* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download P-Element Transformation with period Locus DNA Restores
Epitranscriptome wikipedia , lookup
Designer baby wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Koinophilia wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genealogical DNA test wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
DNA vaccination wikipedia , lookup
Frameshift mutation wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Genetic engineering wikipedia , lookup
Oncogenomics wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
RNA silencing wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Epigenomics wikipedia , lookup
DNA supercoil wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Non-coding RNA wikipedia , lookup
History of RNA biology wikipedia , lookup
Molecular cloning wikipedia , lookup
Microsatellite wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Helitron (biology) wikipedia , lookup
Non-coding DNA wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microevolution wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
History of genetic engineering wikipedia , lookup
Point mutation wikipedia , lookup
Genomic library wikipedia , lookup
Deoxyribozyme wikipedia , lookup
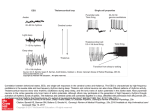
Cell, Vol. 39, 369-376, December 1984 (Part I), Copyright 0 1984 by MIT 0092.8674/84/120369-08 $02,00/O P-Element Transformation with period Locus DNA Restores Rhythmicity to Mutant, Arrhythmic Drosophila melanogaster \ William A. Zehring,* David A. Wheeler,* Pranhitha Reddy,T-Ronald J. Konopka;* Charalambos P. Kyriacou,” Michael Rosbash,* and Jeffrey C. Hall* * Department of Biology + Department of Biochemistry Brandeis University Waltham, Massachusetts 02254 * Department of Biology Clarkson College Potsdam, New York 13676 9 Department of Genetics University of Leicester Leicester LEI 7RH, England Summary Mutations at the period (Per) locus of Drosophila melanogaster disrupt several biological rhythms. Molecular cloning of DNA sequences encompassing the per+ locus has allowed germ-line transformation experiments to be carried out. Certain subsegments of the per region, transduced into the genome of arrhythmic per“ flies, restore rhythmic@ in circadian locomotor behavior and the male’s courtship song. Introduction Biological rhythms are ubiquitous and have been intensively studied at many levels. Yet almost nothing is known about the molecular nature of the “clocks” underlying such rhythms. The isolation of mutations that perturb or abolish rhythms (reviewed by Feldman, 1982) has allowed molecular analyses of these phenomena. The period (per) locus of Drosophila melanogaster, discovered by Konopka and Benzer (1971) is the first gene that was mutated to disrupt biological rhythms. Different mutant alleles at this X-chromosomal locus can lengthen (per” and perLq, shorten (per”), or abolish (per”’ and per”) circadian rhythms (reviewed by Konopka, 1984). These mutations similarly affect a short-term oscillation in the male’s courtship song (Kyriacou and Hall, 1980). Therefore, this locus appears to be centrally involved in more than one of the fly’s rhythms. The per locus has been extensively analyzed genetically and mapped to the 3B1-2 region of the X chromosome (e.g., Smith and Konopka, 1981). We and others have begun molecular analysis of the per region (Reddy et al., 1984; Bargiello and Young, 1984). Some of our previous findings are summarized in Figure IA, including the molecularly mapped sites of certain chromosomal lesions near per. One of these-a breakpoint in the T(1;4)JC43 translocation, which causes aberrant circadian rhythms (Smith and Konopka, 1981)-appears to be in the center of the per region. This breakpoint also leads to lower than normal levels of certain transcripts (Reddy et al., 1984; Bargiello and Young, 1984) whose genomic sources are also depicted in Figure IA. We have suggested that the 0.9 kb RNA species (see large circle in Figure IA) is of special significance because of its abnormally low levels, not only in flies hemizygous for the T(7;4)JC43 breakpoint but also in both peP’ and per”’ adults (Reddy et al., 1984). In addition, the abundance of this RNA species fluctuates dramatically over the course of circadian cycles (Reddy et al., 1984). Against this background, we developed a behavioral assay for per+ DNA in order to probe the biological activity of defined portions of this clock locus. Two of the most extensively characterized behavioral phenotypes known to be affected by per mutations are circadian rhythms of locomotor activity and rhythmic oscillations in the interpulse intervals of the male’s courtship song. Using per” hosts that are arrhythmic for both of these behaviors (Konopka and Benzer, 1971; Kyriacou and Hall, 1980) we carried out a series of P-element-mediated transformations involving DNA obtained from the per region of wild-type flies. We initially intended (and began experiments) to scan this region by testing a series of adjacent and overlapping restriction fragments without any predisposition as to which segment or segments might rescue the effects of this mutation. For reasons referred to above (cf. Reddy et al., 1984) we decided to concentrate on DNA fragments that include or are near the site of