* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Friesland Foods Normal
Molecular ecology wikipedia , lookup
Ligand binding assay wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Signal transduction wikipedia , lookup
Gene nomenclature wikipedia , lookup
Magnesium transporter wikipedia , lookup
Biochemical cascade wikipedia , lookup
Proteolysis wikipedia , lookup
Point mutation wikipedia , lookup
Gene desert wikipedia , lookup
Genomic imprinting wikipedia , lookup
Ridge (biology) wikipedia , lookup
Community fingerprinting wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Gene expression wikipedia , lookup
Biosynthesis wikipedia , lookup
Gene expression profiling wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene regulatory network wikipedia , lookup
Genome evolution wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Transcriptional regulation wikipedia , lookup
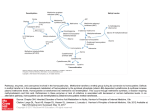
Cysteine and methionine metabolism and its regulation in dairy starter and related bacteria redefined at higher resolution Mengjin Liu1,2†, Celine Prakash2‡, Arjen Nauta1, Roland J Siezen2,3,4,5,6, Christof Francke2,4,5,6,* 1. FrieslandCampina Research, Deventer, the Netherlands 2. Center for Molecular and Biomolecular Informatics (260), NCMLS, Radboud University Nijmegen Medical Center, Nijmegen, The Netherlands 3. NIZO food research, Ede, the Netherlands 4. Kluyver Center for Genomics of Industrial Fermentation, Delft, The Netherlands 5. Netherlands Bioinformatics Center, Nijmegen, The Netherlands 6. TI Food and Nutrition, Wageningen, The Netherlands Corresponding author mailing address: RUNMC, CMBI, P.O.Box 9101, 6500HB, Nijmegen, the Netherlands. Email: [email protected] †present address: Hero-Huishan Nutrition Co.,Ltd. , CA10 Shangri-La,8 Pubei Road, Shenbei, Shenyang, China. ‡present address: Laboratory of Gene Regulation and Inflammation, Singapore Immunology Network (SIgN), Agency for Science, Technology and Research (A*STAR), Biopolis, Immunos #04-00, 8A Biomedical Grove, Singapore 138648 Keywords: Lactic acid bacteria, transcription regulation, sulphur metabolism, T-box, LysR-famliy 1 ABSTRACT Sulphuric volatile compounds provide many dairy products with a characteristic odor and taste. The volatile compounds mainly originate from the catabolism of the sulphur-containing amino acids cysteine and methionine by the lactic acid bacteria applied as starter cultures. To better understand and control the environmental dependencies of sulphuric volatile compound formation by starter bacteria, we have used the available genome sequence and experimental information to systematically evaluate the presence of the key enzymes and to reconstruct the general modes of transcription regulation for the corresponding genes in species of the order Lactobacillales. The genomic organization of the key genes is suggestive of a sub-division of the reaction network into five modules, where we observed distinct differences in the modular composition between the families Lactobacillaceae, Enterococcaceae and Leuconostocaceae and the family Streptococcaceae. These differences are mirrored by the way transcription regulation of the genes is structured in these families. In the Lactobacillaceae, Enterococcaceae and Leuconostocaceae, the main shared mode of transcription regulation is Met T-box (methionine) mediated regulation. In addition, the gene metK, encoding S-adenosylmethionine (SAM) synthetase is controlled via the SMK-box (SAM). The SMK-box is also found upstream of metK in species of the family Streptococcaceae. However, the transcription control of the other modules is mediated via three different LysR-family regulators, MetR/MtaR (methionine), CmbR (O-acetyl-(homo)serine) and HomR (O-acetyl-homoserine). Redefinition of the associated DNA binding motifs helped to identify/disentangle the related regulons, which appeared to perfectly match the proposed sub-division of the reaction network. 2 INTRODUCTION Many of the characteristic flavours in fermented dairy products such as cheese and yoghurt are the result of metabolic reactions involving sulphur-containing amino acids. The micro-organisms applied in these products degrade cysteine and methionine, resulting in the production of flavour components such as methanethiol, dimethyl sulphide (DMS), dimethyl disulphide (DMDS) and dimethyl trisulphide (DMTS). Insight in the regulatory signals and pathways that control the corresponding metabolic fluxes involved in the formation of these flavour compounds and their precursors is essential to rationally control and steer the flavour profiles of said dairy products. The micro-organisms used to produce fermented dairy products belong to the taxonomic order Lactobacillales which includes the families Enterococcaceae, Lactobacillaceae, Leuconostocaceae, and Streptococcaceae. Many of the respective species are characterized by the fact that they produce lactic acid and are therefore known as the lactic acid bacteria (LAB). The transcription of genes encoding the proteins that are involved in cysteine and methionine metabolism in lactic acid bacteria and other Lactobacillales, is controlled by both regulator-binding and RNA riboswitch mechanisms. In various Streptococcaceae the LysR-family transcription regulators MtaR and CmbR have been shown to be involved in activation as well as repression of genes such as cysD, cysK, metA, metC, metE, and metF (e.g. for Lactococcus lactis (24, 68) and Streptococcus mutans (35, 66)). The transcription regulator HomR was reported to control the expression of metB in S. mutans and Streptococcus thermophilus (67). 3 In addition, three types of riboswitches for the regulation of cysteine and methionine metabolism have been reported for low-GC Gram-positive bacteria: the T-box, the S-box and the SMK-box (14, 21, 57, 76, 77). A riboswitch is a structural sequence element at the 5’ untranslated region of an mRNA molecule that can change conformation depending on the binding of an effector molecule. The conformational change can terminate transcription (when forming a terminator structure) or allow read through (when forming an anti-terminator structure) (73, 75). In the case of the T-box, a terminator structure is formed shortly after transcription initiation unless an uncharged tRNA related to a specific amino acid binds to the specifier codon present in the T-box element, whereupon the anti-terminator structure is formed (see (28)). In the case of the S-box and the SMK-box the terminator structure is formed in the presence of S-adenosylmethionine (SAM), whereas in the absence of this molecule transcription will continue (21, 26, 46). Several studies describe the regulation of sulphur-containing amino acid metabolism for specific LAB and other closely related gram-positive bacteria. For instance, Hullo et al. (29) reported on the regulatory mechanisms related to cysteine and methionine conversions in Bacillus subtilis. Sperandio et al. described these relations for Lactococcus lactis (68) and Streptocococus mutans (66, 67). Rodionov et al. (57) and Kovaleva and Gelfand (35) performed a comprehensive comparative in silico study for the transcriptional regulators CmbR and MtaR within gram-positive bacteria. However, the availability of additional experimental and sequence data now allows an overview of the transcriptional control of the key enzymes involved in cysteine and methionine metabolism at a higher resolution. We therefore decided to extend the latter studies and to focus our efforts on the LAB and other Lactobacillales. 