923 A⬎G 15940delT 15950 G⬎A AC loop TC loop Acceptor stem Homo Homo homo 0 0 0 I I ND 14 145 52 MTTP MTTP MTTP 15990 C⬎T 15995 G⬎A 16002 T⬎C AC (P⬎S) AC stem DHU stem CPEO DCM AISA CPEO MM, PEO MM M.M. MM, HCM, RP EM DM, SNHL, FSHD, hydrops fetalis CPEO, fatigue, weakness, EI Sz, SNHL, MR, movement disorder LIMM MM Alzheimer disease/Parkinson disease MM, CPEO Movement disorder Ptosis, CPEO Het Het Het 35 15 19 S (female) D S (male) 71 198 146 AC, anticodon; AISA, acquired idiopathic sideroblastic anemia; ALS, amyotrophic lateral sclerosis; CIP, chronic intestinal pseudoobstruction; CPEO, chronic progressive external ophthalmoplegia; D, de novo; DCM, dilated cardiomyopathy; DD, developmental delay; DHU, dihydrouridine; DM, diabetes mellitus; EI, exercise intolerance; EM, encephalomyopathy; FSGS, focal segmental glomerulosclerosis; FSHD, facioscapulohumeral muscular dystrophy; GH, deficiency growth hormone deficiency; HCM, hypertrophic cardiomyopathy; Het, heteroplasmy; Homo, homoplasmy; I, inherited; KSS, Kearns Sayre syndrome; LIMM, lethal infantile mitochondrial myopathy; LS, Leigh syndrome; MELAS, mitochondrial encephalomyopathy lactic acidosis and stroke-like episodes syndrome; MERRF, myoclonic epilepsy associated with ragged-red fibers; MILS, maternally inherited Leigh syndrome; MM, mitochondrial myopathy; MNGIE, mitochondrial neurogastrointestinal syndrome; MR, mental retardation; MS, multiple sclerosis; ND, not determined; PEM, progressive encephalomyopathy; PEO, progressive external ophthalmoplegia; RP, retinitis pigmentosa; S, sporadic; SNHL, sensorineural hearing loss; Sz, seizure disorder; TIN, tubulointerstitial nephritis. Note: tRNAs for pro, glu, ser(UCN), tyr, cys, asn, ala, and gln are transcribed from the light strand. The nucleotide changes listed in Table 2 refer to the nucleotide and position in the mitochondrial DNA sequence as shown in http://www.mitomap.org We also listed the nucleotide changes, the regions in the tRNA cloverleaf structure where these mutations were present, and the assigned clinical phenotype for these mutations (Table 2). PATHOGENIC MUTATIONS AND BENIGN VARIANTS Based on the newly adopted classification, among all the tRNA mutations analyzed in this study there was a small percentage of mutations that were classified as neutral polymorphisms (6 of 124, 4.8%), whereas the Human Mitochondrial tRNAs remainder of the mutations were classified as possibly (18 of 124, 14.5%), probably (40 of 124, 32.3%), and definitely (60 of 124, 48.4%) pathogenic (Table 3). Only the mutations with definite and probable pathogenicity (scoring 20 points or above) were further reviewed for the purpose of this study. Mitochondrial tRNAPhe Mutations. Mutations in this gene have presented with a myopathic phenotype in the affected subjects. Four heteroplasmic muta- MUSCLE & NERVE February 2008 155 Table 3. Distribution of pathogenic mutations and benign variants among mitochondrial tRNA genes. Definite (ⱖ 30) Probable (20–29) Possible (10–19) Benign variation (ⱕ 9) Total Definite ⫹ probable N (% total) Predominant Phenotype F V L(UUR) I Q M W A N C Y S(UCN) D K G R H S(AGY) L(CUN) 4 0 10 5 2 1 5 3 4 1 2 5 0 7 1 0 2 1 3 1 4 9 6 0 1 1 0 1 1 1 4 2 4 1 1 0 0 1 1 1 3 1 1 0 0 0 0 0 1 2 0 1 1 0 1 0 3 0 0 0 0 0 0 0 0 0 0 0 0 0 1 1 0 0 1 0 6 5 22 12 3 2 6 3 5 2 4 11 2 13 4 1 3 2 7 5 (83%) 4 (80%) 19 (86%) 11 (92%) 2 (67%) 2 (100%) 6 (100%) 3 (100%) 5 (100%) 2 (100%) 3 (75%) 9 (82%) 2 (100%) 11 (85%) 2 (50%) 1 (100%) 2 (67%) 1 (50%) 4 (57%) E T P Total 3 0 1 60 1 1 0 40 0 0 2 18 0 3 0 6 4 4 3 124 4 (100%) 1 (25%) 1 (33%) 100 (82%) MM Multisystemic/LS MELAS/DM/SNHL/LS CM, PEO EM, MM MM MM/MNGIE/LS/EM MM PEO/MM Multisystemic MM/PEO SNHL MM MERRF EM EM Multisystemic DM/SNHL PEO/MM/CM/AISA PEO/MM/DM/SNHL/ EM/HCM EM MM/PEO tRNA gene See Table 2 for definition of abbreviations. tions: 582 T⬎C, 583 G⬎A, 618 T⬎C, and 622 G⬎A were identified in four unrelated patients with mitochondrial myopathy112 and exercise intolerance.30 The 583 G⬎A mutation has also been reported in a patient with a MELAS-like phenotype.57 In a patient with typical clinical, histological, and biochemical features of MERRF, Mancuso et al.94 reported a novel G⬎A transition in nucleotide position 611, affecting the wobble base. The finding of this mutation revealed that MERRF syndrome is not always associated with tRNALys mutations. Mitocondrial tRNAVal Mutations. Patients carrying mutations in this gene have had either multisystemic involvement or a clinical picture consistent with Leigh syndrome. These include a 1642 G⬎A in a MELAS patient, a 1644 G⬎T in a patient with Leigh syndrome, a 1606 G⬎A in patients with multisystemic disorders, and a homoplasmic 1624 C⬎T mutation in a family with Leigh syndrome and incomplete penetrance. The G⬎A substitution at nucleotide position 1606 was found on two occasions by two groups confirming the pathogenicity of this mutation.130,178 These two patients exhibited similar clinical phenotype: hearing loss, followed by ocular, 156 Human Mitochondrial tRNAs CNS, and skeletal muscle involvement with reduced RC enzymes in skeletal muscle. McFarland et al.100 described a family with a homoplasmic 1624 C⬎T mutation that resulted in several neonatal deaths and one surviving child with Leigh syndrome. The mother was minimally symptomatic, but a severe biochemical defect was present in a deceased infant and the mother. The 1624 C⬎T change affected a highly conserved basepair at the stem region of the dihydrouridine (DHU) arm. In addition, there was a striking reduction in the steady-state mitochondrial tRNAVal in different tissues obtained from some of these patients. Northern blot analysis did not reveal evidence of unprocessed intermediates from a large polycistronic RNA unit pointing toward a fast degradation of the mutant tRNA, which would have a major impact on mitochondrial protein synthesis. The marked difference in phenotypic expression between the mother and her children could not be explained by the mtDNA mutation alone since it is homoplasmic in every tissue of all matrilineal family members. Epigenetics or nuclear-encoded elements might act as modifiers. Mitochondrial tRNALeu(UUR) Mutations. Mutations in this gene are associated with diverse phenotypes in- MUSCLE & NERVE February 2008 cluding MELAS syndrome, myopathies, maternally inherited diabetes and deafness, and sensorineural hearing loss (SNHL). More than 22 point mutations have been identified in this 77 nucleotide tRNALeu(UUR) gene. At least 19 of them are of probable or definite pathogenicity. The majority are located in the DHU and anticodon arms. Leucine is the most frequently used amino acid in human mitochondrial-encoded proteins. The usage of UUR and CUN codons is ⬃17%– 83%. The common 3243 A⬎G mutation alters a highly conserved nucleotide and this mutation is predicted to cause a pronounced change in the structure of the tRNALeu(UUR). The impact of this structural change on aminoacylation has been studied.120 Studies using cybrid cell lines that are generated by transferring mitochondria isolated from patients harboring pathogenic mutations into cultured human cells have detected decreased respiratory chain enzyme activities.141 Moreover, additional studies have revealed defects in aminoacylation,74 protein synthesis,24 RNA processing,86 and levels of tRNA modification.202 Mitochondrial processing intermediates, consisting of RNA species referred to as RNA19, could play a significant role in this syndrome, as pointed out in a recent review.139 These mitochondrial processing intermediates corresponding to the 16S ribosomal RNA (rRNA), tRNALeu(UUR), and the nicotinamide adenine dinucleotide dehydrogenase subunit 1 (ND1) genes were observed in cybrid cell lines expressing the 3243 A⬎G.82 Since 16S rRNA is contained in precursor RNA19, this transcript could be incorporated into the ribosomes, leading to ribosome stalling. The percentage corresponding to the 3243 A⬎G mutation in the processing intermediates is higher than the percentage observed in mtDNA, suggesting that the processing intermediates that contain mutations may be more difficult to process than their cellular counterparts.86,87 In addition, this tRNA molecule has unstable structural domains, and one of the effects of the 3243 A⬎G mutation might be the formation of a dimeric complex that interferes with the function of the tRNA within the context of protein-RNA assembly.196 Since the DHU stem of human mitochondrial tRNALeu(UUR) contains only two Watson–Crick pairs, the dimerization of this tRNA caused by this mutation may lead to its disruption. The observed reduction in complex I associated with MELAS syndrome could be linked to a skewing of the tRNALeu anticodon availability. The mitochondrial tRNALeu(UUR) gene harboring the common MELAS mutation lacks the normal taurine modifica- Human Mitochondrial tRNAs tion (5-taurinomethyluridine) at the anticodon wobble position, substantially decreasing UUG(Leu)– dependent protein synthesis.83 This mechanism could explain the reduced translation of UUG-rich genes such as ND6. Most other mitochondrial transcripts preferentially use the UUA(Leu) codon, hence their translation is not substantially affected.84 Deficient wobble modification may be one of the many factors responsible for the phenotypic features of MELAS syndrome. The morbidity and mortality of stroke-like episodes observed in MELAS syndrome could also be associated with the presence of vessels strongly positive for succinate dehydrogenase (SSVs). These SSVs positive for cytochrome c oxidase activity may play an important pathogenic role.62,117 Although there is decreased cytochrome c oxidase activity for each individual mitochondrion, there is a high increase in mitochondrial mass in these SSVs that leads to an increase in total cytochrome c oxidase activity. Nitric oxide (NO) binds the active sites of cytochrome c oxidase and displaces the heme-bound oxygen.193 This could lead to oxygen depletion in the surrounding tissue and decreased unbound NO, with an ensuing microvascular stroke-like pattern. Thus, the stroke-like episodes observed in MELAS syndrome could be caused by alterations in NO homeostasis resulting in microvascular damage. Another common mutation in tRNALeu(UUR) is the 3271 T⬎C mutation at the anticodon stem region, which is also associated with MELAS syndrome and diabetes mellitus. In transmitochondrial cybrid lines generated with this mutation, a lack of posttranscriptional modification,202 decreased steadystate levels,202 and abnormal intermediates of protein synthesis64 have been observed. The anticodon stem of the tRNALeu(UUR) is a loosely structured unit formed by four AU basepairs within the wildtype tRNA.197 The high AU content of the anticodon stem causes weakness in this domain and susceptibility to rupture when mismatch mutations such as the 3271 T⬎C is present. Chemical studies revealed that the anticodon stem was denatured in the mutant.197 This mutation caused an in vitro aminoacylation defect connected to Km abnormalities.197 Mitochondrial tRNAIle Mutations. The mitochondrial tRNAIle gene is the third most commonly mutated among the mitochondrial tRNA genes. Among the 12 reported mutations, 11 of them score above 20. The predominant clinical phenotypes associated with mutations in this mitochondrial tRNA are isolated cardiomyopathy (4295 A⬎G, and 4300 A⬎G),20,105 and progressive external ophthalmople- MUSCLE & NERVE February 2008 157 gia (4274 T⬎C, 4285 T⬎C, 