the T(7;4)JC43 381-2 breakpoint. We now report that either of two overlapping DNA fragments, transduced into an arrhythmic per’ host, rescues behaviorally mutant phenotypes. This is the first identification, by a transformation bioassay, of a functioning part of a metazoan’s genetic locus that could not be molecularly identified from a priori knowledge of a gene product. Results Circadian Rhythms of Transformants Table 1 summarizes behavioral and genetic data on the transformants. Transformation with two of the DNA fragments shown in Figure IA rescued per”‘-induced arrhythmicity of locomotor behavior. These clones came from a portion of the per region that crosses the T(1;4)JC43 breakpoint. One such fragment, 8.0 kb in length, is a subset of the other rescuing DNA segment, which extends 6.6 kb farther to the right (Figure IA). The majority of the individual flies hemizygous for per”, carrying either the 14.6 or 8.0 kb fragment, were rhythmic in their locomotor activity. Moreover, such circadian rhythms were detected in adults from virtually all of the independently isolated “14.6” or “8.0” strains (Table 1). Yet these phenotypes were abnormal in two ways: the proportion of rhythmic cases was relatively low, compared to the very high fraction of wild-type flies that exhibit periodicity in their locomotor activity (Table 1, cf. Konopka and Benzer, 1971; Smith and Konopka, 1981); and the circadian periods for the transformations (examples shown in Figure 2) were routinely longer than the normal 24 hr. In some cases, per” flies carrying an 8.0 kb insert seemed to show at least two cycles of long-period rhythmicity, which then graded into Cdl 370 1981) a pattern of phenotypes similar to those expressed by the “8.0” or “14.6” transformants. D/I/IX95 1.7Kb Adh’XbaI Fragment 100 bp PolyLinker ECoRI BqllI Figure 1. Molecular Map of Subclones in the par Region Used in Transformation, with Reference to Chromosomal Breakpoints and Transcripts from This Region (A) Restriction sites, cut by the endonucleases Eco RI (R, or @ for an artificial, nongenomrc cloning site), Barn HI (B), and Hind III (H) are shown on the horizontal line near the bottom. The genomic sources for the 4.5, 1 .O, 0.9, 2.7, and 1.7 kb RNA species are displayed below this line. See Reddy et al. (1984) for the relevant Northern-blot analyses, including a determination of the directions of transcription, indicated here by the arrowheads; the boundary between genomic subsegments hybridizing to the 2.7 versus the 1.7 kb transcript was marked by a Pst I restriction site. The locations of chromosomal breakpoints associated with the T(7;4)JC43, Df(l)K95, and Df(l)w- @’ aberrations are as described in Reddy et al. (1984) with the latter two lesions indicated here by shaded bars; horizontal lines connected to the bars indicate DNA, as opposed to its absence, which refers to the region to the right of the per-region breakpoint in the Df(7)w-Hddeletion; the Df(l)K95 lesion depicted (with lines running in both directions from it) is an inversion breakpoint associated with this deletion (M. Goldberg, personal communication, and see Reddy et al., 1984). The two Df breakpoints shown may define the boundaries of the per region, since both of these aberrant genotypes complement per mutations (Smith and Konopka, 1981). The left end of this map is distal on the X chromosome (i.e., farthest from the centromere). Solid bars indicate the positions of fragments conferring rhythmicity when transformed into per”’ hosts, whereas open bars indicate regions that did not transform par“’ hosts to rhythmrcity (see Table 1). (B) Map of the pPA-I transformation vector (J. Posakony, unpublished), containing the wild-type allele of the alcohol dehydrogenase (Ad/t+) gene of D. melanogaster and a 100 base pair polylinker located between Adh+ and P-element DNA. arrhythmic behavior (- -> Arrh. in Table 1). This is a phenotype similar to that expressed by certain individuals which are hemizygous for the T(1;4jJC43 per-region breakpoint (Smith and Konopka, 1981). This same genotype also causes stable long-period rhythmicity or complete arrhythmicity for a given individual (Smith and Konopka, Courtship Song Rhythms of Transformants The 14.6 and 8.0 kb subsegments of the per region also rescued the arrhythmic courtship singing behavior of perof males (cf. Kyriacou and Hall, 1980). That is, most of the males carrying either type of insert exhibited regular oscillations of the interpulse intervals (IPls) computed from visual displays (see Figure 3A) of the acoustical output of the wing vibrations they direct at females. The rescue of this short-term song rhythm had a somewhat higher overall penetrance in both sets of transformants than the circadian phenotype (Table 1). Six of the males (carrying 14.6 kb inserts) whose songs were recorded and analyzed had previously been tested for circadian rhythmic@. Three of them were rhythmic in both locomotor and singing behavior, two were arrhythmic for both characters, and one exhibited no circadian rhythm but subsequently sang in a rhythmic manner. As in the case of circadian rhythms, the song rhythm periods were longer than normal (i.e., -70-80 set instead of the usual per+ value of -55-60 set, cf. Kyriacou and Hall, 1980, 1984) in perof males transformed with either