4 In a previous study, we improved the annotation of key enzymes involved in the metabolism of cysteine and methionine in LAB using genome-wide comparative analyses (40). Here, we extend the list of enzymes on basis of the pathway information present in the KEGG database (31). Redefinition of the binding motifs for CmbR, MetR/MtaR and HomR in Lactococci and Streptococci allowed the identification of transcription factor specific binding sites for these regulators. Also, Met-specific T-boxes and SMK-boxes were identified in recently sequenced and published genomes of e.g. L. bulgaricus, L. reuteri and L. casei. The absence of S-boxes (SAM-I) in the Lactoballilales as observed hitherto, was confirmed. Potential structure forming elements associated with the cysK gene and the hom-thrBC operon in various LAB were revealed as presented below. MATERIALS AND METHODS Genomic information, Tools and Data. Genomic information was retrieved from the ERGO resource (as of Dec 2009 (52)) and from the NCBI microbial genome database (as of September 2011 (http://www.ncbi.nlm.nih.gov/genomes/lproks.cgi)). BLAST searches were performed according to (1). Multiple sequence alignments and neighbour-joining trees (corrected for multiple substitutions) were generated using ClustalX (36). BioEdit was used to manipulate the alignments and to toggle between translated protein and nucleotide sequence (version 7.0.9, http://www.mbio.ncsu.edu/BioEdit/bioedit.html). Hidden Markov Models (HMMs) were made and genome-wide HMM searches were performed using the HMMER package (http://hmmer.janelia.org/)(13). Genome context was visualized and upstream sequence data was collected using Microbial Genome Viewer 2 (version 1 in (33), version 2 5 [http://mgv2.cmbi.ru.nl/genome/index.html]; Overmars unpublished). Potential transcription factor binding sites and other regulatory elements of fixed composition and size were searched using the Similar Motif Search approach described by (18). The original data supporting the analyses presented in this paper can be found at www.cmbi.ru.nl/bamics/supplementary/Liuetal_2012_CysMetregulation. Collection of genes related to cysteine and methionine metabolism. The species and strains that were analysed included all complete Lactobacillales genomes published before june 2011 and present within the NCBI database. The KEGG map ‘cysteine and methionine metabolism’ (map 00270) was used to define a core set of enzyme activities. The set extends the set of enzymes we previously defined (40). The protein sequences of experimentally verified members of the set (supplementary file 1) were used to search orthologs/functional equivalents in other species using BLAST. An orthologous relationship and/or functional equivalency was defined on basis of our earlier analyses (40), BLAST e-values, and in some cases multiple sequence alignments followed by clustering on basis of neighbour-joining (as described in (71)). The complete list of enzymes, their function annotation and the relevant experimental literature is given below. The annotation data was taken from the NCBI (55), KEGG (31) and PFAM (17) reference databases. The enzymes have been grouped in five clusters on basis of the composition of the related operons and shared EC numbers. - Enzymes group 1: homoserine dehydrogenase (hom, EC 1.1.1.3, COG0460E, K00003, PF03447 and 00742 (7, 43, 53)); homoserine kinase (thrB, EC 2.7.1.39, COG0083E, K00872, PF08544 and 00288 (43)); aspartate kinase III (thrA (Bsubtilis _yclM), EC 2.7.2.4, COG0527E, 6 K00928, PF01842 and 00696 (7, 34)); threonine synthase (thrC, EC 4.2.3.1, COG0498E, K01733, PF00291 (44, 61, 64, 70)). - Enzymes group 2: serine acetyltransferase (cysE, EC 2.3.1.30, COG1045E, K00640, PF06426 (22, 29, 68)); homoserine O-acetyltransferase (metA, EC 2.3.1.31, COG1897E, K00651, PF04204 (80)); cysteine synthase A and cysteine synthase-like protein (Bsubtilis_cysK and Bsubtilis _ytkP, EC 2.5.1.47, COG0031E, K01738, PF00291 (24, 29, 74)); cystathionine gamma-synthase and O-acetylhomoserine (thiol)-lyase (Bsubtilis_yjcL, EC 2.5.1.48, COG0626E, K01739, PF01053 (3, 32)); O-acetyl-L-homoserine sulfhydrolase and O-acetyl-L-serine sulfhydrolase (cysD, EC 2.5.1.49, COG2873E, K01740, PF01053); cystathionine beta-synthase for the reverse transsulfurase pathway (Bsubtilis_yrhA, EC 4.2.1.22, COG0031E, K01738, PF00291 (29)); cystathionine beta/gamma-lyase and homocysteine gamma-lyase (Bsubtilis_yrhB Ecoli_metB, EC 2.5.1.48 and 4.4.1.8, COG0626E, K01760, PF01053 (12, 16, 29, 30)); cystathionine beta/gamma-lyase (Bsubtilis_yjcJ, EC 4.4.1.8 and 4.4.1.1, COG0626E, K01760, PF01053 (3)); PLP-dependent C-S lyase (Bsubtilis_patB Llactis_ytjE Ecoli_malY, EC 4.4.1.8 and 4.4.1.1, COG1168E, K14155, PF00155 (2, 30, 45)). - Enzymes group 3: 5,10 methylenetedrahdrofolate reductase (metF, EC 1.5.1.20, ?, K00297, PF02219 (62)); bifunctional homocysteine S-methyltransferase 5,10-methylenetetrahydrofolate reductase protein (Bsubtilis_yitJ, EC 2.1.1.10 and 1.5.1.20, COG0646E (cobalamin dependent), K00547, PF02219 and 02574 (41)); homocysteine S-methyltransferase (mmuM, EC 2.1.1.10, COG2040E, K00547, PF02574 (72)); MmuM associated amino acid permease (mmuP, COG0833E, K03293, PF00324); methyltransferase (Bsubtilis_yxjG and Bsubtilis_yxjH Llactis_yhcE, EC 2.1.1.14?, COG0620E (cobalamin-independent), K00548, PF01717 (9, 37)); 7 5-methyltetrahydropteroyltriglutamate--homocysteine S-methyltransferase (metE, EC 2.1.1.14, COG0620E (cobalamin-independent), K00549, PF08267 and 01717 (20, 25)); S-ribosylhomocysteinase (luxS Llactis_ycgE, EC 4.4.1.21, COG1854T, K07173, PF02664; (37, 56)). - Enzymes group 4: C-5 cytosine-specific DNA methylase and SP-beta prophage DNA (cytosine-5-)-methyltransferase (Bsubtilis_ydiO Bsubtilis _ydiP Bsubtilis_mtbP, EC 2.1.1.37, COG0270L, K00558, PF00145 (50, 78)); 5'-methylthioadenosine nucleosidase and S-adenosylhomocysteine nucleosidase (mtn Streptococci_pfs, EC 3.2.2.16 and 3.2.2.9, COG0775F, K01243, PF1048 (10)). - Enzymes group 5: S-adenosylmethionine synthetase (metK, EC 2.5.1.6, COG0192H, K00789, PF02773 and 00438 (21, 47)). Identification of putative regulatory elements and their regulons. Cis-regulatory elements were defined according to the specific footprinting method set out by Francke et al. (19). The method relies on the definition of Groups Of Orthologous Functional Equivalents (GOOFEs) on basis of orthology and conserved genomic context. The comparative linear genome maps generated by the Microbial Genome Viewer were used to visualize and inspect the context. For every GOOFE, the upstream regions (normally ~200 nucleotides) were collected and conserved sequence elements were searched by eye from a multiple sequence alignment and by using MEME (4). The conserved elements were compared and potential regulatory regions identified. In case the conserved elements resembled transcription factor binding motifs reported in literature, experimental data on regulators of the same protein family was searched directly via PubMed (59) or in the reference databases Regulon DB (23) and DBTBS (63). Because members 8 of the same regulator-protein family will in general adopt the same fold, the DNA-binding motif should be similar (i.e. similar composition, and the same size and spacing). Therefore, established binding motifs of regulator-protein family members were taken into account to define the actual binding motif. In addition, we defined the motifs such that they obey general constraints imposed by the molecular nature of the binding process and the helical nature of the DNA molecule. Since most regulator proteins bind to the DNA as a dimer, a binding-site will in general be made up of two monomer binding sites and will have to be either palindromic or represent a direct repeat. Moreover, since the DNA is helical the actual monomer binding-site in general has to be shorter than 7 nucleotides and the two sites that make up the dimer binding-site have to be interspaced by a fixed number of nucleotides. The defined motifs (given in supplementary file 2) were converted to a position frequency matrix, which was used directly to score potential transcription factor binding sites and other regulatory elements of fixed composition and size. In this way the score of any DNA sequence will relate directly to its similarity to the input motif. We have validated and used this approach with success to identify potential binding sites of CcpA and Spo0A in low GC gram positive organisms and the sigma-54 promoter in all organisms (18). A cut-off score of >83% relative similarity and a positioning of maximally around 200 nucleotides upstream (with some exceptions) of the translation start was used to select potential binding sites for the various regulators. The uniform cut-off score was chosen such that the number of false positive assignments should be limited, i.e. such that experimentally validated sites were included and that the number of correctly positioned sites was high (position in terms of distance and orientation with respect to translation start of gene downstream). The identified regulatory 9 elements were related to all genes present in the downstream operon, where an operon was defined as those genes on the same strand that are separated by an intergenic region of less than 250 