4298 G⬎A, and 4309 G⬎A).13,43,156,171 Other phenotypes such as combined cardiomyopathy and PEO (4284 G⬎A),26 multisystemic (4269 A⬎G),170 encephalomyopathic (4267 A⬎G),172 and hypertension and dyslipidemia (4291 T⬎C)195 have been described. Studies of disease-related tRNAIle mutations using cell lines revealed changes in tRNA stability in the presence of the 4269 A⬎G mutation that affects an AU pair in the acceptor stem of the human mitochondrial tRNA.Ile An accelerated tRNA degradation related to the loss of structural integrity caused by the mutation was observed.201 The 4285 T⬎C and the 4298 G⬎A mutations (located in the anticodon stem) diminished the efficiency of aminoacylation by as much as 1,000fold.79 It was also found that mutant tRNAs were able to inhibit the charging of the wildtype tRNAIle80. Two studies have revealed that the cardiomyopathy-associated mutation 4317 A⬎G induces an aberrant secondary structure in the T-stem due to the instability introduced by the native CA pair.32,180 This aberrant structure may decrease aminoacylation. It seems that for this class of mutations in this particular tRNA, a weak structure may amplify the effect of the mutation. Mitochondrial tRNAGln Mutations. Mutations in this gene have been found in patients with either an encephalomyopathic or myopathic presentation. A heteroplasmic G⬎A transition at nucleotide 4332 in the tRNAGln gene was identified in a subject presenting with stroke-like episodes, deafness, leukoencephalopathy, COX-negative ragged-red fibers, and basal ganglia calcifications with late-onset findings that were atypical for MELAS syndrome.6 Dey et al.34 reported a 12-year-old boy with a myopathy in association with a single nucleotide insertion, 4370insA (T⬎AT), in the mitochondrial tRNAGln gene. The muscle biopsy was remarkable, a majority of the fibers were COX-deficient, and there were decreased steady-state levels of subunit II of COX and subunit 1 of complex I. Addition of the extra nucleotide in the anticodon loop of this gene changes seven evolutionary-conserved bases in the anticodon loop to eight. In many cases, tRNAs with eight or nine base anticodon loops have been associated with frameshifting.183 Occasional frameshift during translation may lead to a defect in mitochondrial protein synthesis or misincorporation of amino acids. Mitochondrial tRNAMet Mutations. A 4409 T⬎C mutation was found in a 30-year-old woman with exer- 158 Human Mitochondrial tRNAs cise intolerance and dystrophic changes on skeletal muscle.190 Because of the role of tRNAMet in translation initiation, the high level of the mutation observed in this patient’s skeletal muscle is likely to cause a deficiency of mtDNA-encoded subunits, potentially affecting the capacity for ATP production. In addition, a computerized analysis revealed a markedly altered structure induced by this mutation.190 Another mutation, a G⬎A transition at position 4450 in this gene, was initially described by Lombes et al.92 in a patient with splenic lymphoma. The patient’s lymphocytes revealed abnormal mitochondria and multiple RC defects. Transfer of the mutant mitochondria to Rho0 cells demonstrated a severe respiratory defect. However the link of this mutation to the underlying disease remains unclear. Mitochondrial tRNATrp Mutations. Six pathogenic mutations have been reported in this tRNA gene and cause a diverse clinical picture. A novel pathogenic mutation 5521 G⬎A was found to be associated with late-onset mitochondrial myopathy.157 Histochemical and biochemical studies documented COX-negative ragged-red fibers. This mutation located in the D-stem could affect the tRNA-ribosome interaction, resulting in reduced mitochondrial protein synthesis. Alternatively, since this mutation is located close to the origin of mtDNA light-strand replication, it could interfere with the regulation of the mtDNA light strand, altering the L-strand secondary structure formed after the H-strand displacement. The combination of Leigh syndrome, muscle COX deficiency, complex I deficiency, and a heteroplasmic 5537 insertion T has been described on two occasions in different families by two independent groups.137,182 This mutation affects the structure and stability of the anticodon stem region. Insertions in the tRNA genes are a rare cause of mitochondrial disorders, but these cases suggested that this mutation should be screened in cases of Leigh syndrome and COX deficiency when mutations in SURF1 have been excluded. Other heteroplasmic point mutations in this tRNA gene includes 5532 G⬎A in a patient with a neurogastrointestinal syndrome,96 a novel pathogenic 5540 G⬎A transition in a patient with sporadic encephalomyopathy characterized by spinocerebellar ataxia,158 a 5543 T⬎C causing mitochondrial myopathy, and a 5549 G⬎A mutation associated with dementia, chorea, and diabetes. Complex IV deficiency may be the most common RC defect found in patients affected with mutations in the mitochondrial tRNATrp gene. This finding may be explained MUSCLE & NERVE February 2008 by the higher relative content of tryptophan in COX subunits COI and COIII, 3.1% and 4.6%, respectively, compared to an average of 2.5% in mtDNA encoded complex I subunits.157 Mitochondrial tRNAAla. Mutations in this gene have been associated with a myopathic phenotype. The T⬎C transition at nucleotide position 5628 was identified in a patient with late-onset chronic progressive external ophthalmoplegia (CPEO), dysphagia, and mild proximal myopathy.160 Two mutations were found at the amino acid acceptor stem region: the 5591 G⬎A and the 5650 G⬎A; both cause mitochondrial myopathy.66,40,167 Both mutations create a mismatched basepair that can be designated as G4:U69 and U6:G67 for 5650 G⬎A and 5591 G⬎A, respectively, according to the canonical structure. These mismatches lie next to a G5:U68 wobble in the amino acid acceptor stem of the tRNAAla molecule. One role of the G5:U68 pair is to provide the tRNA conformation that is most favorable for binding and positioning the tRNA acceptor end at the enzyme catalytic site.45 The gain of an additional wobble may distort the conformation of the tRNA and render that structure unsuitable for aminoacylation. Mitochondrial tRNAAsn Mutations. Mutations in this gene have been associated either with a CPEO phenotype or with a myopathic phenotype. The first mutation associated with an A to G substitution in nucleotide position 5692143 was found in a patient presenting with CPEO. The mutation could cause a possible reorganization of the anticodon loop leading to inhibition of assembly of the ribosome-mRNAtRNA complex required for translation. A de novo 5698 G⬎A mutation has been described in two patients presenting with a phenotype consisting of isolated mitochondrial myopathy with chronic progressive ophthalmoplegia.163,161 Two patients with early onset of progressive external ophthalmoplegia and fatigability were described to have the 5703 G⬎A mutation in the tRNAAsn.61,192 Studies using transmitochondrial cybrid cell lines containing this mutation revealed impaired oxidative phosphorylation and protein synthesis with severe reduction in steady-state levels of the tRNAAsn pool.61 This was accompanied by a conformational change in the tRNAAsn that may impair aminoacylation. In contrast, the patient described by Coulbault et al.27 presented with an encephalomyopathy and did not have features of CPEO. A T⬎C substitution at position 5693 in the tRNAAsn gene was found in blood and muscle. It was suggested that this muta- Human Mitochondrial tRNAs tion could affect the anticodon loop structure of the tRNAAsn since it is located next to the anticodon GUU. Another mutation at nucleotide position 5728 in the same tRNA gene was reported as the cause of multiorgan failure.106 Mitochondrial tRNACys Mutations. The 5783 G⬎A was found in a patient with myopathy, cardiomyopathy, renal failure, and sensorineural hearing loss (SNHL).38 Another pathogenic mutation is the 5814 T⬎C mutation reported independently by three groups to be associated with MELAS syndrome and with progressive external ophthalmoplegia in the first two cases.76,95,136 This mutation shares similar features with the 3243 A⬎G mutation in the tRNALeu(UUR)76 in that both mutations are located at the D-stem/D-loop boundary (positions 14 and 13 at the cloverleaf). These nucleotides seem to provide stability to the tertiary structure of mitochondrial tRNAs and regulate the efficiency of aminoacylation.55 Mitochondrial tRNATyr. Patients with mutations in this gene predominantly exhibit a myopathic phenotype, but more cases are required in order to establish a genotype–phenotype correlation. The first reported pathogenic mutation in this gene123 was a somatic 5874 T⬎C transition in a woman with exercise intolerance and bilateral ptosis. The transition changed a highly evolutionarily conserved nucleotide and may alter the transcriptional processing or translation of mitochondrial proteins. This mutation may preferentially affect complex III and IV. This finding could be explained by the fact that the percentage of tyrosine residues is among the highest in cytochrome b, COXI, and COX III subunits. Two additional cases with CPEO, myopathy, and exercise intolerance associated with mutations in this tRNA gene were 5877 C⬎T and 5885delT.126,131 The 5885delT shortened the aminoacyl-acceptor stem.126 Since this 7-basepair stem is strictly conserved among mammalian mitochondrial tRNATyr and is a canonical feature of tRNA in general, a deletion may impair the formation of a proper secondary and tertiary structure. The 5877 C⬎T mutation occurs at the DHU loop of this gene.131 This mutation seems to alter an essential residue for the configuration of the L-shaped tertiary structure of tRNA. Studies using transmitochondrial cybrids revealed decreased oxygen consumption and compromised cell viability. MUSCLE & NERVE February 2008 159 Mitochondrial tRNASer(UCN). Patients carrying mutations in this gene usually present with SNHL. The mutations associated with SNHL include the 7443 A⬎G, 7444 G⬎A, 7445 A⬎G, 7472 insC, the 7480 A⬎G, and the T to C changes at nucleotide positions 7510, 7511, and 7512. The pathogenic mutations located at the boundary between the termination codon of COI and the 3⬘ end of the tRNASer(UCN) gene (7443 A⬎G, 7444 G⬎A, and 7445 A⬎G) affect the processing and stabilization of the precursor tRNASer(UCN) rather than affecting the termination codon of CO1.199 These mutations affect the secondary structure at the 3⬘ tRNA cleavage site for the processing by tRNaseZL, leading to decreased steadystate levels of tRNA.90,200 The 7472 insertion C mutation was found in a family who exhibited SNHL and a MERRF-like phenotype177 and in another patient with isolated myopathy, expanding the clinical phenotype of the mutation.124 The insertion adds a seventh cytosine to a six-cytosine run in the T⌿C loop, potentially altering the secondary structure of the mitochondrial tRNASer(UCN) gene. Analysis of cybrids containing the 7472 insertion C revealed a decrease in the abundance of this particular tRNA, suggesting that it could be the result of altered processing.181 In addition, the 7480 A⬎G mutation was described in a woman who presented with a progressive mitochondrial myopathy and later developed SNHL, dementia, and ataxia.9 This mutation could lengthen the anticodon stem by additional pairing, drastically reducing the size of the anticodon loop, and compromising anticodon function by the