a 14.6 or 8.0 kb fragment from the per region (Table 1, Figure 3B). In one case, the entirety of the male’s -6 min song was not rhythmic, but three strong cycles of regularly oscillating IPls (with a period of -65 set) were computed; the functions applied to these data (cf. Kyriacou and Hall, 1980) showed that the remaining 3 min of IPI fluctuations was arrhythmic (- -> Arrh. in the right-hand part of Table 1). This phenotype is analogous to that exhibited by certain of the transformed individuals monitored for circadian rhythms of locomotor activity (left-hand part of Table I), as described above. Nonrescuing per-Region Subsegments Subsegments of the per region other than the 14.6 and 8.0 kb fragments did not rescue mutant behavioral phenotypes The most extensively analyzed “negative” case is the 9.8 kb fragment (Figure IA); analysis of the relevant transformants showed only one feebly rhythmic animal out of 32 tested (Table 1). For the other per-region subsegments, however, there was only one transformed line (per fragment) available for behavioral testing. Also, the courtship song has not been as extensively analyzed as locomotor activity (Table 1). Discussion Specific Implications of the Transformants’ Phenotypes The most straightforward interpretation of our positive behavioral results from the transformants is that the genomic change in the perO’ mutant is located within the 8.0 kb subsegment of the per region. Since flies from the transformed lines in which the phenotypes caused by peP are ameliorated express f 3 and Behavioral Analysis 26.7 27.8 27.0 : 33 0 14 __ Cl 0 0 5 27.3 26.0 2 * *t f i f 0.1 0.8 0.9 00.3 II 0.1 23.8 ca. _- * _- -_ __ _- 27 0.1 Number Arrhythmic Involving per-Region 28.5-->Rrrh. ca. 2,-->Acfh. _ca. 28 ca. 28-->Arrh. ca. 23 26.0 27.4 3 14 of Transformants umber trong1y __ 24.0 14 0 __ __ __ _- 0 0 0 0 -_ __ __ __ __ 25.9 8 0 0 __ __ _- 24.8 3 + + + __ + + f + + f __ _- 27.0 + 25.0 26.7 25.3 27.3 26.8 26.5 27.6 0.1 0.6 0.6 0.2 0.3 0.4 0.2 1.3 0.4 1.0 Period (h f SE") man 1" 29 7 6 4 5 7 __ hythmic II DNA - 8 ” R 0 0 0 1 __ 0 1 __ __ 7 __ __ 5 0 2 2 1 __ : .c 24.3 ca. ca. 25.1 -_ + __ __ -- __ __ __ __ f f __ __ 0.3 0.6 1.3 0 0.4 0.2 22 22 25.5 + 27.0 __ 23.3 f+ 30.3 Period lh * SEMI 24.9 Locomotor Activity (periodogramsl 3 10 0 12 8 8 : __ 24 : -_ -_ -- __ + __ _- 68.2 __ 0 7 7.3 -- 0 * 2 0 3 1 -- __ __ * __ 1 1 __ -- 1 Y __ __ 0 -- 0 0 __ The transformed strains are named (leftmost column) with respect to the lengths In krlobases of per-region subsegments (see Frgure IA) used to produce me vectors that allowed generation of the transformants, Each strain also is named by the addition of an arbitrary number to desrgnate it as independently isolated line (i.e., more than one strain was usually isolated from the series of injections involving a given per-region fragment). Orientations of per-region inserts (in the transformation vectors and in the genome of transformants) are indicated in the first column as follows: for the 14.8, 7.3, and 8.0 kb inserts, (a) indicates the 0.9 kb RNA is transcribed antiparallel to the Adh+ RNA, and (b) indicates the 0.9 kb RNA is parallel to the Adh+ RNA; for the 9.8 kb Inserts, (a) indicates the 2.7 kb RNA is antiparallel to the Adh+ RNA; for the 5.8 kb insert (b) indicates the 1.7 kb RNA is parallel to the Adh+ RNA; for the 9.0 kb insert (a) indicates the 4.5 kb RNA is antiparallel to the Adh+ RNA (see Figure 1B and Experimental Procedures), The genomic locations of the insert (or inserts) for each strain are listed in the second column; some transformants carry at least one insert on two or more separate chromosomes. Data from many of the activity tests were collected as actograms (see Figure 2) and assessed for rhythmicity versus arrhythmicity as in Konopka and Benzer (1971). The periods were estimated as described rn Expenmental Procedures. The results of the periodogram analyses of circadian rhythmicity (see Experimental Procedures) were divided into three categories: strongly rhythmic (a robust and wall defined peak in the perk&gram, corresponding to a given period which was determined to the nearest half-hour); weakly rhythmic (a relatively small peak, due to a somewhat sloppy rhythm, a rather inactive fly, or both); and arrhythmic (no discernible peak). The overall rhythmicity of the “8.0” strains was stronger, as a result of the periodogram analyses (see Experimental Procedures), than from the appearance of the actograms. Yet an inspection of the digital data (number of activity events in SUCCSSSive time frames), to which the periodogram functions were applied in analyzing these strains, led to the somewhat subjective impression that these flies expressed weaker circadian rhythms (digitally determined) than in the wild type. That is, the overall pattern of results on the “8.0” transformants seemed, from inspection of both kinds of relevant data (analog or digital), to be similar, In the song tests, rhythmicity (or lack of it) in the fluctuations of pulse song IPls was determined as described in Kyriacou and Hall (1980) summarized here in Experimental Procedures and the legend to Figure 3. 9.8:9(a) 9.8:10(a) 9.8:16(a) 9.8:26(a) 9.8:*9(a) Table 1. Genetic Cell 372 perti; P[ per’] Adhf”23pr cn .