nucleotides and which does not contain a termination signal. The analyzed results of the motif searches are given in supplementary file 3. Identification of riboswitches and other structural elements. Hidden Markov Models were constructed for the T-box motif and for the S-box (SAM-I) motif on basis of the available literature (57, 76, 77). Both HMMs were used to scan the selected genomes (cut-off e-value 1 (37)) and the locations of putative boxes were identified. The amino acid specificity of the detected T-boxes was established on basis of the specifier codon as described by Wels et al. (77) and exemplified in Figure S1. Two characteristic SMK-box sequences were defined on basis of (21), as given in Figure 2A, and these were used to scan the selected genomes using the Similar Motif Search procedure (results in supplementary file 3). Only in case both motifs were found directly upstream of a gene and they were complementary we considered the site a putative SMK-box. RESULTS Comparative analysis of the enzymes involved in central cysteine and methionine metabolism. The set of genes related to central cysteine and methionine metabolism was identified on basis of KEGG map00270 (31) and earlier work by others (35, 57) and us (40). Orthologs and homologs of these genes were collected from the genomes of all sequenced species/strains of the LAB and other Lactobacillales as described in Materials & Methods. The related reaction network is given in Figure 1 and the results of the search and analysis procedure 10 are presented in Tables 1 and 2 and given in supplementary file 1. Remarkably, the set of genes (and corresponding proteins) can be divided into five separate groups on basis of genomic organisation, the identity of the EC numbers and the position in the reaction network (see Figure 1). Small differences in operon organization between the families Lactobacillaceae, Enterococcaceae and Leuconostocaceae (Table 1) and the family Streptococcaceae (Table 2) were observed, implying a difference in pathway modularization between the families. The presence-absence list of genes is in agreement with the results obtained in earlier analyses (35, 40, 57). Nevertheless, we observed a few differences. For instance, we found that O-acetylhomoserine sulfhydrolase (gene cysD) is absent in various S. pneumoniae strains including str. TIGR4 and that B12-dependent methionine synthase (gene yxjH) is present in S. gordonii. We also found that the enzyme cysthathionine beta/gamma-lyase of S. pyogenes is more similar to that of B. subtilis (gene yjcJ) than to that of L. lactis (gene ytjE). A multiple sequence alignment of MetA showed that the protein in all Lactobacillales carries the specific Glu residue at position 111 that renders the MetA protein of Bacillus cereus an acetyl-transferase instead of a succinyl transferase (as shown by (80)) (see supplementary file 4). Because we could include more species and strains the general trends in the presence or absence of certain enzymes became more apparent visible and peculiar compositions better to trace. As an example of the latter, orthologs of cystathionine synthase and cystathionine beta/gamma-lyase as represented by genes yrhA and yrhB in B. subtilis were not found among the Streptococci except for the sequenced S. thermophilus strains. We observed that the upstream region associated with the gene yrhA in S. thermophilus was identical (besides a few SNPs) to that of yrhA in Lactobacillus helveticus, L. delbrueckii bulgaricus and that of yrhA on plasmid 11 pLC1 of Lactobacillus rhamnosus Lc 705w (see Figure S2), suggesting a recent plasmid-mediated transfer of the genes and upstream sequences to all the different strains individually or an extreme stability of this particular sequence. We found the enzymes 5'-methylthioadenosine nucleosidase/S-adenosylhomocysteine nucleosidase (corresponding gene designated mtn in B. subtilis and E. coli or pfs in Streptococcaceae) and S-adenosylmethionine synthetase (gene metK) to be present in all analyzed genomes and thus potentially essential. The enzyme S-ribosylhomocysteinase (gene luxS) was found absent only in L. sakei. Besides, serine acetyltransferase (gene cysE) and cysteine synthase A (gene cysK) are present in all Streptococcaceae and N5-methyltetrahydrofolate methyltransferase (gene yxjH or metE2) in all Lactobacillaceae except for L. sakei and L. helveticus. The remaining make-up of cysteine and methionine metabolism appeared more variable between species. L. sakei (Lactobacillaceae) and S. equi (Streptococaceae) have the least extensive enzyme repertoire with 2 and 7 enzymes, respectively. Identification of Riboswitches. Recently, three comprehensive studies described the occurrence and evolution of T-boxes among prokaryotes (28, 76, 77). We have used the T-box HMMs of (77) to search recently acquired genome sequences of e.g. L. bulgaricus, L. brevis, L. reuteri etc. We identified many new T-boxes associated with genes/operons involved in cysteine and methionine metabolism in these genomes. As a control we also scanned the other genomes included in previous studies (76, 77). The conservation of the ATG specifier codon in the multiple sequence alignment of the newly recovered T-boxes (Figure S1) implies that they all respond to the absence of methionine. We found that genes/operons metB (BS_yjcL), metE-metH (BS_metE-yitJ) and hom1-metA-cysD from L. plantarum WCFS1 and genes/operons 12 LEUM_1806-LEUM_1803 (BS_metA-yjcL-yjcJ-yxjG), LEUM_1802 (luxS) and LEUM_1795-LEUM_1794 (metE-metF) from Leuconostoc mesenteroides are regulated by a Met T-box, in agreement with (76, 77). Also cysE, encoding a serine acetyl-transferase was found to be preceded by a Met T-box in B. subtilis, as reported by (54). The association with a Met T-box appeared almost fully conserved within the Lactobacillus species for the genes that encode the proteins responsible for the synthesis of methionine from homocysteine, i.e. metEF, yxjH/yxjG and luxS, and for the genes encoding aspecific methionine ABC import system, where metQ encodes the substrate binding protein. S-adenosyl methionine sensitive SAM-I riboswitches are often found upstream of genes involved in sulphur metabolism and transport in Bacilli and Clostridia (3, 57), but they have not been reported in LAB. In accordance, we did not detect SAM-I riboswitches upstream of genes involved in cysteine and methionine metabolism in the genomes that were analyzed. However, another SAM responsive element was reported by (21) upstream of the metK gene in Lactobacilli and Streptococci, which was named the SMK-box. We used two conserved structure forming stretches of around 6 nucleotides from the reported box (as given in Figure 2A) to search for potential SMK-boxes. We found the two motifs in the correct order upstream of the metK gene in all analyzed genomes, but no additional hits (supplementary file 3). For all Lactobacillaceae and many Streptococcaceae the distance was about 50 nucleotides, whereas in other Streptococci like S. dysgalactiae, S gallolyticus, S. mitis, S. mutans, S. pasteurianis and S. pyogenes this distance was much larger to about 350 nucleotides. This huge variability in spacing, that was also observed by (21) raises interesting questions related to the way in which sequences of such a different length can form similar three-dimensional structures to accommodate SAM binding. 