disruption of codon matching. These SNHL-related tRNASer(UCN) mutations occurred multiple times on different genetic backgrounds at either heteroplasmic or homoplasmic states, and the clinical severity does not appear to correlate with the proportion of mutant heteroplasmy.22,68,69,186,188 In addition to hearing impairment, there are inter- and intrafamilial differences in the penetrance of the neurological dysfunction. These results and the lack of correlation between the mutant load and the phenotype suggest that nuclear genes may play a role in the clinical expression of these mutations. Furthermore, the reason that these mutations preferentially affect the function of the inner ear is currently not known. Interestingly, a heteroplasmic mutation in this same tRNA, 7497 G⬎A, does not cause SNHL but rather a mitochondrial myopathy.51,72 This mutation has been reported in a 13-year-old girl with exercise intolerance and lactic acidosis, and in another two unrelated families with progressive myopathy, 160 Human Mitochondrial tRNAs ragged-red fibers, lactic acidosis, and respiratory chain complexes deficiency.51,72 tRNAAsp Mutations. Two probable pathogenic mutations in the tRNAAsp gene have been reported. The first heteroplasmic pathogenic mutation in this gene was reported154 in a patient with myoclonic epilepsy and psychomotor regression and other affected family members. The mutation abolished a highly conserved pairing in the anticodon domain of the tRNAAsp gene that could affect the double helix, resulting in an alteration of the secondary structure of this tRNA. In addition, another pathogenic mutation in this gene was reported148 in a young girl who presented with isolated progressive exercise intolerance. Mitochondrial The tRNALys is an example of an oversimplified structure with a small DHU loop. The small DHU loop may limit the number of contacts necessary to stabilize the tertiary fold and the intrinsically weak structure of the human mitochondrial tRNALys depends on the existence of modified bases to maintain tRNA integrity. The existence of posttranscriptional modifications plays an important role in stabilizing the structure of mitochondrial tRNALys. Proper folding and aminoacylation require a 1-methyladenine residue at nucleotide 9 to prevent formation of an alternative structure with an extended acceptor stem.65 The 8344 A⬎G mutation located in this gene is present in 80%–90% of patients with myoclonic epilepsy associated with ragged-red fibers (MERRF).36 Cybrid cells carrying this mutation demonstrated reduced protein synthesis as a consequence of diminished tRNA stability and concomitant reduction in the aminoacylation level for the mutant tRNA.Lys37 It has also been found that the levels of posttranscriptional modification in the wobble position of the anticodon tRNALys were decreased, leading to disruptive translation.203 The disruption in translation was related to the defective binding of the mutant 8344 A⬎G to the mRNA-ribosome complex and defective translation of both AAA and AAG codons due to weak codon–anticodon interaction in this mutant tRNA. Further work revealed lack of taurine modification at the wobble uridine of mitochondrial tRNALys from pathogenic cells of MERRF patients carrying 8344 A⬎G mutation.165 The 2-thio group of the wobble uridine is involved in mediating a stable codon–anticodon interaction,4 hence the loss of the 2-thio group associated with this mutation is thought Mitochondrial tRNALys Mutations. MUSCLE & NERVE February 2008 to be one of the main reasons for the weak binding to cognate codons. Two other pathogenic mutations in this same gene, 8356 T⬎C and 8363 G⬎A, seem to account for most of the remaining MERRF cases36 and more recently the 8361 G⬎A has been added to the expanding mutational spectrum of MERRF.129 However, the clinical phenotype associated with mutations in this gene can be broad. The 8361 G⬎A mutation has been associated with cardiomyopathy129 and other mutations in the same gene are associated with exercise intolerance (8300 T⬎C),46 atypical MELAS (8296 A⬎G),133 and a phenotype evocative of mitochondrial neurogastrointestinal encephalomyopathy (MNGIE) syndrome (8313 G⬎A).189 In addition, other clinical presentations include encephalomyopathy (8328 G⬎A),67 myopathy (8362 T⬎G),163 PEO and myopathy (8355 T⬎C),163 and PEO with movement disorder (8342 G⬎A).179 The potential pathomechanistic effects of the 8344 A⬎G mutation were studied in vitro and the mutant transmitochondrial cybrids exhibited poor respiration and defective mitochondrial protein synthesis and respiratory chain enzyme activity in addition to a marked decrease in tRNALys steadystate levels and aminoacylation.5 Mitochondrial tRNAGly Mutations. Two pathogenic mutations in this tRNA gene have been reported. The 9997 T⬎C mutation caused hypertrophic cardiomyopathy.104 The 10010 T⬎C mutation has been associated with diverse clinical phenotypes that include an encephalomyopathic presentation with seizures, choreoathetosis, ataxia, and spastic tetraparesis10; a myopathic presentation with fatigue and elevated serum creatine kinase114; and a multisystemic presentation with exercise intolerance, polyneuropathy, and hypothyroidism.29 This mutation is located within a highly conserved stem region of the DHU arm, which underlies its pathogenicity. Mitochondrial tRNAArg Mutations. The heteroplasmic 10438 A⬎G in the tRNAArg gene was found in a boy affected with mild cognitive impairment, ataxia, nystagmus, and muscle weakness.185 This mutation changed the nucleotide flanking the anticodon. Point mutations flanking the anticodon triplet may affect codon matching and potentially tRNA synthetase recognition, ultimately leading to a decrease in the accuracy and rate of translation.204 The position 10438 in mitochondrial DNA corresponds to position 37 in the tRNA molecule. Through sequencing of human mitochondrial tRNAs, it is known that this nucleotide undergoes posttranscrip- Human Mitochondrial tRNAs tional modification, which suggests functional relevance.15 Mitochondrial tRNAHis. The heteroplasmic 12147 G⬎A mutation was found in a patient with a MELAS phenotype that eventually evolved into a MELAS/ MERRF phenotype.103 The change most likely would perturb secondary and tertiary structure.103 As previously described,60 mutations in the dihydrouridine arm might cause mitochondrial dysfunction by impairing mitochondrial protein synthesis. This may be pertinent for COIII, the richest polypeptide in histidine residues,103 which may account for the associated COX deficiency. Another pathogenic heteroplasmic mutation in tRNAHis is the 12183 G⬎A at the T stem region identified in a patient with pigmentary retinopathy, SNHL, muscle hypotrophy, and ataxia.28 Mitochondrial tRNASer(AGY) Mutations. Two families with novel 12258 C⬎A mutation in the tRNASer(AGY) gene have been reported. In the first case the patient had a history of cataracts, SNHL, cerebellar ataxia, and diabetes.93 The second time the mutation was reported in a large pedigree of five generations with 24 affected matrilineal family members presenting with retinitis pigmentosa and progressive SNHL.97 The 12258 C⬎A substitution altered a highly conserved basepair in the amino acid acceptor stem that potentially could affect the secondary and tertiary structures and therefore the function of this tRNA.162 It is interesting to note that the only reported pathogenic mutation in the tRNASer(AGY) gene, 12258 C⬎A change, causes SNHL, similar to mutations in tRNASer(UCN). It is not clear why mutations in the tRNASer genes predominantly cause SNHL. tRNALeu(CUN) Mutations. The myopathic phenotype is predominantly associated with the mutations in this tRNALeu(CUN). The 12297 T⬎C mutation, located right next to the anticodon, caused dilated cardiomyopathy. The heteroplasmic 12315 G⬎A mutation was diagnosed in a young woman affected with exercise intolerance, bilateral ptosis, and PEO.77 The mutation disrupts a highly conserved G-C basepair in the T⌿C stem of the molecule. This mutation had been previously reported in a middle-aged man who had developed ptosis, CPEO, weakness, retinitis pigmentosa, hearing loss, and peripheral neuropathy.44 Another heteroplasmic mutation 12320 A⬎G affecting the T⌿C loop was identified in a patient with an isolated skeletal myopathy.194 In vivo, this mutaMitochondrial MUSCLE & NERVE February 2008 161 tion increased over time from 70%–90%. Concurrently, the patient’s myopathy progressed from mild to profound. The somatic 12334 G⬎A mutation affecting the acceptor stem was identified in a subject who exhibited an isolated skeletal myopathy.191 The patient presented with exercise intolerance, ragged-red fibers, and deficiencies of complex III and IV in skeletal muscle. Mitochondrial tRNAGlu Mutations. A 14687 T⬎C somatic mutation affecting the T⌿C loop of the mitochondrial tRNAGlu was associated with mitochondrial myopathy, respiratory failure, COX-negative ragged-red fibers, and complex I and IV deficiencies.16 The 14696 A⬎G mutation was detected in a patient with psychomotor retardation, hypotonia, dysphasia, combined complex I and III deficiencies, and ragged-red fibers. This mutation changes a nucleotide in the DHU stem, creating a novel basepair and reducing the wobble.185 The heteroplasmic 14709 T⬎C mutation in the tRNAGlu gene has been reported several times, presenting with a very heterogeneous clinical phenotype including diabetes, myopathy, ataxia, congenital encephalopathy, and a presentation suggestive of facioscapulohumeral muscular dystrophy.56,59,102 Some of the clinical diversity could be accounted for by the intratissue variability of the mutant load, but the involvement of nuclear factors cannot be discounted. It has been documented that the mutation has attained homoplasmy within a family and in a variety of tissues.102 Cybrid studies revealed that this mutation alone was not sufficient to lead to impairment of mitochondrial function in these homoplasmic cybrid clones.121 Reduced levels of mutant tRNAs were observed. This nucleotide located immediately next to the anticodon may be important to maintain the stability of the tRNA for codon-binding. The 14710 G⬎A mutation3 was associated with a phenotype consisting of retinopathy, PEO, ptosis, and mitochondrial myopathy. This mutation resulted in an anticodon swap from Glu (CUU) to Lys (UUU) in this particular gene, potentially having an effect on tRNA aminoacylation. This is one of the few mitochondrial tRNA mutations that changes an anticodon amino acid. The other one is the 15990 C⬎T in the tRNAPro gene, that changes to serine (see below).109 Mitochondrial tRNAThr Mutations. The only mutation in the mitochondrial tRNAThr gene that meets crite- 162 Human Mitochondrial tRNAs ria for probable pathogenicity is the 15915 G⬎A. This mutation was found in a patient with mitochondrial encephalomyopathy, myoclonus, ragged-red fibers, and progressive calcification in the basal ganglia and cerebral atrophy.116,144 Mitochondrial tRNAPro Mutations. The 15990 C⬎T mutation in the tRNAPro gene has been associated with myopathy and CPEO.109 This mutation would convert the proline anticodon into a serine anticodon resulting in altered charging properties and impairment of aminoacylation. The tRNA anticodon swap could affect protein synthesis by enhancing misprocessing of polycistronic primary transcripts, amino acid misincorporation, and reduced or defective charging of the mitochondrial