- 14.6:21 I . I ~~26.5 h ~=27 h ...z... -. perd;P[per’] Adhf”23pr cn 14.6:63 . I-. ~=26 h rx23.7 -_- h per” Figure 2. Actograms of Locomotor Activity The genotypes of the transformants shown (as opposed to the per + and per” controls) are according to the nomenclature prescribed in Spradling and Rubin (1983). In the actograms time moves from left to right and from top to bottom (see Expenmental Procedures). The data displayed represent recordings made during freerunning (Le., after entrainment of the adults by light/dark cycles). The black bar in the center of each set of traces corresponds to the time when the lights had been turned on during each of the three entrainment days. The normal (per’) fly had activrty offsets and onsets, which are arranged rn an approximately vertical manner (slope -0, see Experimental Procedures), near the beginning and the end (respectively) of the subjective day (hence 7 = -24 hr). However, the activity offsets for the transformants move progressively to the right on successive days; these shifting offsets, which are later and later than in the normal case, therefore define a positively sloping line (see Experimental Procedures), and this is reflected in the relatively long periods (8s) depicted here for the transformants from different “14.6” strains (also see Table 1) either long-period rhythms or are arrhythmic, we infer that the expression of per-region functions is subnormal in these two types of transformants. This conclusion is based on the supposition that long-period perL variants seem to be hypomorphic mutations (summarized in Konopka, 1984) and that hemizygosity for the per-region breakpoint in 7(7;4)JC43 leads to an overall array of locomotor activity phenotypes not unlike those exhibited by our transformants. The chromosomally aberrant genotype, moreover, causes hypomorphic expression of three transcripts from the per region (Reddy et al., 1984). Hypomorphic expression of per can also be achieved simply by constructing females that are heterozygous for peP’, or a deletion of the locus, and per+. In these cases, circadian periods are slightly longer than normal (-25 hr), but essentially every individual is rhythmic (Konopka and Benzer, 1971; Smith and Konopka, 1981). We infer that the array of behavioral phenotypes expressed by our transformants is not due merely to the fact that the various transformed flies carried only one dose of transduced per+ DNA (also, some of the flies we tested carried more than one dose, but were still long-period; see Experimental Procedures). We do not have enough molecular information to interpret the apparently subnormal activity of the two rescuing DNA segments in the transformants. One possibility is that sequences adjacent to these pieces of DNA are necessary for completely normal per+ expression. It is also conceivable that the various genomic sites at which the 8.0 and 14.6 kb fragments from the per region have integrated ;raFformation of per IPI A lead to subnormal expression; this would include the possibility that autosomal locations for the inserts (i.e., the great majority of the cases listed in Table 1) are somehow associated with relatively poor transcription of the relevant RNAs, which are normally transcribed from the X chromosome. It is known, however, that ectopic locations for autosomal enzyme-encoding loci in D. melanogaster usually (Scholnick et al., 1983; Spradling and Rubin, 1983) but not in every case (Goldberg et al., 1983) allow for nearly normal expression of these transduced genes; and that autosomal DNA inserts of the (usually X-chromosomal) white (w) locus typically-but, again, not always-give apparently normal levels of w+ expression (Hazelrigg et al., 1984). The failure of a given fragment to affect the behavior of the per0 hosts must be interpreted with caution. For example, the 9.8 kb fragment fails to rescue the per“’ mutation; thus the 2.7 kb RNA species (Figure 1A) would seem to be eliminated as a candidate for the transcript that is directly affected by per” mutations. Yet the various RNAs transcribed from this region have not been mapped with enough precision to show whether an individual segment of the per region contains all of the coding (and noncoding) information required to express properly a given transcript. Experiments to assess the capacity of the various transduced DNA fragments to express, in vivo, individual transcripts are in progress. The results of this study, taken together with our previous findings (Reddy et al., 1984) suggest that either the 0.9 kb transcript or the 1.O kb RNA species (see Figure 1A) is the transcript directly affected by peP’, and that this mutational defect is rescued in Pans by introduction into the genome of wild-type DNA homologous to either or both of these transcripts. We propose further that the 1 .O kb transcript may be the key RNA species; this is because the 7.3 kb DNA fragment-which looks as if it should be able to produce the 0.9 kb RNA but not the 1 .O kb RNA (see Figure lA)-did not rescue per”‘-induced arrhythmicity (although only one such “7.3” strain was recovered, Table 1). This speculation is correlated with, and influenced by, previous genetic and behavioral data implying that Xchromosomal material to the left of the T(l;4)JC43 breakpoint (to which the 1.0 but not the 0.9 kb transcript is homologous) is sufficient