13 LysR-family Regulators related to cysteine and methionine metabolism. MetR from S. mutans, MtaR from Streptococcus agalactiae, CmbR from L. lactis and HomR from S. mutans have been identified as being important regulators of the genes related to cysteine and methionine metabolism in Streptococci (24, 66, 67). They belong to the LysR-family of transcription regulators. A neighbour-joining tree of LysR-family proteins was constructed on basis of a comprehensive BLAST search for homologs (cut-off e-value: 1*e-5). There was a clear division of protein sequences in three subclusters, corresponding to MetR/MtaR, CmbR and HomR, within the resulting NJ-tree as observed also by (35). The tree (Figure 3) clearly shows that MetR of S. mutans and MtaR of S. agalactiae are orthologous and that CmbR and HomR are their closest relatives. The phylogenetic analysis further implies that CysR from S. mutans is orthologous to L. lactis although CmbR was proposed to be a new regulator separated from the CmbR cluster (67). Our assessment of orthology between the two transcription factors is supported by the similarity of their cognate DNA binding-motif (see below). For the other species, only Lactobacillus delbrueckii, L. plantarum and Enterococcus faecalis possessed a homolog. The former two are clearly orthologous to MetR/MtaR, whereas the Enterococcal regulator seems most related to CmbR. The LysR-family proteins have a domain architecture that is common for transcriptional regulators in prokaryotes. The member proteins consist of a signal-molecule binding domain followed by a helix-turn-helix (HTH) DNA-binding domain. The family contains a number of well-studied proteins, including AlsR, CcpC, CitR, GltR, YwfK and CmbR (42). We have compared the results of DNA-binding studies for LysR-family members for a number of Firmicutes (data in Table S1). A straightforward alignment and comparison of the reported 14 binding sites reveals a common motif structure of the cis-elements, namely ATNNNN---NNNNAT. The motif displays a clear dyad symmetry and a conserved spacing of three nucleotides. This architecture agrees well with the observation made by Schell et al. (60) that LysR-family members generally recognize a box with a conserved sequence T-N11-A, located 50-80 bp upstream of the transcriptional start site. In fact, LysR-family members in general assemble as tetramers (dimer of dimers) and thus their binding locus most often is composed of two adjacent dimer binding-sites where the spacing between the two sites may vary (42). The motifs for LysR binding that have been defined more recently in some cases deviate from this family-consensus and we have therefore redefined them as described below, taking the mechanistic/molecular characteristics of the binding into account. The motifs were then used to search the Lactobacilli genomes for putative binding-sites and the results are given in Tables 1, 2 and 3. MetR/MtaR. A 17-bp palindromic conserved sequence TATAGTTtnaAACTATA was identified upstream of metY, metA, metQ, metI and the metEF operon in Streptococci and upstream of the metEF operon in L. lactis (35, 57). However at that time, the regulatory protein which should bind to the so-called MET-box had not yet been identified. Rodionov et al. (57) initially proposed that the transcriptional regulator MtaR, known to be involved in methionine uptake in S. agalactiae, was a good candidate. Indeed, later it was later shown that MetR is the regulator protein that binds to the MET-box in S. mutans (66). In fact, MetR and MtaR can be inferred to fulfil the same role as they are orthologs (Figure 3). The MET-box motif was identified in the upstream regions of atmB, metE, cysD, metA, and smu.1487 in S. mutans and 15 binding of MetR to these MET-boxes was confirmed using gel mobility shift assays and base substitutions in the MET-boxes (66). We have analyzed the upstream regions of the genes orthologous to metE and metH (mmuM) in the selected genomes for the presence of a sequence similar to the MET-box. Like others we observed two MetR/MtaR-binding-sites in the upstream region of the metH (mmuM) and metE genes in the L. lactis and S. thermophilus genomes. The first site is often located around 60-70bp from the transcriptional start and displays an activating role (66). The second putative binding site, which is closer to the transcriptional start, is far less conserved. The observed organization of the binding-sites appears general for LysR-family members and relates to a mechanism that requires tetramer formation of the transcription factor (42). It has clear implications for the dynamics of LysR-family mediated transcriptional regulation as described for the enterobacterial nitrogen assimilation control protein Nac by (58). The dissected sites were used to redefine the binding motif for MetR/MtaR in both lactococcal and streptococcal strains (Figure 2B; supplementary file 2). We used the experimentally identified binding sites of all LysR-family regulators to guide the definition (see above and Materials & Methods). The putative binding sites of MetR/MtaR showed an overall palindromic structure, ATA-N9-TAT, which is typical for the LysR-family. The most conserved element ATAGTT is located upstream, whereas the downstream element N3-XXCTAT shows somewhat less conservation. The complete MetR/MtaR dimer binding motif we thus define is ‘ATAGTT-N3-XXCTAT’. The newly defined MetR/MtaR binding-motif is covered completely by the earlier defined MET-box, yet it is two nucleotides shorter so that it agrees with the LysR-family characteristic. The recovery of putative binding-sites in genome-wide searches is 16 very sensitive with respect to the precise composition of the search motif, and therefore we used both the newly defined motif and the extended motif (one nucleotide at each side) reported by (57) to identify MetR/MtaR binding sites. We observed only a few differences between the two searches (not shown) although the latter search seemed to be more discriminative and the results given in Tables 2 and 3 and supplementary file 3 therefore relate to the second search. The observed positive effect of the addition of the flanking nucleotides in the search and their conservation could well be related to potential effects of the flanking nucleotides on the molecular structure of the binding site (where the physical binding occurs) and therefore on binding-affinity. A clear MetR/MtaR binding motif was discerned in the upstream region of metA, yjcLpatB, metEF/metEyitJ, mmuM (metH), yxjH and the operon related to ABC mediated transport of methionine in most Streptococcal genomes (see Table 2 and 3 and supplementary file 3) as reported earlier (57). These regulatory relations seem to be conserved (with a few exceptions as described below), i.e. when the genes are present the MetR/MtaR binding-site is also present. However, some sites score relatively low which could be indicative of lost function, but could also be very well related to slightly altered binding preferences between species. In the case of S. gordonii, S. parasanguinis and S. sanguinis the SMK-box upstream of metK is preceded by a MetR/MtaR binding-site. As S. equi, S. dysgalactiae, S. parauberis and S. uberis lack the related genes, MetR/MtaR-mediated regulation seems restricted to methionine import via the ABC transport system. Another regulatory connection that was conserved in at least three species was observed in S. mitis, S oralis and S. pneumoniae for the genes fhs and folD, encoding formate-tetrahydrofolate ligase and a bifunctional methylenetetrahydrofolate dehydrogenase 17 (NADP+) / methenyltetrahydrofolate cyclohydrolase, respectively, which are important in the biosynthesis of the co-factor tetrahydrofolate. In L. lactis MetR/MtaR-mediated regulation seems to be restricted to metEF although most of the other genes are present in the L. lactis genomes. In the Lactobacilli L. delbrueckii subsp. bulgaricus and L. plantarum only one gene seems to be connected to MetR/MtaR. In L. delbrueckii subsp. bulgaricus the gene metE2 is preceded by an obvious binding-site. Remarkably, in other Lactobacilli the gene is preceded by a T-box. As the gene is adjacent to the gene encoding the regulator it could well be that this unit has been acquired from some Streptococcus. In fact, also on the L. plantarum genome the genes metE and metR are neighbours. Suprisingly, in L. plantarum the MetR/MtaR binding-site appears to have been lost. However, a binding-site was found upstream of trxB1 (lp_0761), a gene encoding a thioredoxin reductase, more remotely connected to sulphur metabolism. Similarly, a binding site was found upstream of coaE in L. lactis, which encodes dephospho-coenzyme A kinase, the enzyme that catalyzes the final step in the biosynthesis of the thiol-compound coenzyme A. CmbR/CysR. CmbR positively regulates the metC-cysK (i.e. yrhB-cysK1) operon in L. lactis (24Mic). In fact, the expression of eighteen genes is affected by a cmbR knockout in this species (68). The genes, which are grouped in several transcriptional units, include cysD, cysM (i.e. cysK2), yhcE (i.e. yxjG/yxjH), metC-cysK (i.e. yrhB-cysK1) and metA-metB1-ytjE (i.e. metA-yjcL-patB) and the methionine (plpABCDydcBD) and cystine (yjgCDE) ABC transport systems (68). It was shown that binding of CmbR to the metB2-cysK (i.e. yrhB-cysK1) promoter is stimulated by O-acetyl-L-serine and by low cysteine and methionine concentrations in L. lactis (15, 24). In vitro promoter binding studies confirmed strong binding of CmbR to the metB2-cysK 18 (i.e. yrhB-cysK1) promoter and weaker binding to the promoter of cysD and cysM and of the methionine (plpA) and cystine (yjgC) ABC transport system operons. Hardly any binding was observed for metA-metB1-ytjE (i.e. metA-yjcL-patB) and yhcE (i.e. yxjG/yxjH) (Figure 2 in (68)). In S. mutans cysK, tcyD (i.e tcyJ/K), metBC (i.e. yjcL-patB) and homR expression was shown to be affected upon deletion of cmbR (67). As CmbR belongs to the LysR-family of transcription regulators, one would expect its binding site to have a common LysR-family motif. However, all studies dedicated to CmbR binding to date have resulted in a distinctly different CmbR-binding motif definition. In the first study, a deletion analysis showed that a direct repeat of ATAAAAAAA is required for metC activation by CmbR (24). In the second study, the upstream regions of seven transcriptional units found to be regulated by CmbR in L. lactis were analyzed. A first consensus binding sequence TWAAAAATTNNTA was proposed, centered 46 to 53 bp upstream of the transcriptional start, with a second consensus TWAAAWANNTNNA, located 8 to 10 bp upstream (68). The approximate location of CmbR binding in the metB2 and cysD upstream regions was determined by gel-shift experiments (24, 68). A recent publication by Kovaleva et al. (35) describes an in silico analysis of upstream regions of cysteine biosynthesis genes in streptococcal species and, to increase the recognition power, defines the CmbR-binding motif as TGATA-N9-TATCA-N2-4-TGATA. To resolve the discrepancies resulting from the previous analyses, we re-analyzed the reported CmbR-binding sites in L. lactis and Streptococcus species. We identified the consensus binding sequences in the upstream region of the CmbR-regulated cysK gene and its paralogs/orthologs from lactococcal and streptococcal genomes (Figure 2B; supplementary file 19 2). Since two sub-clusters with respect to the cysK gene can be distinguished by phylogeny and gene context in the L. lactis strains, at first a putative CmbR-binding site was identified for each sub-cluster. In case of the so-called cysM1 cluster, CmbR-binding sites from other streptococci such as S. agalactiae and S. gordonii were also taken into account. Nevertheless, all the putative CmbR-binding sites in both Lactococcus and Streptococcus could be summarized by a motif of dyad symmetry ATA-N9-TAT, with a preference for a long stretch of adenines following the starting “ATA”. This putative CmbR binding motif has the general LysR-family signature and, in addition, all the previous experimental work on CmbR binding supports this assignment. Moreover, like in the case of MetR/MtaR a second more degenerate site is found directly downstream. Similarly, we observed a positive effect of the addition of the flanking nucleotides in the search for putative CmbR binding sites and we have thus used the extended motif in our searches (Figure 2B). In the NJ-tree, the CmbR subfamily is most closely related to the MetR/MtaR subfamily (Figure 2). Therefore, one might expect that their binding sites resemble each other. Indeed the motifs are remarkably similar. However, there were some small differences especially in the conserved flanking residues. Like in the case of MetR/MtaR we have thus used the extended CmbR motif to search for additional binding sites (in this case 3 nucleotides on each side). The results are depicted in Tables 1, 2 and 3 and supplemental file X3. We retrieved the known binding sites in the region upstream of metB2-cysK in the Lactococcus and cysK in the Streptococcus strains. We also identified a novel site upstream of cysD in the Streptococcus thermophilus strains, whereas in for example S. mutans the regulation of cysD seems mediated by MetR/MtaR. Another conserved relation of CmbR was found with homologs of the gene 20 encoding the ABC transport related cystine substrate binding protein (6, 51). In many Streptococci, two CmbR regulated copies of the gene are present, the first related to tcyA or yxeM of B. subtilis and the second to tcyJ and tcyK of B. subtilis. The first is associated with the metR/mtaR regulated methionine ABC transport related operon (48, 79) and the second with genes encoding the permease and ATP-binding sub-unit of a separate cystine ABC transport system (related to tcyLMN in B. subtilis (6)). Remarkably the scoring suggests that in S. suis also the regulation of metBC, metEF and the methionine ABC transport related operon is CmbR-dependent instead of MetR/MtaR-dependent like in the other Streptococci. For S. mutans our observations fit the regulon derived on basis of the cmbR knockout experiments (67). In contrast, the number of putative CmbR binding sites detected in L. lactis was low compared to the number of reported putative sites (68). Only the highest affinity promoter was retrieved. Apparently, the defined motif is not very discriminative in the analysis of the L. lactis genomes. Other regulatory connections that were conserved in at least three species were observed with genes encoding pyruvate formate lyase, dihydrofolate reductase, a glutamine amidotransferase, a beta-lactamase, glyceraldehyde 3-P dehydrogenase and a specific RNA methyl transferase in various species (Table S2). HomR. HomR is the third transcriptional regulator of the LysR-family that is closely related to MetR/MtaR and CmbR and which was found only in S. gallolyticus, S. mutans, S. pasteurianus and S. thermophilus. The expression of S. mutans metBC encoding cysteine biosynthesis genes and the tcyDEFGH (i.e. tcyJKLMN) cluster encoding a cysteine transport system is specifically affected in a homR knockout mutant (67). We examined the upstream 21 region of metB in the HomR containing Streptococci and could identify a clear HomR binding-motif according to the common structure of LysR-family DNA elements (as given in Figure 2B). The binding-motif resembles that of MetR/MtaR, the most obvious differences being located in the flanking nucleotides. Similar to MetR/MtaR and CmbR, HomR exerts control over metB expression via a second binding site located downstream with lower similarity. We have used the extended HomR-related motif to search additional binding-sites (supplementary file 3). Besides the conserved relation with regulation of the metBC (i.e. yjcL-patB) operon, we found a HomR specific site upstream of folD in S. pasteurianus and panE (encoding a 2-dehydropantoate 2-reductase) in S. thermophilus. In addition, the relative scoring is suggestive of some overlap with the regulation via MetR/MtaR for the metEyitJ operon in S. gallolyticus, S. mutans, and S. pasteurianus and for metA in S. thermophilus. CodY. A nutritional regulator involved in the global control of amino acid metabolism in Gram-positive bacteria is CodY (65, 69). It has been extensively studied in L. lactis (11, 27) and other Streptococci (8, 38, 39) and was reported to control the expression of hom, thrA, thrB, thrC and cysD in these organisms. We defined a CodY binding motif on basis of the data provided in (11). The motif is given in Figure 2C and is almost identical to the experimentally defined motif of B. subtilis (5). The motif was used to search for putative CodY binding-sites upstream of the genes related to cysteine and methionine metabolism. The results of the search are listed in Tables 1 and 2 and supplementary file 3. We found similar motifs upstream of thrA, thrC and/or the hom-thrB operon in many of the Streptococcal genomes. However, the upstream regions are relatively dissimilar between the species and as a result for various other Streptococci it is not particularly clear whether the CodY binding-site is conserved (not shown). We found a potential 22 site upstream of the metA-metB1-ytjE (i.e. metA-yjcL-patB) operon in L. lactis and the metB1-ytjE (i.e. yjcL-patB) operon in S. thermophilus. Novel elements upstream of cbs and thrB genes. A comparative analysis (see Materials & Methods) of the upstream regions of the other genes involved in cysteine and methionine metabolism did yield three additional putative regulatory elements. The yrhA-yrhB operon in various Lactobacilli strains such as Lb. salivarius, Lb. plantarum, Lb. bulgaricus, Lb. acidophilus, as well as in O. oeni, and Leuconostoc is preceded by a conserved palindromic motif AAAGGGCGCGAA-N(11-18)-TTCGCGCCtTTT (Figure 