tRNAPro gene. This mutation could also have effects on the posttranscriptional modification of other nucleotides and potentially lead to hypomethylation or no methylation in nucleotide 37 in the mitochondrial tRNAPro gene.15 This hypomodification may attenuate the severity of disease and act as a negative determinant interfering with the recognition of a tRNA by a noncognate aminoacyl-tRNA synthetase,125 leading to mischarging of the mutant tRNA instead of a complete lack of charging. ASCERTAINMENT BIAS Mitochondrial tRNA mutations have been traditionally characterized by the focal accumulation of large numbers of morphologically and biochemically abnormal mitochondrial that present as ragged-red fibers on skeletal muscle histology.153 This finding may have led to an ascertainment bias since screening for mutations in mitochondrial tRNA genes is usually done when ragged-red fibers are found, perhaps contributing to the disproportionately large number of reported mitochondrial tRNA mutations. PATHOGENICITY AND LOCATION OF MUTATIONS IN THE THREE-LEAVES CLOVER STRUCTURE OF tRNA The different distribution patterns of pathogenic and neutral variants in the tRNA structure may prove useful in devising an algorithm to predict the pathogenic effects of novel mitochondrial tRNA changes. After reviewing and scoring the published mutations for pathogenicity based on our scoring system, it is apparent that a significant percentage of mutations are not supported by sufficient evidence to be considered pathogenic. These are variations that score below 20 points and are considered to be of benign or possible pathogenicity. Of the listed mutations on MITOMAP and PubMed, 4.8% are con- MUSCLE & NERVE February 2008 sidered neutral variants. If we add the mutations with possible pathogenicity (14.5%), it is found that about 19% of the surveyed mutations either constitute benign variants or have weak evidence for pathogenicity (Tables 3, 4). Some of these published mutations were reported before the canonical criteria35 were established. It is interesting to note that in the article published by McFarland et al.101 26% of the reviewed mutations listed as pathogenic on MITOMAP did not have enough evidence to be considered pathogenic. The same group also stated that only 16% of the analyzed mutations were pathogenic beyond doubt,101 in contrast to our review, where around 48% of the analyzed mutations have definite pathogenicity (Tables 3, 4). The difference in results could be accounted by the different scoring criteria used in both studies and by the fact that their analysis only included mutations listed on MITOMAP as pathogenic as of December 1, 2003, whereas our review includes mutations listed on the same database as pathogenic as of January 31, 2007. In addition, mutations retrieved from PubMed and not contained in MITOMAP were also included. The reports of newly described mutations have included a more careful functional analysis of their pathogenicity. A small number of common mutations are responsible for the majority of mitochondrial disorders due to mitochondrial DNA mutations. Three mitochondrial tRNAs (tRNAIle, tRNALeu(UUR), and tRNALys) concentrate almost half (41%) of the known mutations associated with definite and probable pathogenicity (Table 3). This finding has been noted before,58 and the percentage found in our review is in agreement with previous reviews196 where mutations in those three genes accounted for almost half of the mutations deemed to be associated with pathogenicity. Among the three genes the tRNALeu(UUR) gene is a hotspot for pathogenic mutations consistent with the findings of a previous study.141 Moreover, we observed the highest sequence variability (22 variants) and pathogenic mutation (definite plus probable) to nonpathogenic variant (possible plus unlikely) ratio (19:3 ⫽ 6.3) occurred in the mitochondrial tRNALeu(UUR) gene. However, these observations may also reflect an ascertainment bias since, in some studies of mitochondrial patients with defined clinical syndromes (MERRF, MELAS, mitochondrial myopathy, PEO, and Leigh syndrome), only the mitochondrial tRNALeu(UUR) and tRNALys genes were targeted to look for known mutations previously associated with these particular clinical phenotypes, thus ignoring the potential contribution of other mitochondrial tRNA genes to these phenotypes. The distribution of pathogenic mutations encompasses all of the 22 mitochondrial tRNA genes (http://www.mitomap/org) (Table 3). Our review revealed that although the distribution of mitochondrial tRNA mutations includes any domain of the cloverleaf structure, most of the pathogenic mutations occur in stem structures since among the 100 mutations (definite plus probable) found in mitochondrial tRNA genes, 68% occurred in stem structures. There was also a predominance of possible pathogenic mutations in stem structures (12 of 18; 66.6%). In the case of benign polymorphisms, mutations in loop structures (5 of 6; 83.3%) predominated over mutations in stem regions (1 of 6; 16.6%). McFarland et al.101 also commented on the fact that the majority of pathogenic mutations in tRNAs are located in stem structures (73%), with mutation hotspots in the anticodon and the aminoacyl acceptor stems.101 According to our review, 45 definite and probable mutations (45 of 100; 45%) occur in Table 4. Distributions of tRNA mutations according to affected region of cloverleaf structure. Acceptor stem AC stem AC loop TC stem TC loop DHU stem DHU loop Variable Connector region (Acceptor/DHU stem) DHU/AC stem Total Definite (ⱖ 30) Probable (20–29) Possible (10–19) Benign (ⱕ 9) Total 15 12 9 5 5 10 1 3 0 0 60 9 9 4 4 2 4 5 1 1 1 40 3 3 2 3 2 3 1 1 0 0 18 1 0 1 0 3 0 1 0 0 0 6 28 24 16 12 12 176 8 5 1 1 124 See Table 2 for abbreviations. Human Mitochondrial tRNAs MUSCLE & NERVE February 2008 163 the acceptor and anticodon stems, edging closer to 50% of the total number of pathogenic mutations. Alterations in these regions may exert significant impact on codon binding and amino acid charging thus impairing mitochondrial protein synthesis. Few mitochondrial tRNA genes harbor mutations in the loop structures. However, an exception occurs in the mitochondrial tRNALeu(UUR) gene, where five pathogenic mutations (one definite and four probable) are located in the dihydrouridine loop (DHU loop) (see Table 2). Of interest, the structure of this particular mitochondrial tRNA gene might explain why mutations occurring in the loop may significantly affect the function. The tertiary structure of this mitochondrial tRNA gene is different from the structure of the cytosolic tRNAs and this difference is achieved by a process of posttranscriptional modifications.159 This tRNA has the largest DHU loop and a large T⌿C loop. An alteration in the base composition of these large loops may render this structure vulnerable by altering the tertiary hydrogen bonds in these loops that are critical in maintaining the stability of this tRNA molecule. Structural anomalies caused by the 3243 A⬎G mutation were noted in the folding of the DHU-arm of the mitochondrial tRNALeu(UUR), but structural anomalies were not observed when the mutation was in the stem structure as in the case of the 3271 T⬎C.159 Another example of mutations that occur in large loop structures is the 8344 A⬎G mutation in the tRNALys gene. This mitochondrial tRNA has the largest T⌿C loop, and a mutation in this structure may also affect the integrity of the tRNA. Thus, when pathogenic mutations occur in loop structures, they tend to occur in loops that are unusual in size and influence the tertiary folding or function of the tRNA. It is interesting to note that substitutions in the anticodon triplet are extremely rare. Only three pathogenic mutations, the 611 G⬎A in the tRNAPhe, the 14710 G⬎A in the tRNAGlu, and the 15990 C⬎T in the tRNAPro have been determined to affect any of the three bases of the anticodon.3,94,109 The 611 G⬎A involves a wobble base change and it does not change the amino acid to be charged. The other two anticodon mutations indeed alter the amino acid to be charged as specified by the tRNA. Due to the fundamental role played by the anticodon in selecting tRNA identity, a high frequency of mutations is not expected in these preserved residues, as they would lead to lethal cellular consequences incompatible with life. Although most of the pathogenic mitochondrial tRNA mutations listed in Table 3 (definite plus probable) were found in highly conserved sites, consis- 164 Human Mitochondrial tRNAs tent with previous studies,41,142,163 the known pathogenic 8344 A⬎G transition in the mitochondrial tRNALys gene affects a nonconserved nucleotide. This implies that the presence of a variant at a conserved site is not an absolute requirement to predict its pathogenicity.41 At the level of the primary sequence the changes introduced by the mutations in mitochondrial tRNA genes are known to be mainly transitions (replacement of a purine by a purine or a pyrimidine by a pyrimidine) rather than transversions (replacement of a purine by a pyrimidine or vice versa). Upon the analysis of our data we found that an overwhelming majority of pathogenic mutations were transitions (89%), with the remainder transversions (5%), and deletions/insertions (6%). These findings support the notion that transversions may be extremely detrimental from a cellular standpoint and that “milder” changes are better tolerated.41 It is plausible to speculate that it might be easier to achieve a transition than a transversion due to the structural similarity. The predominance of chemically “mild” mutations is a concept that was first introduced by Florentz and Sissler.41 In addition, among the transitions there is a predominance of AG changes (57 of 89; 64%) when compared to TC changes (36%). One of the most common oxidation products of mtDNA, 8-oxoguanine, has been found at high levels in human cells under conditions of oxidative stress,2 suggesting a possible role of oxidative stress in mitochondrial DNA damage and generation of mutations. However, it is not known whether this could be one of the mechanisms responsible for the predominance of AG changes. Although most of the mutations analyzed in this review appeared to be inherited, eight patients were found to exhibit somatic mutations that probably arose during early embryonic development. In this small cohort there was a predominance of females (8 of 11; 72.7%). However, more patients with somatic mutations in mitochondrial tRNA genes will need to be analyzed in order to know whether there is a clear gender trend due to a biological mechanism that is still elusive to us. HOMOPLASMY OF PATHOGENIC MUTATIONS Among the 100 pathogenic mutations described in this review, 11 occurred in the homoplasmic state, in which clinical expression and disease severity do not correlate with mutant load in affected tissues. Apparently, nuclear genes play a role in the disease expression of these homoplasmic mutations. Homoplasmic pathogenic mutations have also been reported in MUSCLE & NERVE February 2008 mitochondrial genes encoding for mRNAs. The increasing frequency of reports of homoplasmic mutations of definite and probable pathogenicity weakens a rigid concept of heteroplasmy as sine qua non of pathogenicity, requiring a more comprehensive functional approach to clearly determine pathogenicity in these cases. CORRELATION BETWEEN CLINICAL PHENOTYPE AND NATURE OF tRNA MUTATION There is a wide