to confer weak, long-period rhythmicity in circadian behavior (Smith and Konopka, 1981). If the 1 .O kb RNA is the species affected directly by peP’ and in need of rescue, our present results and those reported earlier (Reddy et al., 1984) imply that the 100 ms B Per “‘; P[pw+] Adbfnz3prcn 14.6~63 r=66.1 s 46r per” P[perf];Adhfn”Prcn per”‘; P [per + 8.0~2 Adhfnz3prcn; P[perf] r=63.1 s 9.8:lO cl 48 0 0 36 Figure 3. Oscillatrons of Interpulse min Intervals in the Male’s Courtshrp Song Males of the indicated transformed genotypes (cf. Table 1 and Figure 2) had their songs recorded and analyzed as described in Experimental Procedures. (A) An example of a pulse train, with eight tone pulses, hence seven interpulse intervals (IPls), is depicted. In this species of Drosophila, pulses are typically generated at the rate of -20-30/set (e.g., Kyriacou and Hall, 1980). (6) The two rhythmic songs (for a male from a “14.6” transformed line, and one from an “8.0” line) are indicated by the sinusoidal (dashed) lines, accompanied by the best estimates of the periods (T’S). No line was drawn with respect to the IPI fluctuations for the male from a “9.8” transformed lrne (cf. Table I), because there was no discernible pattern to these changing IPls. Cell 374 dramatic effects of arrhythmic per mutations on the abundance of the 0.9 kb transcript are not the primary cause of arrhythmicity. More complex models are possible and cannot be excluded at present. Among these is the possibility that the per”’ mutation does not map within the 8.0 kb subsegment of the per-region but has an adverse effect in cis on a transcript or transcripts coded for by this subsegment of the per region. Such a model implies that one or more transcripts coded for by this fragment (presumably the 1 .O and/or the 0.9 kb RNA species) are able to provide for rhythmic behaviors when the 8.0 kb segment is not adjacent to the per"' mutation. Therefore, the site of this mutation would not necessarily define a portion of the per locus that, in the wild-type, is required for downstream regulation of the transcription involved in the control of normal rhythmicity. This possibility (and others) can be examined by molecularly defining the site of the peror mutation. General Implications of the Molecular Expression of the per Locus As for other biological rhythms in Drosophila-and their regulation by expression of the per locus-no single transcript seems sufficient to control all the relevant phenotypes For instance, erratic heartbeating has been detected in perO’ larvae (E. 0. Aceves-Piha and M. S. Livingstone, unpublished; see Livingstone, 1981). The RNAs transcribed from the portions of the per region immediately adjacent to the T(1;4)JC43 breakpoint (Figure 1A) would appear to contain inadequate information to influence this phenotype, because none of these transcripts is detectable earlier than the late pupal stage of development (Reddy et al., 1984; Bargiello and Young, 1984). Two of the other per-region transcripts shown in Figure IA, the 2.7 and 1.7 kb species, are present during all stages of the life cycle, though only the former RNA is detectable in the fly’s head (see Reddy et al., 1984, for discussion of this and other evidence suggesting that the 1.7 kb transcript may be associated with the expression of a separate gene, located to the right of per, called /(l)zw6). Therefore, the 2.7 kb transcript is the only obvious candidate for participating in the control of rhythmic phenotypes that are entrained or expressed relatively early in development and are also disturbed by per mutations (see further discussion of these several phenotypes in Konopka, 1984). Another rhythmic phenomenon associated with the per locus, the light/dark-mediated fluctuation in the 0.9 kb transcript’s abundance (Reddy et al., 1984) is almost certainly related to the fly’s circadian rhythms of locomotor activity. However, the correlation of this molecular phenotype with the male’s song rhythms is ostensibly obscure, since that oscillating behavior is insensitive to changing environmental cues (Kyriacou and Hall, 1980). For these reasons, and because of the molecular complexity of the per region’s transcripts, we suspect this locus may contribute more than one functional product to the control of the fly’s biological clocks, Consequently, transformation experiments must be extended to an analysis of various other per-mutant phenotypes, such as eclosion and the larval heartbeat. Also, additional transformed lines must be generated both by injection of as yet unanalyzed portions of the per region and by genetic and molecular construction of various combinations of DNA fragments. The results presented here represent an important first step in this extensive program and lead to the strong prediction that transcripts (and proteins?) coded for by the 8.0 kb subset of the per locus are components of the control of Drosophila’s biological clocks. Experimental Procedures Construction of Transformation Vectors In preparation for generation of the transformants, overlapping Eco RI or Barn HI restriction fragments were ligated to the P-element transformation vector pPA-1 (Figure 1 B, J. Posakony, unpublished) that had been linearized by digestion with either Eco RI or Barn HI. Ligated DNA was transformed into