4A). As described earlier all S. thermophilus strains carry an orthologous operon with a completely conserved upstream region. We could not detect identical sites elsewhere on the genome. Considering the variable spacing between the complementary stretches the motif probably represents a structural element. Similarly, a multiple sequence alignment of the upstream regions of thrB from Lb. acidophilus and Lb. gasseri, revealed another previously unidentified conserved motif (Figure 4B). The putative motif consists of inverted repeats separated by 15 bp (ATTGTAAC-N15-GTTACAAT). Interestingly, each part of the inverted repeat complements itself (e.g. in the first part, ATTG is followed immediately by its complement sequence TAAC). Again no similar sites were found elsewhere on the genome. Another potential structural motif was recovered upstream of thrB in L. lactis (Figure 4C). The 12 nucleotide sequence is found more than 100 times seemingly randomly distributed in all L. lactis genomes and less than 2 times in all other analyzed genomes (not shown). The motif was discovered before and called Highly Repetitive Motif by (49). The absolute conservation and occurrence implies functional importance to L. lactis although it has yet to be discovered what the precise regulatory role might be. 23 DISCUSSION We have performed a genome-wide in silico analyses to reveal the transcription regulatory interactions that control the expression of the genes encoding various key enzymes involved in cysteine and methionine metabolism in all sequenced species of the order Lactobacillales. The associated regulators we could identify were the known regulatory proteins such as CmbR, MetR/MtaR, HomR and CodY as well as known RNA riboswitches. In addition, we found two potential regulatory structure forming elements. We redefined specific binding motifs for CmbR, MetR/MtaR and HomR in Lactococci and Streptococci. All motifs follow the characteristic LysR-family signature (ATA-N9-TAT) and are substantiated by the available experimental data. The motifs allowed a computational separation of the various binding-sites within the Streptococci. The identified regulator-gene associations overlapped well with the subdivision of the reaction network that was made on basis of the operon composition and on basis of EC numbers (compare Figure 1 with Tables 1 and 2). A clear diversity in the transcriptional regulation network between the different families was observed. In various Lactobacillaceae like Lb. plantarum the biosynthesis of methionine, as well as its precursors i.e. homocysteine and cystathionine, is responsive to low methionine levels through T-box mediated regulation. Similarly, the ABC transport of methionine is regulated via a T-box riboswitch. We could not identify new general sites related to the synthesis of cysteine from homocysteine and the degradation of both cysteine and methionine in the Lactobacillaceae. This may indicate that most of the relevant regulators are known or that potential other elements might be less conserved or species specific. In Streptococci, the gene-regulator associations 24 nicely fit the established inducer substrates. MetR/MtaR controls the expression of the genes that encode biosynthesis and transport of methionine, whereas CmbR relates to the control of cysteine biosynthesis and cystine transport. HomR appears specifically dedicated to the control of yjcL-patB expression in four Streptococci. The latter relationship makes sense as the genes encode the enzymes responsible for the conversion of acetyl-homoserine to cystathionine and further. In S. thermophilus, an additional gene controlled by HomR encodes the protein that catalyzes the production of acetyl homoserine. The intracellular levels of homoserine seem to be controlled by the global regulator CodY. The biosynthesis of flavour compounds, e.g. H2S, methanethiol, DMS, DMDS, DMTS, is catalyzed directly by cystathionine beta/gamma lyase, which is encoded in most Streptococci by a single gene, the expression of which is controlled by MetR/MtaR (methionine). Of course, changing the levels of the flavour precursors, methionine, cysthathionine and cysteine will also affect the synthesis rate of flavour compounds. These levels are determined by the activity of cysteine synthase (cysK), cystathionine synthase (yrhA), homocysteine S-methyltransferase (yitJ and mmuM) and S-adenosylmethionine synthetase (metK) and thus controlled by CmbR (O-acetyl-(homo)serine), MetR/MtaR (methionine) and an SMK-box (S-adenosylmethionine), respectively. Inclusion of the above reaction network and the regulatory interconnections into a quantitative metabolic network model may help to rationalize new strategies for controlling the sulphuric flavour formation in various LAB-fermented food products. ACKNOWLEDGEMENTS 25 This work was supported by grant CSI4017 from the Casimir program of the Ministry of Economic Affairs, the Netherlands. 26 REFERENCES 1. 2. 3. 4. 5. 6. 7. 8. 9. 10. 11. 12. 13. 14. 15. Altschul, S. F., T. L. Madden, A. A. Schaffer, J. Zhang, Z. Zhang, W. Miller, and D. J. Lipman. 1997. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25:3389-3402. Auger, S., M. P. Gomez, A. Danchin, and I. Martin-Verstraete. 2005. The PatB protein of Bacillus subtilis is a C-S-lyase. Biochimie 87:231-238. Auger, S., W. H. Yuen, A. Danchin, and I. Martin-Verstraete. 2002. The metIC operon involved in methionine biosynthesis in Bacillus subtilis is controlled by transcription antitermination. Microbiology 148:507-518. Bailey, T. L., M. Boden, F. A. Buske, M. Frith, C. E. Grant, L. Clementi, J. Ren, W. W. Li, and W. S. Noble. 2009. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37:W202-208. Belitsky, B. R., and A. L. Sonenshein. 2008. Genetic and biochemical analysis of CodY-binding sites in Bacillus subtilis. J. Bacteriol. 190:1224-1236. Burguiere, P., S. Auger, M. F. Hullo, A. Danchin, and I. Martin-Verstraete. 2004. Three different systems participate in L-cystine uptake in Bacillus subtilis. J. Bacteriol. 186:4875-4884. Cahyanto, M. N., H. Kawasaki, M. Nagashio, K. Fujiyama, and T. Seki. 2006. Regulation of aspartokinase, aspartate semialdehyde dehydrogenase, dihydrodipicolinate synthase and dihydrodipicolinate reductase in Lactobacillus plantarum. Microbiology 152:105-112. Caymaris, S., H. J. Bootsma, B. Martin, P. W. Hermans, M. Prudhomme, and J. P. Claverys. 2010. The global nutritional regulator CodY is an essential protein in the human pathogen Streptococcus pneumoniae. Mol. Microbiol. 78:344-360. Chi, B. K., K. Gronau, U. Mader, B. Hessling, D. Becher, and H. Antelmann. 2011. S-Bacillithiolation Protects Against Hypochlorite Stress in Bacillus subtilis as Revealed by Transcriptomics and Redox Proteomics. Mol. Cell. Proteomics 10:M111 009506. Choi-Rhee, E., and J. E. Cronan. 2005. A nucleosidase required for in vivo function of the S-adenosyl-L-methionine radical enzyme, biotin synthase. Chem. Biol. 12:589-593. den Hengst, C. D., S. A. van Hijum, J. M. Geurts, A. Nauta, J. Kok, and O. P. Kuipers. 2005. The Lactococcus lactis CodY regulon: identification of a conserved cis-regulatory element. J. Biol. Chem. 280:34332-34342. Dobric, N., G. K. Limsowtin, A. J. Hillier, N. P. Dudman, and B. E. Davidson. 2000. Identification and characterization of a cystathionine beta/gamma-lyase from Lactococcus lactis ssp. cremoris MG1363. FEMS Microbiol. Lett. 182:249-254. Durbin, R., S. Eddy, A. Krogh, and G. and Mitchison. 1998. Biological Sequence Analysis: Probabilistic Models of Proteins and Nucleic Acids. Cambridge University Press. Epshtein, V., A. S. Mironov, and E. Nudler. 2003. The riboswitch-mediated control of sulfur metabolism in bacteria. Proc. Natl. Acad. Sci. U S A 100:5052-5056. Fernandez, M., M. Kleerebezem, O. P. Kuipers, R. J. Siezen, and R. van Kranenburg. 2002. Regulation of the metC-cysK operon, involved in sulfur metabolism in Lactococcus lactis. J. Bacteriol. 184:82-90. 27 16. 17. 18. 19. 20. 21. 22. 23. 24. 25. 26. Fernandez, M., W. van Doesburg, G. A. Rutten, J. D. Marugg, A. C. Alting, R. van Kranenburg, and O. P. Kuipers. 2000. Molecular and functional analyses of the metC gene of Lactococcus lactis, encoding cystathionine beta-lyase. Appl. Environ. Microbiol. 66:42-48. Finn, R. D., J. Mistry, J. Tate, P. Coggill, A. Heger, J. E. Pollington, O. L. Gavin, P. Gunasekaran, G. Ceric, K. Forslund, L. Holm, E. L. Sonnhammer, S. R. Eddy, and A. Bateman. 