range of clinical manifestations that are characteristic of disorders caused by mtDNA mutations, and in particular of patients harboring mutations in the mitochondrial tRNA genes. As described earlier, there appears to be little correlation between the clinical manifestations observed in a particular patient and the specific mitochondrial tRNA harboring that mutation. However, mutations at the same position in the cloverleaf structure of different tRNA genes may be associated with similar clinical phenotypes.142 Mutations at nucleotide position 4284 in the tRNAIle and at nucleotide position 5703 in the tRNAAsn, both at position 27 of the tRNA structure, are associated with PEO. In addition, mutations at nucleotide position 3303 of the tRNALLys eu(UUR) and nucleotide position 8363 of the tRNA , both at position 72, are associated with cardiomyopathy. Similarly, mutations at nucleotide position 4269 of the tRNAIle and at nucleotide position 9997 of the tRNAGly, both at position 7, are associated with cardiomyopathy. Encephalomyopathy has been observed in association with mutations at nucleotide positions 3271 of the tRNALeu(UUR) and 5549 of the tRNATrp, which correspond to position 39. Lastly, multisystemic involvement was observed with mutations at nucleotide positions 1606 of tRNAVal and nucleotide position 7512 of tRNASer(UCN) that corresponded to position 5 of the cloverleaf structure. These findings may add further evidence to support the hypothesis that the genotype–phenotype correlation for tRNA mutations may be based on the disruption of secondary and tertiary structures. tRNA STRUCTURES ARE FRAGILE, SUSCEPTIBLE TO DETRIMENTAL EFFECTS OF NUCLEOTIDE SUBSTITUTIONS In a previous review of mutations in mitochondrial DNA encoded mRNA genes, 74 mutations scored as probable or definite pathogenic in a total of 11,340 basepairs of 13 mRNA gene regions.199 In the current review, 90 pathogenic mutations were found in a total of 1,505 basepairs coding for tRNA genes. These data translate into 6.5 and 60 Human Mitochondrial tRNAs mutations per kb for mRNA and tRNA genes, respectively. There is almost 10-fold higher frequency in generating pathogenic mutations in tRNA genes than in mRNA genes. If it is assumed that the mutation events occur randomly regardless of the nucleotide location, these results suggest that nucleotide substitutions in tRNA genes are more detrimental from a structural and functional standpoint than mutations in mRNA genes. Nucleotide substitutions in tRNA genes may affect their fragile structure, unlike nucleotide substitutions in mRNA genes that do not lead to a disruption in mitochondrial protein synthesis. CONCLUSIONS Further analysis of an expanded series of tRNA mutants will be required in order to generate a consensus or framework that would allow the description of the cellular and phenotypic effects of disease-related tRNA mutations. It is important to consider the effects of pathogenic mutations on the structure of mitochondrial tRNA genes because the structural distortions probably underlie the biochemical and clinical phenotypes observed in these patients. The human mitochondrial tRNAs represent a class of tRNAs with intrinsically weak structures that may be the result of expedited evolution and streamlining of the human mitochondrial genome. The inherent conformational fragility and instability of these tRNAs may amplify the effects of single-base substitutions, by disrupting a native conformation of structural elements or by promoting the formation of intermediate, unstable, non-native structures. REFERENCES 1. Allen JF, Raven JA. Free-radical-induced mutation vs redox regulation: costs and benefits of genes in organelles. J Mol Evol 1996;42:482– 492. 2. Ames BN. Endogenous oxidative DNA damage, aging, and cancer. Free Radic Res Commun 1989;7:121–128. 3. Anitori R, Manning K, Quan F, Weleber RG, Buist NR, Shoubridge EA, et al. Contrasting phenotypes in three patients with novel mutations in mitochondrial tRNA genes. Mol Genet Metab 2005;84:176 –188. 4. Ashraf SS, Sochacka E, Cain R, Guenther R, Malkiewicz A, Agris PF. Single atom modification (O–⬎S) of tRNA confers ribosome binding. RNA 1999;5:188 –194. 5. Bacman SR, Atencio DP, Moraes CT. Decreased mitochondrial tRNALys steady-state levels and aminoacylation are associated with the pathogenic G8313A mitochondrial DNA mutation. Biochem J 2003;374:131–136. 6. Bataillard M, Chatzoglou E, Rumbach L, Sternberg D, Tournade A, Laforet P, et al. Atypical MELAS syndrome associated with a new mitochondrial tRNA glutamine point mutation. Neurology 2001;56:405– 407. 7. Berkovic SF, Shoubridge EA, Andermann F, Andermann E, Carpenter S, Karpati G. Clinical spectrum of mitochondrial DNA mutation at base pair 8344. Lancet 1991;338:457. MUSCLE & NERVE February 2008 165 8. Bernier FP, Boneh A, Dennett X, Chow CW, Cleary MA, Thorburn DR. Diagnostic criteria for respiratory chain disorders in adults and children. Neurology 2002;59:1406 –1411. 9. Bidooki S, Jackson MJ, Johnson MA, Chrzanowska-Lightowlers ZM, Taylor RW, Venables G, et al. Sporadic mitochondrial myopathy due to a new mutation in the mitochondrial tRNASer(UCN) gene. Neuromuscul Disord 2004;14:417– 420. 10. Bidooki SK, Johnson MA, Chrzanowska-Lightowlers Z, Bindoff LA, Lightowlers RN. Intracellular mitochondrial triplasmy in a patient with two heteroplasmic base changes. Am J Hum Genet 1997;60:1430 –1438. 11. Bindoff LA, Howell N, Poulton J, McCullough DA, Morten KJ, Lightowlers RN, et al. Abnormal RNA processing associated with a novel tRNA mutation in mitochondrial DNA. A potential disease mechanism. J Biol Chem 1993;268:19559 – 19564. 12. Blakely EL, Poulton J, Pike M, Wojnarowska F, Turnbull DM, McFarland R, et al. Childhood neurological presentation of a novel mitochondrial tRNA(Val) gene mutation. J Neurol Sci 2004;225:99 –103. 13. Borthwick GM, Taylor RW, Walls TJ, Tonska K, Taylor GA, Shaw PJ, et al. Motor neuron disease in a patient with a mitochondrial tRNAIle mutation. Ann Neurol 2006;59:570 – 574. 14. Brown MD, Torroni A, Shoffner JM, Wallace DC. Mitochondrial tRNA(Thr) mutations and lethal infantile mitochondrial myopathy. Am J Hum Genet 1992;51:446 – 447. 15. Brule H, Holmes WM, Keith G, Giege R, Florentz C. Effect of a mutation in the anticodon of human mitochondrial tRNAPro on its post-transcriptional modification pattern. Nucleic Acids Res 1998;26:537–543. 16. Bruno C, Sacco O, Santorelli FM, Assereto S, Tonoli E, Bado M, et al. Mitochondrial myopathy and respiratory failure associated with a new mutation in the mitochondrial transfer ribonucleic acid glutamic acid gene. J Child Neurol 2003; 18:300 –303. 17. Campos Y, Gamez J, Garcia A, Andreu AL, Rubio JC, Martin MA, et al. A new mtDNA mutation in the tRNA(Leu(UUR)) gene associated with ocular myopathy. Neuromuscul Disord 2001;11:477– 480. 18. Campos Y, Garcia A, del Hoyo P, Jara P, Martin MA, Rubio JC, et al. Two pathogenic mutations in the mitochondrial DNA tRNA Leu(UUR) gene (T3258C and A3280G) resulting in variable clinical phenotypes. Neuromuscul Disord 2003;13:416 – 420. 19. Cardaioli E, Da Pozzo P, Radi E, Dotti MT, Federico A. A novel heteroplasmic tRNA(Leu(CUN)) mtDNA point mutation associated with chronic progressive external ophthalmoplegia. Biochem Biophys Res Commun 2005;327:675– 678. 20. Casali C, Santorelli FM, D’Amati G, Bernucci P, DeBiase L, DiMauro S. A novel mtDNA point mutation in maternally inherited cardiomyopathy. Biochem Biophys Res Commun 1995;213:588 –593. 21. Chalmers RM, Lamont PJ, Nelson I, Ellison DW, Thomas NH, Harding AE, et al. A mitochondrial DNA tRNA(Val) point mutation associated with adult-onset Leigh syndrome. Neurology 1997;49:589 –592. 22. Chapiro E, Feldmann D, Denoyelle F, Sternberg D, Jardel C, Eliot MM, et al. Two large French pedigrees with non syndromic sensorineural deafness and the mitochondrial DNA T7511C mutation: evidence for a modulatory factor. Eur J Hum Genet 2002;10:851– 856. 23. Chinnery PF, Johnson MA, Taylor RW, Durward WF, Turnbull DM. A novel mitochondrial tRNA isoleucine gene mutation causing chronic progressive external ophthalmoplegia. Neurology 1997;49:1166 –1168. 24. Chomyn A, Martinuzzi A, Yoneda M, Daga A, Hurko O, Johns D, et al. MELAS mutation in mtDNA binding site for transcription termination factor causes defects in protein synthesis and in respiration but no change in levels of up- 166 Human Mitochondrial tRNAs 25. 26. 27. 28. 29. 30. 31. 32. 33. 34. 35. 36. 37. 38. 39. 40. 41. stream and downstream mature transcripts. Proc Natl Acad Sci U S A 1992;89:4221– 4225. Ciafaloni E, Ricci E, Servidei S, Shanske S, Silvestri G, Manfredi G, et al. Widespread tissue distribution of a tRNALeu(UUR) mutation in the mitochondrial DNA of a patient with MELAS syndrome. Neurology 1991;41:1663–1664. Corona P, Lamantea E, Greco M, Carrara F, Agostino A, Guidetti D, et al. Novel heteroplasmic mtDNA mutation in a family with heterogeneous clinical presentations. Ann Neurol 2002;51:118 –122. Coulbault L, Herlicoviez D, Chapon F, Read MH, Penniello MJ, Reynier P, et al. A novel mutation in the mitochondrial tRNA Asn gene associated with a lethal disease. Biochem Biophys Res Commun 2005;329:1152–1154. Crimi M, Galbiati S, Perini MP, Bordoni A, Malferrari G, Sciacco M, et al. A mitochondrial tRNA(His) gene mutation causing pigmentary retinopathy and neurosensorial deafness. Neurology 2003;60:1200 –1203. Crimi M, Galbiati S, Sciacco M, Bordoni A, Natali MG, Raimondi M, et al. Mitochondrial-DNA nucleotides G4298A and T10010C as pathogenic mutations: the confirmation in two new cases. Mitochondrion 2004;3:279 –283. Darin N, Kollberg G, Moslemi AR, Tulinius M, Holme E, Gronlund MA, et al. Mitochondrial myopathy with exercise intolerance and retinal dystrophy in a sporadic patient with a G583A mutation in the mt tRNA(phe) gene. Neuromuscul Disord 2006;16:504 –506. de Coo IF, Sistermans EA, de Wijs IJ, Catsman-Berrevoets C, Busch HF, Scholte HR, et al. A mitochondrial tRNA(Val) gene mutation (G1642A) in a patient with mitochondrial myopathy, lactic acidosis, and stroke-like episodes. Neurology 1998;50:293–295. Degoul F, Brule H, Cepanec C, Helm M, Marsac C, Leroux J, et al. Isoleucylation properties of native human mitochondrial tRNAIle and tRNAIle transcripts. Implications for cardiomyopathy-related point mutations (4269, 4317) in the tRNAIle gene. Hum Mol Genet 1998;7:347–354. Deschauer M, Swalwell H, Strauss M, Zierz S, Taylor RW. Novel mitochondrial transfer RNA(Phe) gene mutation associated with late-onset neuromuscular disease. Arch Neurol 2006;63:902–905. Dey R, Tengan CH, Morita MP, Kiyomoto BH, Moraes CT. A novel myopathy-associated mitochondrial DNA mutation altering the conserved size of the tRNA(Gln) anticodon loop. Neuromuscul Disord 2000;10:488 – 492. DiMauro S, Schon EA. Mitochondrial DNA mutations in human disease. Am J Med Genet 2001;106:18 –26. DiMauro S, Hirano M, Kaufmann P, Tanji K, Sano M, Shungu DC, et al. Clinical features and genetics of myoclonic epilepsy with ragged red fibers. Adv Neurol 2002;89: 217–229. Enriquez JA, Chomyn A, Attardi G. MtDNA mutation in MERRF syndrome causes defective aminoacylation of tRNA(Lys) and premature translation termination. Nat Genet 1995;10:47–55. Feigenbaum A, Bai RK, Doherty ES, Kwon H, Tan D, Sloane A, et al. Novel mitochondrial DNA mutations associated with myopathy, cardiomyopathy, renal failure, and deafness. Am J Med Genet A 2006;140:2216 –2222. Finnila S, Autere J, Lehtovirta M, Hartikainen P, Mannermaa A, Soininen H, et al. Increased risk of sensorineural hearing loss and migraine in patients with a rare mitochondrial DNA variant 4336A⬎G in tRNAGln. J Med Genet 2001;38:400 – 405. Finnila S, Tuisku S, Herva R, Majamaa K. A novel mitochondrial DNA mutation and a mutation in the Notch3 gene in a patient with myopathy and CADASIL. J Mol Med 2001;79: 641– 647. Florentz C, Sissler M. Disease-related versus polymorphic mutations in human mitochondrial tRNAs. Where is the difference? EMBO Rep 2001;2:481– 486. MUSCLE & NERVE February 2008 42. Florentz C, Sohm B, Tryoen-Toth P, Putz J, Sissler M. Human mitochondrial tRNAs in health and disease. Cell Mol Life Sci 2003;60:1356 –1375. 