the DHl strain of E. coli. Plasmid DNA was extracted from ampicillinresistant colonies and analyzed by restriction digestion as described by Reddy et al. (1984). Restriction endonucleases and T4 DNA polymsrase were purchased from New England Biolabs and Bethesda Research Labs and used as recommended by the manufacturers. By using cuts made in the polylinker of the transformation vector (Figure IB), clones containing the fragments indicated in Figure IA were inserted in the two possible orientations with respect to the Adh gene; the orientations were confirmed by analysis of restriction digests, Generation of Transformants The transformants were produced by microinjection of cloned DNA (cf. Spradling and Rubin, 1982; Rubin and Spradling, 1982). We used a series of transformation vectors in which the separate DNA fragments from the per region (Figure 1A) had been cloned into the plasmid shown in Figure 1B. All DNA transformation cocktails, with the exception of that involving the 5.8 kb fragment, also contained the helper P-element plasmid p?r25.7wc, which is able to provide “transposase” function but is unable to insert into the D. melanogaster genome (Karess and Rubin, 1984). The “5.8” transformations utilized the helper P-element plasmid pr25.1, able both to provide transposase function and to be inserted into a chromosome. For both types of helper plasmid, this DNA was present at a concentration of 50 ag/ml. In the cases of transformations involving the 5.8, 7.3, and 9.0 kb fragments (Figure IA), roughly equal amounts of the per-recombinant plasmids, containing single inserts in both possible orientations, were present in the transformation cocktails at a final concentration of 500 pg/ ml. The 9.8 and 8.0 kb fragments were cloned into pPA-I (Figure 1B) in each orientation, but injections were carried out separately with each orientation of the insert, The 14.6 kb fragment was inserted in only one orientation, with the 0.9 kb RNA antiparallel to the Adh+ RNA. In all these cases, where a single orientation of the fragment in the vector was injected, the DNA was again at 500 pg/ml. Each transformant line was examined by Southern blotting (data not shown) to determine the orientation of the inserted DNA wrth respect to the Aclh+ gene present in the vector (see Table 1). Microinjection of transforming DNA was carried out using the techniques of Rubin and Spradling (1982) as applied by Goldberg et al. (1983). The latter authors rescued the effects of an alcohol-dehydrogenase-null (Adh’“23) mutation (cf. Benyajati et al., 1983). Our injections were performed on embryos homozygous for this mutation (linked to the eye color markers pr and cn); in most cases, the injectees were also homozygous or hemizygous for per”‘. Adults were collected as they emerged rn the cultures that resulted from the development of the injected embryos. Such flies were mated srngly to pep’; Adh’“23 pr cn (or, when appropriate, Adh”‘23 pr cn) adults to produce a series of strains in the next generation (called Gl). These Gl strains were screened for acquired resistance to a 6% ethanol-water solution in a test descrrbed by Vigue and Sofer (1974). It was important to age the adults for 4 days before ethanol treatment (to avord the misleadrng sun4val of nontransformed Adh”‘= individuals). Transformation 375 of per Basic Characterization and Manipulation of Transformants Strains recognized to contain Adh+ inserts have been maintained by regular selection for ethanol resistance, but were behaviorally tested soon after therr isolation. The chromosomal location of each insert was determined by applying standard genetic techniques for assessment of X linkage versus autosomal inheritance. In the latter cases, crosses involving autosomaldominant-containing balancer chromosomes led to an assignment of 2nd. versus 3rd.chromosomal linkage of the insert(s) for a given strain, using ethanol selection and spectrophotometric assays (re ADH activity, cf. Ursprung et al., 1970) of the appropriate segregants. For the transformants carrying the 5.8, 7.3, and 9.0 kb fragments, which were initially obtained in a per’ background, it was necessary to cross individual ethanol-resistant males to per”‘; Adh “‘23prcn females and select ethanol-resistant male progeny (hemizygous for per”) prior to behavioral testing. All other transformations were carried out in the per”‘; Adh”‘*3 pr cn genetic background. For all the transformed strains, the presence of the insert from the per region was confirmed (data not shown) by Southern blot analysis, using appropriate probes (Reddy et al., 1984) and techniques similar to those previously reported (e.g. Goldberg et al., 1983; Hazelrigg et al., 1984). After selecting individual Adh+ males from a given strain containing an insert from the per region, these flies were monitored for circadian rhythms for locomotor activity or were recorded with respect to courtship singing behavior, as described below. Analysis of Circadian Rhythms Single male flies were selected (if appropriate, see Table 1) and exposed to 3 days of entrainment in light/dark cycles of 12 hr/12 hr. At the end of the third entrainment day, a given transformed or control fly was marntained for 4-7 additional days in constant darkness, when the “freerunning” endogenous rhythms (if