2010. The Pfam protein families database. Nucleic Acids Res. 38:D211-222. Francke, C., T. Groot Kormelink, Y. Hagemeijer, L. Overmars, V. Sluijter, R. Moezelaar, and R. J. Siezen. 2011. Comparative analyses imply that the enigmatic Sigma factor 54 is a central controller of the bacterial exterior. BMC Genomics 12:385. Francke, C., R. Kerkhoven, M. Wels, and R. J. Siezen. 2008. A generic approach to identify Transcription Factor-specific operator motifs; Inferences for LacI-family mediated regulation in Lactobacillus plantarum WCFS1. BMC Genomics 9:145. Fu, T. M., J. Almqvist, Y. H. Liang, L. Li, Y. Huang, and X. D. Su. 2011. Crystal structures of cobalamin-independent methionine synthase (MetE) from Streptococcus mutans: a dynamic zinc-inversion model. J. Mol. Biol. 412:688-697. Fuchs, R. T., F. J. Grundy, and T. M. Henkin. 2006. The S(MK) box is a new SAM-binding RNA for translational regulation of SAM synthetase. Nat. Struct. Mol. Biol. 13:226-233. Gagnon, Y., R. Breton, H. Putzer, M. Pelchat, M. Grunberg-Manago, and J. Lapointe. 1994. Clustering and co-transcription of the Bacillus subtilis genes encoding the aminoacyl-tRNA synthetases specific for glutamate and for cysteine and the first enzyme for cysteine biosynthesis. J. Biol. Chem. 269:7473-7482. Gama-Castro, S., H. Salgado, M. Peralta-Gil, A. Santos-Zavaleta, L. Muniz-Rascado, H. Solano-Lira, V. Jimenez-Jacinto, V. Weiss, J. S. Garcia-Sotelo, A. Lopez-Fuentes, L. Porron-Sotelo, S. Alquicira-Hernandez, A. Medina-Rivera, I. Martinez-Flores, K. Alquicira-Hernandez, R. Martinez-Adame, C. Bonavides-Martinez, J. Miranda-Rios, A. M. Huerta, A. Mendoza-Vargas, L. Collado-Torres, B. Taboada, L. Vega-Alvarado, M. Olvera, L. Olvera, R. Grande, E. Morett, and J. Collado-Vides. 2011. RegulonDB version 7.0: transcriptional regulation of Escherichia coli K-12 integrated within genetic sensory response units (Gensor Units). Nucleic Acids Res. 39:D98-105. Golic, N., M. Schliekelmann, M. Fernandez, M. Kleerebezem, and R. van Kranenburg. 2005. Molecular characterization of the CmbR activator-binding site in the metC-cysK promoter region in Lactococcus lactis. Microbiology 151:439-446. Gonzalez, J. C., R. V. Banerjee, S. Huang, J. S. Sumner, and R. G. Matthews. 1992. Comparison of cobalamin-independent and cobalamin-dependent methionine synthases from Escherichia coli: two solutions to the same chemical problem. Biochemistry 31:6045-6056. Grundy, F. J., and T. M. Henkin. 1998. The S box regulon: a new global transcription termination control system for methionine and cysteine biosynthesis genes in gram-positive bacteria. Mol. Microbiol. 30:737-749. 28 27. 28. 29. 30. 31. 32. 33. 34. 35. 36. 37. 38. 39. 40. Guedon, E., B. Sperandio, N. Pons, S. D. Ehrlich, and P. Renault. 2005. Overall control of nitrogen metabolism in Lactococcus lactis by CodY, and possible models for CodY regulation in Firmicutes. Microbiology 151:3895-3909. Gutierrez-Preciado, A., T. M. Henkin, F. J. Grundy, C. Yanofsky, and E. Merino. 2009. Biochemical features and functional implications of the RNA-based T-box regulatory mechanism. Microbiol. Mol. Biol. Rev. 73:36-61. Hullo, M. F., S. Auger, O. Soutourina, O. Barzu, M. Yvon, A. Danchin, and I. Martin-Verstraete. 2007. Conversion of methionine to cysteine in Bacillus subtilis and its regulation. J. Bacteriol. 189:187-197. Irmler, S., S. Raboud, B. Beisert, D. Rauhut, and H. Berthoud. 2008. Cloning and characterization of two Lactobacillus casei genes encoding a cystathionine lyase. Appl. Environ. Microbiol. 74:99-106. Kanehisa, M., S. Goto, Y. Sato, M. Furumichi, and M. Tanabe. 2011. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. Kanzaki, H., M. Kobayashi, T. Nagasawa, and H. Yamada. 1986. Distribution of two kinds of cystathionine gamma-synthase in various bacteria. FEMS Microbiol. Lett. 33:65-68. Kerkhoven, R., F. H. van Enckevort, J. Boekhorst, D. Molenaar, and R. J. Siezen. 2004. Visualization for genomics: the Microbial Genome Viewer. Bioinformatics 20:1812-1814. Kobashi, N., M. Nishiyama, and H. Yamane. 2001. Characterization of aspartate kinase III of Bacillus subtilis. Biosci. Biotechnol. Biochem. 65:1391-1394. Kovaleva, G. Y., and M. S. Gelfand. 2007. Transcriptional regulation of the methionine and cysteine transport and metabolism in Streptococci. FEMS Microbiol. Lett. 276:207-215. Larkin, M. A., G. Blackshields, N. P. Brown, R. Chenna, P. A. McGettigan, H. McWilliam, F. Valentin, I. M. Wallace, A. Wilm, R. Lopez, J. D. Thompson, T. J. Gibson, and D. G. Higgins. 2007. Clustal W and Clustal X version 2.0. Bioinformatics 23:2947-2948. Lebeer, S., S. C. De Keersmaecker, T. L. Verhoeven, A. A. Fadda, K. Marchal, and J. Vanderleyden. 2007. Functional analysis of luxS in the probiotic strain Lactobacillus rhamnosus GG reveals a central metabolic role important for growth and biofilm formation. J. Bacteriol. 189:860-871. Lemos, J. A., M. M. Nascimento, V. K. Lin, J. Abranches, and R. A. Burne. 2008. Global regulation by (p)ppGpp and CodY in Streptococcus mutans. J. Bacteriol. 190:5291-5299. Liu, F., L. Du, P. Du, and G. Huo. 2009. Possible promoter regions within the proteolytic system in Streptococcus thermophilus and their interaction with the CodY homolog. FEMS Microbiol. Lett. 297:164-172. Liu, M., A. Nauta, C. Francke, and R. J. Siezen. 2008. Comparative genomics of enzymes in flavor-forming pathways from amino acids in lactic acid bacteria. Appl. Environ. Microbiol. 74:4590-4600. 29 41. 42. 43. 44. 45. 46. 47. 48. 49. 50. 51. 52. 53. 54. Lu, C., F. Ding, A. Chowdhury, V. Pradhan, J. Tomsic, W. M. Holmes, T. M. Henkin, and A. Ke. 2010. SAM recognition and conformational switching mechanism in the Bacillus subtilis yitJ S box/SAM-I riboswitch. J. Mol. Biol. 404:803-818. Maddocks, S. E., and P. C. Oyston. 2008. Structure and function of the LysR-type transcriptional regulator (LTTR) family proteins. Microbiology 154:3609-3623. Madsen, S. M., B. Albrechtsen, E. B. Hansen, and H. Israelsen. 1996. Cloning and transcriptional analysis of two threonine biosynthetic genes from Lactococcus lactis MG1614. J. Bacteriol. 178:3689-3694. Malumbres, M., L. M. Mateos, C. Guerrero, and J. F. Martin. 1995. Molecular cloning of the hom-thrC-thrB cluster from Bacillus sp. ULM1: expression of the thrC gene in Escherichia coli and Corynebacteria, and evolutionary relationships of the threonine genes. Folia Microbiol. (Praha) 40:595-606. Martinez-Cuesta, M. C., C. Pelaez, J. Eagles, M. J. Gasson, T. Requena, and S. B. Hanniffy. 2006. YtjE from Lactococcus lactis IL1403 Is a C-S lyase with alpha, gamma-elimination activity toward methionine. Appl. Environ. Microbiol. 72:4878-4884. McDaniel, B. A., F. J. Grundy, I. Artsimovitch, and T. M. Henkin. 2003. Transcription termination control of the S box system: direct measurement of S-adenosylmethionine by the leader RNA. Proc. Natl. Acad. Sci. U S A 100:3083-3088. McDaniel, B. A., F. J. Grundy, V. P. Kurlekar, J. Tomsic, and T. M. Henkin. 2006. Identification of a mutation in the Bacillus subtilis S-adenosylmethionine synthetase gene that results in derepression of S-box gene expression. J. Bacteriol. 188:3674-3681. Merlin, C., G. Gardiner, S. Durand, and M. Masters. 2002. The Escherichia coli metD locus encodes an ABC transporter which includes Abc (MetN), YaeE (MetI), and YaeC (MetQ). J. Bacteriol. 184:5513-5517. Mrazek, J., L. H. Gaynon, and S. Karlin. 2002. Frequent oligonucleotide motifs in genomes of three Streptococci. Nucleic Acids Res. 30:4216-4221. Ohshima, H., S. Matsuoka, K. Asai, and Y. Sadaie. 2002. Molecular organization of intrinsic restriction and modification genes BsuM of Bacillus subtilis Marburg. J. Bacteriol. 184:381-389. Ohtsu, I., N. Wiriyathanawudhiwong, S. Morigasaki, T. Nakatani, H. Kadokura, and H. Takagi. 2010. The L-cysteine/L-cystine shuttle system provides reducing equivalents to the periplasm in Escherichia coli. J. Biol. Chem. 285:17479-17487. Overbeek, R., N. Larsen, T. Walunas, M. D'Souza, G. Pusch, E. Selkov, K. Liolios, V. Joukov, D. Kaznadzey, I. Anderson, A. Bhattacharyya, H. Burd, W. Gardner, P. Hanke, V. Kapatral, N. Mikhailova, O. Vasieva, A. Osterman, V. Vonstein, M. Fonstein, N. Ivanova, and N. Kyrpides. 2003. The ERGO (TM) genome analysis and discovery system. Nucleic Acids Res. 31:164-171. Parsot, C., and G. N. Cohen. 1988. Cloning and nucleotide sequence of the Bacillus subtilis hom gene coding for homoserine dehydrogenase. Structural and evolutionary relationships with Escherichia coli aspartokinases-homoserine dehydrogenases I and II. J. Biol. Chem. 263:14654-14660. Pelchat, M., and J. Lapointe. 