43. Franceschina L, Salani S, Bordoni A, Sciacco M, Napoli L, Comi GP, et al. A novel mitochondrial tRNA(Ile) point mutation in chronic progressive external ophthalmoplegia. J Neurol 1998;245:755–758. 44. Fu K, Hartlen R, Johns T, Genge A, Karpati G, Shoubridge EA. A novel heteroplasmic tRNAleu(CUN) mtDNA point mutation in a sporadic patient with mitochondrial encephalomyopathy segregates rapidly in skeletal muscle and suggests an approach to therapy. Hum Mol Genet 1996;5:1835– 1840. 45. Gabriel K, Schneider J, McClain WH. Functional evidence for indirect recognition of G.U in tRNA(Ala) by alanyl-tRNA synthetase. Science 1996;271:195–197. 46. Gambello MJ, Bai RK, Chen TJ, Dimachkie M, Wong LJ. Exercise intolerance associated with a novel 8300T ⬎ C mutation in mitochondrial transfer RNAlys. Muscle Nerve 2006;34:437– 443. 47. Gattermann N, Retzlaff S, Wang YL, Berneburg M, Heinisch J, Wlaschek M, et al. A heteroplasmic point mutation of mitochondrial tRNALeu(CUN) in non-lymphoid haemopoietic cell lineages from a patient with acquired idiopathic sideroblastic anaemia. Br J Haematol 1996;93:845– 855. 48. Goldstein JD, Shanske S, Bruno C, Perszyk AA. Maternally inherited mitochondrial cardiomyopathy associated with a C-to-T transition at nucleotide 3303 of mitochondrial DNA in the tRNA(Leu(UUR)) gene. Pediatr Dev Pathol 1999;2: 78 – 85. 49. Goto Y, Tojo M, Tohyama J, Horai S, Nonaka I. A novel point mutation in the mitochondrial tRNA(Leu)(UUR) gene in a family with mitochondrial myopathy. Ann Neurol 1992;31: 672– 675. 50. Goto Y, Tsugane K, Tanabe Y, Nonaka I, Horai S. A new point mutation at nucleotide pair 3291 of the mitochondrial tRNA(Leu(UUR)) gene in a patient with mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes (MELAS). Biochem Biophys Res Commun 1994;202: 1624 –1630. 51. Grafakou O, Hol FA, Otfried Schwab K, Siers MH, ter Laak H, Trijbels F, et al. Exercise intolerance, muscle pain and lactic acidaemia associated with a 7497G⬎A mutation in the tRNASer(UCN) gene. J Inherit Metab Dis 2003;26:593– 600. 52. Grasbon-Frodl EM, Kosel S, Sprinzl M, von Eitzen U, Mehraein P, Graeber MB. Two novel point mutations of mitochondrial tRNA genes in histologically confirmed Parkinson disease. Neurogenetics 1999;2:121–127. 53. Grasso M, Diegoli M, Brega A, Campana C, Tavazzi L, Arbustini E. The mitochondrial DNA mutation T12297C affects a highly conserved nucleotide of tRNA(Leu(CUN)) and is associated with dilated cardiomyopathy. Eur J Hum Genet 2001;9:311–315. 54. Hadjigeorgiou GM, Kim SH, Fischbeck KH, Andreu AL, Berry GT, Bingham P, et al. Shanske S, Bonilla E, DiMau A new mitochondrial DNA mutation (A3288G) in the tRNA(Leu(UUR)) gene associated with familial myopathy. J Neurol Sci 1999;164:153–157. 55. Hamann CS, Hou YM. Probing a tRNA core that contributes to aminoacylation. J Mol Biol 2000;295:777–789. 56. Hanna MG, Nelson I, Sweeney MG, Cooper JM, Watkins PJ, Morgan-Hughes JA, et al. Congenital encephalomyopathy and adult-onset myopathy and diabetes mellitus: different phenotypic associations of a new heteroplasmic mtDNA tRNA glutamic acid mutation. Am J Hum Genet 1995;56: 1026 –1033. 57. Hanna MG, Nelson IP, Morgan-Hughes JA, Wood NW. MELAS: a new disease associated mitochondrial DNA mutation and evidence for further genetic heterogeneity. J Neurol Neurosurg Psychiatry 1998;65:512–517. Human Mitochondrial tRNAs 58. Hanna MG, Nelson IP. Genetics and molecular pathogenesis of mitochondrial respiratory chain diseases. Cell Mol Life Sci 1999;55:691–706. 59. Hao H, Bonilla E, Manfredi G, DiMauro S, Moraes CT. Segregation patterns of a novel mutation in the mitochondrial tRNA glutamic acid gene associated with myopathy and diabetes mellitus. Am J Hum Genet 1995;56:1017–1025. 60. Hao H, Moraes CT. Functional and molecular mitochondrial abnormalities associated with a C–⬎ T transition at position 3256 of the human mitochondrial genome. The effects of a pathogenic mitochondrial tRNA point mutation in organelle translation and RNA processing. J Biol Chem 1996;271:2347–2352. 61. Hao H, Moraes CT. A disease-associated G5703A mutation in human mitochondrial DNA causes a conformational change and a marked decrease in steady-state levels of mitochondrial tRNA(Asn). Mol Cell Biol 1997;17:6831– 6837. 62. Hasegawa H, Matsuoka T, Goto Y, Nonaka I. Strongly succinate dehydrogenase-reactive blood vessels in muscles from patients with mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes. Ann Neurol 1991;29: 601– 605. 63. Hattori Y, Goto Y, Sakuta R, Nonaka I, Mizuno Y, Horai S. Point mutations in mitochondrial tRNA genes: sequence analysis of chronic progressive external ophthalmoplegia (CPEO). J Neurol Sci 1994;125:50 –55. 64. Hayashi J, Ohta S, Takai D, Miyabayashi S, Sakuta R, Goto Y, et al. Accumulation of mtDNA with a mutation at position 3271 in tRNA(Leu)(UUR) gene introduced from a MELAS patient to HeLa cells lacking mtDNA results in progressive inhibition of mitochondrial respiratory function. Biochem Biophys Res Commun 1993;197:1049 –1055. 65. Helm M, Giege R, Florentz C. A Watson-Crick base-pairdisrupting methyl group (m1A9) is sufficient for cloverleaf folding of human mitochondrial tRNALys. Biochemistry 1999;38:13338 –13346. 66. Horvath R, Lochmuller H, Scharfe C, Do BH, Oefner PJ, Muller-Hocker J, et al. A tRNA(Ala) mutation causing mitochondrial myopathy clinically resembling myotonic dystrophy. J Med Genet 2003;40:752–757. 67. Houshmand M, Lindberg C, Moslemi AR, Oldfors A, Holme E. A novel heteroplasmic point mutation in the mitochondrial tRNA(Lys) gene in a sporadic case of mitochondrial encephalomyopathy: de novo mutation and no transmission to the offspring. Hum Mutat 1999;13:203–209. 68. Hutchin TP, Parker MJ, Young ID, Davis AC, Pulleyn LJ, Deeble J, et al. A novel mutation in the mitochondrial tRNA(Ser(UCN)) gene in a family with non-syndromic sensorineural hearing impairment. J Med Genet 2000;37:692– 694. 69. Hutchin TP, Navarro-Coy NC, Van Camp G, Tiranti V, Zeviani M, Schuelke M, et al. Multiple origins of the mtDNA 7472insC mutation associated with hearing loss and neurological dysfunction. Eur J Hum Genet 2001;9:385–387. 70. Ingman M, Kaessmann H, Paabo S, Gyllensten U. Mitochondrial genome variation and the origin of modern humans. Nature 2000;408:708 –713. 71. Ionasescu VV, Hart M, DiMauro S, Moraes CT. Clinical and morphologic features of a myopathy associated with a point mutation in the mitochondrial tRNA(Pro) gene. Neurology 1994;44:975–977. 72. Jaksch M, Klopstock T, Kurlemann G, Dorner M, Hofmann S, Kleinle S, et al. Progressive myoclonus epilepsy and mitochondrial myopathy associated with mutations in the tRNA(Ser(UCN)) gene. Ann Neurol 1998;44:635– 640. 73. Jaksch M, Lochmuller H, Schmitt F, Volpel B, ObermaierKusser B, Horvath R. A mutation in mt tRNALeu(UUR) causing a neuropsychiatric syndrome with depression and cataract. Neurology 2001;57:1930 –1931. 74. Janssen GM, Maassen JA, van Den Ouweland JM. The diabetes-associated 3243 mutation in the mitochondrial tRNA(Leu(UUR)) gene causes severe mitochondrial dys- MUSCLE & NERVE February 2008 167 75. 76. 77. 78. 79. 80. 81. 82. 83. 84. 85. 86. 87. 88. 89. 90. 91. 92. 168 function without a strong decrease in protein synthesis rate. J Biol Chem 1999;274:29744 –29748. Jeppesen TD, Schwartz M, Hansen K, Danielsen ER, Wibrand F, Vissing J. Late onset of stroke-like episode associated with a 3256C–⬎T point mutation of mitochondrial DNA. J Neurol Sci 2003;214:17–20. Karadimas C, Tanji K, Geremek M, Chronopoulou P, Vu T, Krishna S, et al. A5814G mutation in mitochondrial DNA can cause mitochondrial myopathy and cardiomyopathy. J Child Neurol 2001;16:531–533. Karadimas CL, Salviati L, Sacconi S, Chronopoulou P, Shanske S, Bonilla E, et al. Mitochondrial myopathy and ophthalmoplegia in a sporadic patient with the G12315A mutation in mitochondrial DNA. Neuromuscul Disord 2002;12:865– 868. Kawarai T, Kawakami H, Kozuka K, Izumi Y, Matsuyama Z, Watanabe C, et al. A new mitochondrial DNA mutation associated with mitochondrial myopathy: tRNA(Leu)(UUR) 3254C-to-G. Neurology 1997;49:598 – 600. Kelley SO, Steinberg SV, Schimmel P. Functional defects of pathogenic human mitochondrial tRNAs related to structural fragility. Nat Struct Biol 2000;7:862– 865. Kelley SO, Steinberg SV, Schimmel P. Fragile T-stem in disease-associated human mitochondrial tRNA sensitizes structure to local and distant mutations. J Biol Chem 2001; 276:10607–10611. King MP, Attardi G. Human cells lacking mtDNA: repopulation with exogenous mitochondria by complementation. Science 1989;246:500 –503. King MP, Koga Y, Davidson M, Schon EA. Defects in mitochondrial protein synthesis and respiratory chain activity segregate with the tRNA(Leu(UUR)) mutation associated with mitochondrial myopathy, encephalopathy, lactic acidosis, and strokelike episodes. Mol Cell Biol 1992;12:480 – 490. Kirino Y, Yasukawa T, Ohta S, Akira S, Ishihara K, Watanabe K, et al. Codon-specific translational defect caused by a wobble modification deficiency in mutant tRNA from a human mitochondrial disease. Proc Natl Acad Sci U S A 2004; 101:15070 –15075. Kirino Y, Goto Y, Campos Y, Arenas J, Suzuki T. Specific correlation between the wobble modification deficiency in mutant tRNAs and the clinical features of a human mitochondrial disease. Proc Natl Acad Sci U S A 2005;102:7127– 7132. Kleinle S, Schneider V, Moosmann P, Brandner S, Krahenbuhl S, Liechti-Gallati S. A novel mitochondrial tRNA(Phe) mutation inhibiting anticodon stem formation associated with a muscle disease. Biochem Biophys Res Commun 1998; 247:112–115. Koga A, Koga Y, Akita Y, Fukiyama R, Ueki I, Yatsuga S, et al. Increased mitochondrial processing intermediates associated with three tRNA(Leu(UUR)) gene mutations. Neuromuscul Disord 2003;13:259 –262. Koga Y, Yoshino M, Kato H. MELAS exhibits dominant negative effects on mitochondrial RNA processing. Ann Neurol 1998;43:835. Kogelnik AM, Lott MT, Brown MD, Navathe SB, Wallace DC. MITOMAP: a human mitochondrial genome database— 1998 update. Nucleic Acids Res 1998;26:112–115. Lauber J, Marsac C, Kadenbach B, Seibel P. Mutations in mitochondrial tRNA genes: a frequent cause of neuromuscular diseases. Nucleic Acids Res 1991;19:1393–1397. Levinger L, Giege R, Florentz C. Pathology-related substitutions in human mitochondrial tRNA(Ile) reduce precursor 3⬘ end processing efficiency in vitro. Nucleic Acids Res 2003; 31:1904 –1912. Levinger L, Morl M, Florentz C. Mitochondrial tRNA 3⬘ end metabolism and human disease. Nucleic Acids Res 2004;32: 5430 –5441. Lombes A, Bories D, Girodon E, Frachon P, Ngo MM, Breton-Gorius J, et al. The first pathogenic mitochondrial me- Human Mitochondrial tRNAs 93. 94. 95. 96. 97. 98. 99. 100. 101. 102. 103. 104. 105. 106. 107. 108. 