any) of locomotor activity are expressed. For each Individual fly, activity was monitored by methods similar to those previously reported (Konopka and Benzer, 1971; Smith and Konopka, 1981; Jackson, 1983). An infrared light beam was passed through the short axis of a cylindrical glass tube (15 mm long X 4 mm diameter), and a photoelectric cell picked up any interruptions in the light beam caused by the fly moving Into it. These momentary signals were amplified and recorded on chart paper as a function of time. In addition, for many of the data processed and then summarized in Table 1, activity events were recorded digitally as described in Smith and Konopka (1981) and Sulzman (1982). Subsequent to collection of data by the event recorders, markings on the chart paper were analyzed by cutting strips corresponding to 24 hr segments and placing each subsequent strip in a vertical column (cf. Figure 2). During entrainment, per”’ (as well as wild-type) adults are rather rhythmic in their activity (data not shown in Figure 2) in that the flies are simply more active rn response to light (hence these behavioral fluctuations are not, strictly speaking, endogenous rhythms). The period length (in hours) for a given fly was defined as 7 = 24 + s, where s is the slope (in hours per day) of a line drawn so that it passes approximately through the activity “offsets” during freerunning. An offset is the termination of densely recorded markings within a given rest/activity cycle. By convention, the lines were drawn through these offsets with the chart paper strips oriented vertically (so that offsets occurring about the same time in each 24 hr time frame define azero slope, and those occurring later and later in successive cycles define a positive slope; cf. Figure 2). Most of the flies tested behaviorally were not known to be heterozygous or homozygous for an insert or inserts. Yet, in some cases, individuals were subsequently outcrossed to Adhmz3 pr cn flies, and the Fi tested for ethanol resistance. Two such crosses (involving “14.6” transformants) revealed that the behaviorally tested fly had been homozygous for an insert (or inserts); yet the behavioral phenotypes (longer than normal periods, i.e., both 27 hr) were similar to those determined for other “14.6” individuals, whose progeny revealed that they had carned only one “dose” of an Adh+containing autosome (mean T, for 17 rhythmrc flies, 26.9 f 0.2 hr; arrhythmic, n = 9). In conjunction with tests of circadian rhythmrcity of ethanol-treated transformants, the same treatments of per+; Adh+ adults (from a CantonS strain, which was the source of normal controls and from which the original per mutations were derived, Konopka and Benrer, 1971) showed that the treated flies were rhythmic in either behavioral character, with no period lengthening (cf. such effects, in other organrsms, when they are monitored for circadian rhythms rn the chronic presence of alcohol; eg, Biinnrng and Moser, 1973). Additional controls showed that the Adh’“23 pr cn chromosome (when homozygous or heterozygous with a Canton-Sderived 2nd chromosome) had no effect on normal circadian rhythmicity (in a per+ background) or on the arrhythmicity induced by per”‘. A few control transformants were obtained in which the hybrid P-element/Adh+ vector (see Figure lB), with no per-region insert, was injected: individuals from the resultant ethanol-resistant strains (when hemizygous for per”‘) tested as arrhythmic (data not shown). The activity data collected digitally (see above) were analyzed using periodogram functions (cf. Smith and Konopka, 1981; Sulzman, 1982) to determine objectively whether the activity of a given fly was rhythmic and to obtain the best estimate of the circadian period. In the preliminary tests of this type, the data were collected and the periodograms generated with no knowledge of whether the transformed strains in question appeared, from the actogram analyses, to contain rhythmic flies (Le., these tests of the digital data were done “blrnd”). Analysis of Courtship Song Rhythms Before being tested in courtship, males were aged, ethanol treated (see above), and in some cases monrtored for circadian rhythms. Subsequent to these treatments and tests, males were maintained singly for one more day in unyeasted food vials, and then each was paired with a single, young, virgin female from a wild-type (Canton-S) stock. Courtship typically began shortly after Introduction of the two flies into the recording chamber, consistrng of a small, nylon-mesh-covered plastic cell that was placed close to an electret microphone; thus device had been modrfred to be sensitrve to the “parkcle velocrty” component of sound waves (see Bennet-Clark, 1984, for brologrcal and electronic details). Sounds produced by the courting male’s wrng vibratrons were amplified and recorded (on magnetic tape) for -4-7 mm of courtship. The signals were then transferred to oscillograph paper, on which pulse trains appeared as exemplrfred in Frgure 3A. Analysrs of the successrve Intervals between pulses (IPls) was carried out by the method of Kyrracou and Hall (1980) in which a mean for all the IPls occurrrng within a given 10 set time frame IS computed. The means for successrve frames were plotted (as shown rn Figure 38) wrth the provrso