109. thionine tRNA point mutation is discovered in splenic lymphoma. Hum Mutat 1998;(Suppl 1):S175–183. Lynn S, Wardell T, Johnson MA, Chinnery PF, Daly ME, Walker M, et al. Mitochondrial diabetes: investigation and identification of a novel mutation. Diabetes 1998;47:1800 – 1802. Mancuso M, Filosto M, Mootha VK, Rocchi A, Pistolesi S, Murri L, et al. A novel mitochondrial tRNAPhe mutation causes MERRF syndrome. Neurology 2004;62:2119 –2121. Manfredi G, Schon EA, Bonilla E, Moraes CT, Shanske S, DiMauro S. Identification of a mutation in the mitochondrial tRNA(Cys) gene associated with mitochondrial encephalopathy. Hum Mutat 1996;7:158 –163. Maniura-Weber K, Taylor RW, Johnson MA, ChrzanowskaLightowlers Z, Morris AA, Charlton CP, et al. A novel point mutation in the mitochondrial tRNA(Trp) gene produces a neurogastrointestinal syndrome. Eur J Hum Genet 2004;12: 509 –512. Mansergh FC, Millington-Ward S, Kennan A, Kiang AS, Humphries M, Farrar GJ, et al. Retinitis pigmentosa and progressive sensorineural hearing loss caused by a C12258A mutation in the mitochondrial MTTS2 gene. Am J Hum Genet 1999;64:971–985. Mariotti C, Tiranti V, Carrara F, Dallapiccola B, DiDonato S, Zeviani M. Defective respiratory capacity and mitochondrial protein synthesis in transformant cybrids harboring the tRNA(Leu(UUR)) mutation associated with maternally inherited myopathy and cardiomyopathy. J Clin Invest 1994; 93:1102–1107. Masucci JP, Davidson M, Koga Y, Schon EA, King MP. In vitro analysis of mutations causing myoclonus epilepsy with ragged-red fibers in the mitochondrial tRNA(Lys)gene: two genotypes produce similar phenotypes. Mol Cell Biol 1995; 15:2872–2881. McFarland R, Clark KM, Morris AA, Taylor RW, Macphail S, Lightowlers RN, et al. Multiple neonatal deaths due to a homoplasmic mitochondrial DNA mutation. Nat Genet 2002;30:145–146. McFarland R, Elson JL, Taylor RW, Howell N, Turnbull DM. Assigning pathogenicity to mitochondrial tRNA mutations: when “definitely maybe” is not good enough. Trends Genet 2004;20:591–596. McFarland R, Schaefer AM, Gardner JL, Lynn S, Hayes CM, Barron MJ, et al. Familial myopathy: new insights into the T14709C mitochondrial tRNA mutation. Ann Neurol 2004; 55:478 – 484. Melone MA, Tessa A, Petrini S, Lus G, Sampaolo S, di Fede G, et al. Revelation of a new mitochondrial DNA mutation (G12147A) in a MELAS/MERFF phenotype. Arch Neurol 2004;61:269 –272. Merante F, Tein I, Benson L, Robinson BH. Maternally inherited hypertrophic cardiomyopathy due to a novel Tto-C transition at nucleotide 9997 in the mitochondrial tRNA(glycine) gene. Am J Hum Genet 1994;55:437– 446. Merante F, Myint T, Tein I, Benson L, Robinson BH. An additional mitochondrial tRNA(Ile) point mutation (A-to-G at nucleotide 4295) causing hypertrophic cardiomyopathy. Hum Mutat 1996;8:216 –222. Meulemans A, Seneca S, Lagae L, Lissens W, De Paepe B, Smet J, et al. A novel mitochondrial transfer RNA(Asn) mutation causing multiorgan failure. Arch Neurol 2006;63: 1194 –1198. Meulemans A, Seneca S, Smet J, De Paepe B, Lissens W, Van Coster R, et al. A new family with the mitochondrial tRNAGLU gene mutation m.14709T⬎C presenting with hydrops fetalis. Eur J Paediatr Neurol 2007;11:17–20. Mitchell AL, Elson JL, Howell N, Taylor RW, Turnbull DM. Sequence variation in mitochondrial complex I genes: mutation or polymorphism? J Med Genet 2006;43:175–179. Moraes CT, Ciacci F, Bonilla E, Ionasescu V, Schon EA, DiMauro S. A mitochondrial tRNA anticodon swap associated with a muscle disease. Nat Genet 1993;4:284 –288. MUSCLE & NERVE February 2008 110. Moraes CT, Ciacci F, Bonilla E, Jansen C, Hirano M, Rao N, et al. Two novel pathogenic mitochondrial DNA mutations affecting organelle number and protein synthesis. Is the tRNA(Leu(UUR)) gene an etiologic hot spot? J Clin Invest 1993;92:2906 –2915. 111. Morten KJ, Cooper JM, Brown GK, Lake BD, Pike D, Poulton J. A new point mutation associated with mitochondrial encephalomyopathy. Hum Mol Genet 1993;2:2081–2087. 112. Moslemi AR, Lindberg C, Toft J, Holme E, Kollberg G, Oldfors A. A novel mutation in the mitochondrial tRNA(Phe) gene associated with mitochondrial myopathy. Neuromuscul Disord 2004;14:46 –50. 113. Nelson I, Hanna MG, Alsanjari N, Scaravilli F, MorganHughes JA, Harding AE. A new mitochondrial DNA mutation associated with progressive dementia and chorea: a clinical, pathological, and molecular genetic study. Ann Neurol 1995;37:400 – 403. 114. Nishigaki Y, Bonilla E, Shanske S, Gaskin DA, DiMauro S, Hirano M. Exercise-induced muscle “burning,” fatigue, and hyper-CKemia: mtDNA T10010C mutation in tRNA(Gly). Neurology 2002;58:1282–1285. 115. Nishigaki Y, Tadesse S, Bonilla E, Shungu D, Hersh S, Keats BJ, et al. A novel mitochondrial tRNA(Leu(UUR)) mutation in a patient with features of MERRF and Kearns-Sayre syndrome. Neuromuscul Disord 2003;13:334 –340. 116. Nishino I, Seki A, Maegaki Y, Takeshita K, Horai S, Nonaka I, et al. A novel mutation in the mitochondrial tRNA(Thr) gene associated with a mitochondrial encephalomyopathy. Biochem Biophys Res Commun 1996;225:180 –185. 117. Ohama E, Ohara S, Ikuta F, Tanaka K, Nishizawa M, Miyatake T. Mitochondrial angiopathy in cerebral blood vessels of mitochondrial encephalomyopathy. Acta Neuropathol (Berl) 1987;74:226 –233. 118. Ozawa M, Nishino I, Horai S, Nonaka I, Goto YI. Myoclonus epilepsy associated with ragged-red fibers: a G-to-A mutation at nucleotide pair 8363 in mitochondrial tRNA(Lys) in two families. Muscle Nerve 1997;20:271–278. 119. Pandya A, Xia XJ, Erdenetungalag R, Amendola M, Landa B, Radnaabazar J, et al. Heterogenous point mutations in the mitochondrial tRNA Ser(UCN) precursor coexisting with the A1555G mutation in deaf students from Mongolia. Am J Hum Genet 1999;65:1803–1806. 120. Park H, Davidson E, King MP. The pathogenic A3243G mutation in human mitochondrial tRNALeu(UUR) decreases the efficiency of aminoacylation. Biochemistry 2003; 42:958 –964. 121. Perucca-Lostanlen D, Taylor RW, Narbonne H, Mousson de Camaret B, Hayes CM, Saunieres A, et al. Molecular and functional effects of the T14709C point mutation in the mitochondrial DNA of a patient with maternally inherited diabetes and deafness. Biochim Biophys Acta 2002;1588: 210 –216. 122. Pesole G, Gissi C, De Chirico A, Saccone C. Nucleotide substitution rate of mammalian mitochondrial genomes. J Mol Evol 1999;48:427– 434. 123. Pulkes T, Siddiqui A, Morgan-Hughes JA, Hanna MG. A novel mutation in the mitochondrial tRNA(TYr) gene associated with exercise intolerance. Neurology 2000;55:1210 – 1212. 124. Pulkes T, Liolitsa D, Eunson LH, Rose M, Nelson IP, Rahman S, et al. New phenotypic diversity associated with the mitochondrial tRNA(SerUCN) gene mutation. Neuromuscul Disord 2005;15:364 –371. 125. Putz J, Florentz C, Benseler F, Giege R. A single methyl group prevents the mischarging of a tRNA. Nat Struct Biol 1994;1:580 –582. 126. Raffelsberger T, Rossmanith W, Thaller-Antlanger H, Bittner RE. CPEO associated with a single nucleotide deletion in the mitochondrial tRNA(Tyr) gene. Neurology 2001;57:2298–2301. 127. Reid FM, Vernham GA, Jacobs HT. A novel mitochondrial point mutation in a maternal pedigree with sensorineural deafness. Hum Mutat 1994;3:243–247. Human Mitochondrial tRNAs 128. Rossmanith W, Tullo A, Potuschak T, Karwan R, Sbisa E. Human mitochondrial tRNA processing. J Biol Chem 1995; 270:12885–12891. 129. Rossmanith W, Raffelsberger T, Roka J, Kornek B, Feucht M, Bittner RE. The expanding mutational spectrum of MERRF substitution G8361A in the mitochondrial tRNALys gene. Ann Neurol 2003;54:820 – 823. 130. Sacconi S, Salviati L, Gooch C, Bonilla E, Shanske S, DiMauro S. Complex neurologic syndrome associated with the G1606A mutation of mitochondrial DNA. Arch Neurol 2002; 59:1013–1015. 131. Sahashi K, Yoneda M, Ohno K, Tanaka M, Ibi T, Sahashi K. Functional characterisation of mitochondrial tRNA(Tyr) mutation (5877–⬎GA) associated with familial chronic progressive external ophthalmoplegia. J Med Genet 2001;38: 703–705. 132. Sakuta R, Goto Y, Horai S, Nonaka I. Mitochondrial DNA mutations at nucleotide positions 3243 and 3271 in mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes: a comparative study. J Neurol Sci 1993; 115:158 –160. 133. Sakuta R, Honzawa S, Murakami N, Goto Y, Nagai T. Atypical MELAS associated with mitochondrial tRNA(Lys) gene A8296G mutation. Pediatr Neurol 2002;27:397– 400. 134. Santorelli FM, Mak SC, Vazquez-Acevedo M, Gonzalez-Astiazaran A, Ridaura-Sanz C, Gonzalez-Halphen D, et al. A novel mitochondrial DNA point mutation associated with mitochondrial encephalocardiomyopathy. Biochem Biophys Res Commun 1995;216:835– 840. 135. Santorelli FM, Schlessel JS, Slonim AE, DiMauro S. Novel mutation in the mitochondrial DNA tRNA glycine gene associated with sudden unexpected death. Pediatr Neurol 1996;15:145–149. 136. Santorelli FM, Siciliano G, Casali C, Basirico MG, Carrozzo R, Calvosa F, et al. Mitochondrial tRNA(Cys) gene mutation (A5814G): a second family with mitochondrial encephalopathy. Neuromuscul Disord 1997;7:156 –159. 137. Santorelli FM, Tanji K, Sano M, Shanske S, El-Shahawi M, Kranz-Eble P, et al. Maternally inherited encephalopathy associated with a single-base insertion in the mitochondrial tRNATrp gene. Ann Neurol 1997;42:256 –260. 138. Scaglia F, Vogel H, Hawkins EP, Vladutiu GD, Liu LL, Wong LJ. Novel homoplasmic mutation in the mitochondrial tRNATyr gene associated with atypical mitochondrial cytopathy presenting with focal segmental glomerulosclerosis. Am J Med Genet A 2003;123:172–178. 139. Scaglia F, Northrop JL. The mitochondrial myopathy encephalopathy, lactic acidosis with stroke-like episodes (MELAS) syndrome: a review of treatment options. CNS Drugs 2006;20:443– 464. 140. Scheffler IE. Mitochondria make a come back. Adv Drug Deliv Rev 2001;49:3–26. 141. Schon EA, Koga Y, Davidson M, Moraes CT, King MP. The mitochondrial tRNA(Leu)(UUR)) mutation in MELAS: a model for pathogenesis. Biochim Biophys Acta 1992;1101: 206 –209. 142. Schon EA, Bonilla E, DiMauro S. Mitochondrial DNA mutations and pathogenesis. J Bioenerg Biomembr 1997;29:131– 149. 143. Seibel P, Lauber J, Klopstock T, Marsac C, Kadenbach B, Reichmann H. Chronic progressive external ophthalmoplegia is associated with a novel mutation in the mitochondrial tRNA(Asn) gene. Biochem Biophys Res Commun 1994;204: 482– 489. 144. Seki A, Nishino I, Goto Y, Maegaki Y, Koeda T. Mitochondrial encephalomyopathy with 15915 mutation: clinical report. Pediatr Neurol 1997;17:161–164. 145. Seneca S, Lissens W, Liebaers I, van den Bergh P, Nassogne MC, Benatar A, et al. Pitfalls in the diagnosis of mtDNA mutations. J Med Genet 1998;35:963–964. MUSCLE & NERVE February 2008 169 146. Seneca S, Ceuterik-De Groote C, Van Coster R, De Meirleir L. A novel mitochondrial transfer RNA proline mutation. J Inherit Metab Dis 2000;23:853– 854. 147. Seneca S, Verhelst H, De Meirleir L, Meire F, Ceuterick-De Groote C, Lissens W, et al. A new mitochondrial point mutation in the transfer RNA(Leu) gene in a patient with a clinical phenotype resembling Kearns-Sayre syndrome. Arch Neurol 2001;58:1113–1118. 148. Seneca S, Goemans N, Van Coster R, Givron P, Reybrouck T, Sciot R, et al. A mitochondrial tRNA aspartate mutation causing isolated mitochondrial myopathy. Am J Med Genet A 2005;137:170 –175. 149. Sevior KB, Hatamochi A, Stewart IA, Bykhovskaya Y, AllenPowell DR, Fischel-Ghodsian N, et al. Mitochondrial A7445G mutation in two pedigrees with palmoplantar keratoderma and deafness. Am J Med Genet 1998;75:179 –185. 150. Shaag A, Saada A, Steinberg A, Navon P, Elpeleg ON. Mitochondrial encephalomyopathy associated with a novel mutation in the mitochondrial tRNA(leu)(UUR) gene (A3243T). Biochem Biophys Res Commun 1997;233:637– 639. 151. Shin WS, Tanaka M, Suzuki J, Hemmi C, Toyo-oka T. A novel homoplasmic mutation in mtDNA with a single evolutionary origin as a risk factor for cardiomyopathy. Am J Hum Genet 2000;67:1617–1620. 152. Shoffner JM, Bialer MG, Pavlakis SG, Lott M, Kaufman A, Dixon J, et al. Mitochondrial encephalomyopathy associated with a single nucleotide pair deletion in the mitochondrial tRNALeu(UUR) gene. Neurology 1995;45:286 –292. 153. Shoubridge EA. Mitochondrial DNA diseases: histological and cellular studies. J Bioenerg Biomembr 1994;26:301–310. 154. Shtilbans A, El-Schahawi M, Malkin E, Shanske S, Musumeci O, DiMauro S. A novel mutation in the mitochondrial DNA transfer ribonucleic acidAsp gene in a child with myoclonic epilepsy and psychomotor regression. J Child Neurol 1999; 14:610 – 613. 155. Silvestri G, Santorelli FM, Shanske S, Whitley CB, Schimmenti LA, Smith SA, et al. A new mtDNA mutation in the tRNA(Leu(UUR)) gene associated with maternally inherited cardiomyopathy. Hum Mutat 1994;3:37– 43. 156. Silvestri G, Servidei S, Rana M, Ricci E, Spinazzola A, Paris E, et al. A novel mitochondrial DNA point mutation in the tRNA(Ile) gene is associated with progressive external ophtalmoplegia. Biochem Biophys Res Commun 1996;220:623– 627. 157. Silvestri G, Rana M, DiMuzio A, Uncini A, Tonali P, Servidei S. A late-onset mitochondrial myopathy is associated with a novel mitochondrial DNA (mtDNA) point mutation in the tRNA(Trp) gene. Neuromuscul Disord 1998;8:291–295. 158. Silvestri G, Mongini T, Odoardi F, Modoni A, deRosa G, Doriguzzi C, et al. A new mtDNA mutation associated with a progressive encephalopathy and cytochrome c oxidase deficiency. Neurology 2000;54:1693–1696. 159. Sohm B, Frugier M, Brule H, Olszak K, Przykorska A, Florentz C. Towards understanding human mitochondrial leucine aminoacylation identity. J Mol Biol 2003;328:995– 1010. 160. Spagnolo M, Tomelleri G, Vattemi G, Filosto M, Rizzuto N, Tonin P. A new mutation in the mitochondrial tRNA(Ala) gene in a patient with ophthalmoplegia and dysphagia. Neuromuscul Disord 2001;11:481– 484. 161. Spinazzola A, Carrara F, Mora M, Zeviani M. Mitochondrial myopathy and ophthalmoplegia in a sporadic patient with the 5698G–⬎A mitochondrial DNA mutation. Neuromuscul Disord 2004;14:815– 817. 162. Steinberg S, Gautheret D, Cedergren R. Fitting the structurally diverse animal mitochondrial tRNAs(Ser) to common three-dimensional constraints. J Mol Biol 1994;236:982–989. 163. Sternberg D, Chatzoglou E, Laforet P, Fayet G, Jardel C, Blondy P, et al. Mitochondrial DNA transfer RNA gene sequence variations in patients with mitochondrial disorders. Brain 2001;124:984 –994. 170 Human Mitochondrial tRNAs 164. Sue CM, Tanji K, Hadjigeorgiou G, Andreu AL, Nishino I, Krishna S, et al. Maternally inherited hearing loss in a large kindred with a novel T7511C mutation in the mitochondrial DNA tRNA(Ser(UCN)) gene. Neurology 1999;52:1905–1908. 165. Suzuki T, Suzuki T, Wada T, Saigo K, Watanabe K. Taurine as a constituent of mitochondrial tRNAs: new insights into the functions of taurine and human mitochondrial diseases. EMBO J 2002;21:6581– 6589. 166. Suzuki Y, Suzuki S, Hinokio Y, Chiba M, Atsumi Y, Hosokawa K, et al. Diabetes associated with a novel 3264 mitochondrial tRNA(Leu)(UUR) mutation. Diabetes Care 1997;20:1138 – 1140. 167. Swalwell H, Deschauer M, Hartl H, Strauss M, Turnbull DM, Zierz S, et al. Pure myopathy associated with a novel mitochondrial tRNA gene mutation. Neurology 2006;66:447– 449. 168. Sweeney MG, Bundey S, Brockington M, Poulton KR, Winer JB, Harding AE. Mitochondrial myopathy associated with sudden death in young adults and a novel mutation in the mitochondrial DNA leucine transfer RNA(UUR) gene. Q J Med 1993;86:709 –713. 169. Tanaka M, Ino H, Ohno K, Hattori K, Sato W, Ozawa T, et al. Mitochondrial mutation in fatal infantile cardiomyopathy. Lancet 1990;336:1452. 170. Taniike M, Fukushima H, Yanagihara I, Tsukamoto H, Tanaka J, Fujimura H, et al. Mitochondrial tRNA(Ile) mutation in fatal cardiomyopathy. Biochem Biophys Res Commun 1992;186:47–53. 171. Taylor RW, Chinnery PF, Bates MJ, Jackson MJ, Johnson MA, Andrews RM, et al. A novel mitochondrial DNA point mutation in the tRNA(Ile) gene: studies in a patient presenting with chronic progressive external ophthalmoplegia and multiple sclerosis. Biochem Biophys Res Commun 1998;243:47– 51. 172. Taylor RW, Schaefer AM, McFarland R, Maddison P, Turnbull DM. A novel mitochondrial DNA tRNA(Ile) (A4267G) mutation in a sporadic patient with mitochondrial myopathy. Neuromuscul Disord 2002;12:659 – 664. 173. Taylor RW, Giordano C, Davidson MM, d’Amati G, Bain H, Hayes CM, et al. A homoplasmic mitochondrial transfer ribonucleic acid mutation as a cause of maternally inherited hypertrophic cardiomyopathy. J Am Coll Cardiol 2003;41: 1786 –1796. 174. Taylor RW, Schaefer AM, McDonnell MT, Petty RK, Thomas AM, Blakely EL, et al. Catastrophic presentation of mitochondrial disease due to a mutation in the tRNA(His) gene. Neurology 2004;62:1420 –1423. 175. Terasaki F, Tanaka M, Kawamura K, Kanzaki Y, Okabe M, Hayashi T, et al. A case of cardiomyopathy showing progression from the hypertrophic to the dilated form: association of Mt8348A–⬎G mutation in the mitochondrial tRNA(Lys) gene with severe ultrastructural alterations of mitochondria in cardiomyocytes. Jpn Circ J 2001;65:691– 694. 176. Tessa A, Vilarinho L, Casali C, Santorelli FM. MtDNA-related idiopathic dilated cardiomyopathy. Eur J Hum Genet 1999; 7:847– 848. 177. Tiranti V, Chariot P, Carella F, Toscano A, Soliveri P, Girlanda P, et al. Maternally inherited hearing loss, ataxia and myoclonus associated with a novel point mutation in mitochondrial tRNASer(UCN) gene. Hum Mol Genet 1995;4: 1421–1427. 178. Tiranti V, D’Agruma L, Pareyson D, Mora M, Carrara F, Zelante L, et al. A novel mutation in the mitochondrial tRNA(Val) gene associated with a complex neurological presentation. Ann Neurol 1998;43:98 –101. 179. Tiranti V, Carrara F, Confalonieri P, Mora M, Maffei RM, Lamantea E, et al. A novel mutation (8342G–⬎A) in the mitochondrial tRNA(Lys) gene associated with progressive external ophthalmoplegia and myoclonus. Neuromuscul Disord 1999;9:66 –71. 180. Tomari Y, Hino N, Nagaike T, Suzuki T, Ueda T. Decreased CCA-addition in human mitochondrial tRNAs bearing a MUSCLE & NERVE February 2008 181. 182. 183. 184. 185. 186. 187. 188. 189. 190. 191. pathogenic A4317G or A10044G mutation. J Biol Chem 2003;278:16828 –16833. Toompuu M, Yasukawa T, Suzuki T, Hakkinen T, Spelbrink JN, Watanabe K, et al. The 7472insC mitochondrial DNA mutation impairs the synthesis and extent of aminoacylation of tRNASer(UCN) but not its structure or rate of turnover. J Biol Chem 2002;277:22240 –22250. Tulinius M, Moslemi AR, Darin N, Westerberg B, Wiklund LM, Holme E, et al. Leigh syndrome with cytochrome-c oxidase deficiency and a single T insertion nt 5537 in the mitochondrial tRNATrp gene. Neuropediatrics 2003;34:87– 91. Tuohy TM, Thompson S, Gesteland RF, Atkins JF. Seven, eight and nine-membered anticodon loop mutants of tRNA(2Arg) which cause ⫹1 frameshifting. Tolerance of DHU arm and other secondary mutations. J Mol Biol 1992; 228:1042–1054. Tzen CY, Tsai JD, Wu TY, Chen BF, Chen ML, Lin SP, et al. Tubulointerstitial nephritis associated with a novel mitochondrial point mutation. Kidney Int 2001;59:846 – 854. Uusimaa J, Finnila S, Remes AM, Rantala H, Vainionpaa L, Hassinen IE, et al. Molecular epidemiology of childhood mitochondrial encephalomyopathies in a Finnish population: sequence analysis of entire mtDNA of 17 children reveals heteroplasmic mutations in tRNAArg, tRNAGlu, and tRNALeu(UUR) genes. Pediatrics 2004;114:443– 450. Van Camp G, Smith RJ. Maternally inherited hearing impairment. Clin Genet 2000;57:409 – 414. van den Bosch BJ, de Coo IF, Hendrickx AT, Busch HF, de Jong G, Scholte HR, et al. Increased risk for cardiorespiratory failure associated with the A3302G mutation in the mitochondrial DNA encoded tRNALeu(UUR) gene. Neuromuscul Disord 2004;14:683– 688. Verhoeven K, Ensink RJ, Tiranti V, Huygen PL, Johnson DF, Schatteman I, et al. Hearing impairment and neurological dysfunction associated with a mutation in the mitochondrial tRNASer(UCN) gene. Eur J Hum Genet 1999;7:45–51. Verma A, Piccoli DA, Bonilla E, Berry GT, DiMauro S, Moraes CT. A novel mitochondrial G8313A mutation associated with prominent initial gastrointestinal symptoms and progressive encephaloneuropathy. Pediatr Res 1997;42:448 – 454. Vissing J, Salamon MB, Arlien-Soborg P, Norby S, Manta P, DiMauro S, et al. A new mitochondrial tRNA(Met) gene mutation in a patient with dystrophic muscle and exercise intolerance. Neurology 1998;50:1875–1878. Vives-Bauza C, Gamez J, Roig M, Briones P, Cervera C, Solano A, et al. Exercise intolerance resulting from a musclerestricted mutation in the mitochondrial tRNA(Leu (CUN)) gene. Ann Med 2001;33:493– 496. Human Mitochondrial tRNAs 192. Vives-Bauza C, Del Toro M, Solano A, Montoya J, Andreu AL, Roig M. Genotype-phenotype correlation in the 5703G⬎A mutation in the tRNA(ASN) gene of mitochondrial DNA. J Inherit Metab Dis 2003;26:507–508. 193. Vos MH, Lipowski G, Lambry JC, Martin JL, Liebl U. Dynamics of nitric oxide in the active site of reduced cytochrome c oxidase aa3. Biochemistry 2001;40:7806 –7811. 194. Weber K, Wilson JN, Taylor L, Brierley E, Johnson MA, Turnbull DM, et al. A new mtDNA mutation showing accumulation with time and restriction to skeletal muscle. Am J Hum Genet 1997;60:373–380. 195. Wilson FH, Hariri A, Farhi A, Zhao H, Petersen KF, Toka HR, et al. A cluster of metabolic defects caused by mutation in a mitochondrial tRNA. Science 2004;306:1190 –1194. 196. Wittenhagen LM, Kelley SO. Impact of disease-related mitochondrial mutations on tRNA structure and function. Trends Biochem Sci 2003;28:605– 611. 197. Wittenhagen LM, Roy MD, Kelley SO. The pathogenic U3271C human mitochondrial tRNA(Leu(UUR)) mutation disrupts a fragile anticodon stem. Nucleic Acids Res 2003; 31:596 – 601. 198. Wong LJ, Liang MH, Kwon H, Bai RK, Alper O, Gropman A. A cystic fibrosis patient with two novel mutations in mitochondrial DNA: mild disease led to delayed diagnosis of both disorders. Am J Med Genet 2002;113:59 – 64. 199. Wong LJC. Pathogenic mitochondrial DNA mutations in protein-coding genes. Muscle Nerve 2007;36:279 –293. 200. Yan H, Zareen N, Levinger L. Naturally occurring mutations in human mitochondrial pre-tRNASer(UCN) can affect the transfer ribonuclease Z cleavage site, processing kinetics, and substrate secondary structure. J Biol Chem 2006;281: 3926 –3935. 201. Yasukawa T, Hino N, Suzuki T, Watanabe K, Ueda T, Ohta S. A pathogenic point mutation reduces stability of mitochondrial mutant tRNA(Ile). Nucleic Acids Res 2000;28:3779 – 3784. 202. Yasukawa T, Suzuki T, Ueda T, Ohta S, Watanabe K. Modification defect at anticodon wobble nucleotide of mitochondrial tRNAs(Leu)(UUR) with pathogenic mutations of mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes. J Biol Chem 2000;275:4251– 4257. 203. Yasukawa T, Suzuki T, Ishii N, Ohta S, Watanabe K. Wobble modification defect in tRNA disturbs codon-anticodon interaction in a mitochondrial disease. EMBO J 2001;20:4794 – 4802. 204. Yoon KL, Aprille JR, Ernst SG. Mitochondrial tRNA(thr) mutation in fatal infantile respiratory enzyme deficiency. Biochem Biophys Res Commun 1991;176:1112–1115. MUSCLE & NERVE February 2008 171