that at least 15 pulses were generated by a male srnging durrng that particular 10 set period. Srnusordal curves (dashed lines In Figure 38) with the best-fitting periods and amplrtudes, were drawn only if a nonlinear regressron analysis and accompanying F-ratio statistic revealed that the IPI fluctuations were significantly rhythmic at the 5% level or beyond. Thus krnd of analysis typically reveals that most, but not all, fires expressing per+ sing with regularly oscillating IPls (Kynacou and Hall, 1980, 1984). All of the present data collectrons and analyses (for transformants and controls) were performed “blrnd,” such that the genotype associated with each song was decoded only after a formal assessment of rhythmicity versus arrhythmicity. Acknowledgments We thank J. Posakony for the P-element/Adh+ vector, the ~~25.1 plasmid, and the Adhfiz3 stock of Drosophrla melanogaster. We are grateful to G. Rubin for the pa257wc plasmid. We greatly appreciate electronic aid from D. Wight, H. Bennet-Clark, and M. Gorczyca and are extremely thankful for the data acquisition and analysis software donated by F. Sulzman. We thank J. Posakony, A. James, and M. Gray for comments on the manuscript. This work was supported by National Institutes of Health grant GM-33205 to M. R. and J. C. H., by a Clarkson College grant to R. J. K., and by SERC grant 94595 to C. P. K.; W. A. Z. is supported by a National Institutes of Health postdoctoral fellowship, and P. R. by a Dretzin predoctoral fellowship. The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 U.S.C. Section 1734 solely to indicate this fact. Recerved September 7, 1984 References Bargrello, T. A., and Young, M. W. (1984). Molecular genetics of a biological clock in Drosophila. Proc. Nat. Acad. Sci. USA 81, 2142-2146. Cell 376 Bennet-Clark, small Insects 463. J. C. (1984). A particle velocity microphone for the song of and other acoustic measurements. J. Exp. Biol. 108, 459- Benyajati, C., Place, A. R., Wangh, N., Pent& E., and Sofer, W. (1983). Formaldehyde mutagenesis in Drosophila: molecular analysis of ADHnegative mutants. Mut. Res. 7 17, 1-7. Bunning, E., and Moser, I. (1973). Light-induced phase shifts of circadian leaf movements of Phasaohx comparison with the effects of potassium and of ethyl alcohol. Proc. Nat. Acad. Sci. USA 70, 3387-3389. Feldman, J. F. (1982). Genetic Plant Physiol. 33, 583-608. approaches to circadian clocks. Ann. Rev. Goldberg, D. A., Posakony, J. W., and Maniatis, T. (1983). Correct developmental expression of a cloned alcohol dehydrogenase gene transduced into the Drosophila germ line. Cell 34, 59-73. Hazelrigg, T., Levis, R., and Rubin, G. M. (1984). Transformation of white locus DNA in Drosophila: dosage compensation, zeste interaction, and position effects. Cell 36, 469-481. Jackson, F. R. (1983). The isolation of biological rhythm mutations autosomes of Drosophila melanogasfer. J. Neurogenet. 7, 3-15. Karess, R. E., and Rubin, G. M. (1984). Analysis functions in Drosophila. Cell 38, 135-146. Konopka, R. J. (1984). Neurogenetics Evolutionary Genetics of Invertebrate York: Plenum Press), in press. of P transposable on the element of Drosophila circadian rhythms. In Behavior, M. D. Huettel, ed. (New Konopka, R. J., and Benzer, S. (1971). Clock mutants melanogaster. Proc. Nat. Acad. Sci. USA 68, 2112-2116. of Drosophila Kyriacou, C. P., and Hall, J. C. (1980). Circadian rhythm mutations in Drosophila melanogasfer affect a short-term fluctuation in the male’s courtship song. Proc. Nat. Acad. Sci. USA 77, 6929-6933. Kyriacou, C. P., and Hall, J. C. (1984). Action potential mutations biological clock in Drosophila. Nature, in press. stop a Livingstone, M. S. (1981). Two mutations in Drosophila affect the synthesis of octopamine, dopamine, and serotonin by altering the activities of two amino-acid decarboxylases. Neurosci. Abstr. 7, 351. Reddy, P., Zehring, W. A., Wheeler, D. A., Pirrotta, V., Hadfield, C., Hall, J. C., and Rosbash, M. (1984). Molecular analysis of the period locus in Drosophila melanogaster and identification of a transcript involved in biological rhythms. Cell 38, 701-710. Rubin, G. M., and Spradling, A. C. (1982). Genetic transformation Drosophila with transposable element vectors. Science 278, 348-353. of Scholnick, S. B., Morgan, B. A., and Hirsh, J. (1983). The cloned dopa decarboxylase gene is developmentally regulated when reintegrated into the Drosophila genome. Ceil 34, 37-45. Smith, R. F., and Konopka, R. J. (1981). Circadian clock phenotypes of chromosomal aberrations with a breakpoint at the per locus. Mol. Gen. Genet. 783, 243-251. Spradling, A. C., and Rubin, G. M. (1982). Transposition of cloned P elements into Drosophila germ line chromosomes. Science 278, 341-347. Spradling, A. C., and Rubin, G. M. (1983). The effect of chromosomal position on the expression of the Drosophila xanthine dehydrogenase gene. Cell 34, 47-57. Sulzman, F. M. (1982). Microcomputer Comput. Biol. Med. 72. 253-261. monitoring of circadian rhythms, Ursprung, H., Sofer, W., and Burroughs, N. (1970). Ontogeny and tissue distribution of alcohol dehydrogenase in Drosophila melanogaster. Wilhelm Roux’s Arch. 164, 201-208. Vigue, C., and Sofer, W. (1974). Ad/?“? a temperature-sensitive the Adh locus in Drosophila. Biochem. Genet. 74, 127-135. mutant at