1999. In vivo and in vitro processing of the Bacillus subtilis transcript coding for glutamyl-tRNA synthetase, serine acetyltransferase, and cysteinyl-tRNA synthetase. RNA 5:281-289. 30 55. 56. 57. 58. 59. 60. 61. 62. 63. 64. 65. 66. 67. 68. Pruitt, K. D., T. Tatusova, G. R. Brown, and D. R. Maglott. 2011. NCBI Reference Sequences (RefSeq): current status, new features and genome annotation policy. Nucleic Acids Res. Rajan, R., J. Zhu, X. Hu, D. Pei, and C. E. Bell. 2005. Crystal structure of S-ribosylhomocysteinase (LuxS) in complex with a catalytic 2-ketone intermediate. Biochemistry 44:3745-3753. Rodionov, D. A., A. G. Vitreschak, A. A. Mironov, and M. S. Gelfand. 2004. Comparative genomics of the methionine metabolism in Gram-positive bacteria: a variety of regulatory systems. Nucleic Acids Res. 32:3340-3353. Rosario, C. J., R. L. Frisch, and R. A. Bender. 2010. The LysR-type nitrogen assimilation control protein forms complexes with both long and short DNA binding sites in the absence of coeffectors. J. Bacteriol. 192:4827-4833. Sayers, E. W., T. Barrett, D. A. Benson, E. Bolton, S. H. Bryant, K. Canese, V. Chetvernin, D. M. Church, M. Dicuccio, S. Federhen, M. Feolo, L. Y. Geer, W. Helmberg, Y. Kapustin, D. Landsman, D. J. Lipman, Z. Lu, T. L. Madden, T. Madej, D. R. Maglott, A. Marchler-Bauer, V. Miller, I. Mizrachi, J. Ostell, A. Panchenko, K. D. Pruitt, G. D. Schuler, E. Sequeira, S. T. Sherry, M. Shumway, K. Sirotkin, D. Slotta, A. Souvorov, G. Starchenko, T. A. Tatusova, L. Wagner, Y. Wang, W. John Wilbur, E. Yaschenko, and J. Ye. 2010. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 38:D5-16. Schell, M. A. 1993. Molecular biology of the LysR family of transcriptional regulators. Annu. Rev. Microbiol. 47:597-626. Schildkraut, I., and S. Greer. 1973. Threonine synthetase-catalyzed conversion of phosphohomoserine to alpha-ketobutyrate in Bacillus subtilis. J. Bacteriol. 115:777-785. Sheppard, C. A., E. E. Trimmer, and R. G. Matthews. 1999. Purification and properties of NADH-dependent 5, 10-methylenetetrahydrofolate reductase (MetF) from Escherichia coli. J. Bacteriol. 181:718-725. Sierro, N., Y. Makita, M. de Hoon, and K. Nakai. 2008. DBTBS: a database of transcriptional regulation in Bacillus subtilis containing upstream intergenic conservation information. Nucleic Acids Res. 36:D93-96. Skarstedt, M. T., and S. B. Greer. 1973. Threonine synthetase of Bacillus subtilis. The nature of an associated dehydratase activity. J. Biol. Chem. 248:1032-1044. Sonenshein, A. L. 2005. CodY, a global regulator of stationary phase and virulence in Gram-positive bacteria. Curr. Opin. Microbiol. 8:203-207. Sperandio, B., C. Gautier, S. McGovern, D. S. Ehrlich, P. Renault, I. Martin-Verstraete, and E. Guedon. 2007. Control of methionine synthesis and uptake by MetR and homocysteine in Streptococcus mutans. J. Bacteriol. 189:7032-7044. Sperandio, B., C. Gautier, N. Pons, D. S. Ehrlich, P. Renault, and E. Guedon. 2010. Three Paralogous LysR-Type Transcriptional Regulators Control Sulfur Amino Acid Supply in Streptococcus mutans. J. Bacteriol. 192:3464-3473. Sperandio, B., P. Polard, D. S. Ehrlich, P. Renault, and E. Guedon. 2005. Sulfur amino acid metabolism and its control in Lactococcus lactis IL1403. J. Bacteriol. 187:3762-3778. 31 69. 70. 71. 72. 73. 74. 75. 76. 77. 78. 79. 80. Stenz, L., P. Francois, K. Whiteson, C. Wolz, P. Linder, and J. Schrenzel. 2011. The CodY pleiotropic repressor controls virulence in gram-positive pathogens. FEMS Immunol. Med. Microbiol. 62:123-139. Tang, D. W., L. F. Li, Y. M. Yu, X. Y. Liu, X. D. Su, X. Zhao, and Y. H. Liang. 2007. Preparation, crystallization and preliminary X-ray analysis of threonine synthase from Streptococcus mutans. Protein Pept. Lett. 14:836-838. Teusink, B., F. H. van Enckevort, C. Francke, A. Wiersma, A. Wegkamp, E. J. Smid, and R. J. Siezen. 2005. In silico reconstruction of the metabolic pathways of Lactobacillus plantarum: comparing predictions of nutrient requirements with those from growth experiments. Appl. Environ. Microbiol. 71:7253-7262. Thanbichler, M., B. Neuhierl, and A. Bock. 1999. S-methylmethionine metabolism in Escherichia coli. J. Bacteriol. 181:662-665. Tucker, B. J., and R. R. Breaker. 2005. Riboswitches as versatile gene control elements. Curr. Opin. Struct. Biol. 15:342-348. van der Ploeg, J. R., M. Barone, and T. Leisinger. 2001. Functional analysis of the Bacillus subtilis cysK and cysJI genes. FEMS Microbiol. Lett. 201:29-35. Vitreschak, A. G., D. A. Rodionov, A. A. Mironov, and M. S. Gelfand. 2004. Riboswitches: the oldest mechanism for the regulation of gene expression? Trends Genet. 20:44-50. Vitreschak, A. G., A. A. Mironov, V.A. Lyubetsky and M. S. Gelfand. 2004. Comparative genomic analysis of T-box regulatory systems in bacteria. RNA 14:717-735. Wels, M., T. Groot Kormelink, M. Kleerebezem, R. J. Siezen, and C. Francke. 2008. An in silico analysis of T-box regulated genes and T-box evolution in prokaryotes, with emphasis on prediction of substrate specificity of transporters. BMC Genomics 9:330. Xing, L., Y. Zhu, P. Fang, J. Wang, F. Zeng, X. Li, and M. Teng. 2011. Crystallization and preliminary crystallographic studies of UbiG, an O-methyltransferase from Escherichia coli. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 67:727-729. Zhang, Z., J. N. Feige, A. B. Chang, I. J. Anderson, V. M. Brodianski, A. G. Vitreschak, M. S. Gelfand, and M. H. Saier, Jr. 2003. A transporter of Escherichia coli specific for L- and D-methionine is the prototype for a new family within the ABC superfamily. Arch. Microbiol. 180:88-100. Zubieta, C., K. A. Arkus, R. E. Cahoon, and J. M. Jez. 2008. A single amino acid change is responsible for evolution of acyltransferase specificity in bacterial methionine biosynthesis. J. Biol. Chem. 283:7561-7567. 32 FIGURE LEGENDS Figure 1. Generalized cysteine and methionine metabolism in the Lactobacillales. For most of the studied species only part of the depicted reactions can take place as can be concluded from Tables 1 and 2. The map is divided in to differently coloured boxes on basis of the operon composition and EC numbers in line with the same Tables. Abbreviations: DMS, dimethyl sulfide; DMDS, dimethyl disulfide; and DMTS, dimethyl trisulfide. Figure 2. Binding motifs of various cysteine and methionine metabolism related regulators in the Lactobacillales. (A) The characteristic SMK-box motifs upstream of the metK gene in Lactobacilli and Streptococci as reported by (21). (B) Redefined MetR, CmbR and HomR dimer binding motifs in Lactobacilli and Streptococci. See the main text for details. (C) CodY binding motif in Lactobacilli and Streptococci. The motif was created on basis of the data provided in (11). The original data can be found in supplementary file 2. The motifs were created with WebLogo (frequency representation; no correction applied; http://weblogo.berkeley.edu). Figure 3. Bootstrapped (n=1000) partial neighbor-joining tree of LysR family transcription regulators in representative Lactobacillales. Only the branches of the CmbR, MetR/MtaR and HomR sub-families are shown. NCBI RefSeq GI codes precede the species/strain names (synchronized with Tables 1 and 2). Experimentally validated regulatory proteins are indicated by red dots. Abbreviations: S. = Streptococcus; L. = Lactococcus; Lb. = Lactobacillus; E. = Enterococcus. 33 Figure 4. Potential regulatory motifs related to the control of cys and met metabolism. The motifs were identified upstream of (A) the yrhAB operon in LAB; (B) thrB in LAB; and C) in L. lactis. The motifs are palindromic and therefore could relate to structure forming elements. The matching parts of the sequence have been underlined in different colours. The motifs were created with WebLogo (frequency representation; no correction applied; http://weblogo.berkeley.edu). 34 serine-acetyl transferase (cysE) homoserine O-acetyl transferase (metA) cysteine synthase (cysK/ytkP) cystathionine gamma synthase (yjcL) O-acetyl-homoserine sulfohydrolase (cysD) cystathionine beta synth. rev. pathway (yrhA) 1T C 1 cystathionine beta/gamma-lyase (yrhB) 1C 1 cystathionine beta/gamma lyase (yjcJ) PLP-dependent C-S lyase (patB) 1 2 4.4.1.8/ 4.4.1.1 4.4.1.8 /4.4.1.1 4.4.1.8 /4.4.1.1 1.5.1.20 5,10 methylenethf reductase (metF) 1 1 T 1 1 1(2) 1 1 1 1 1 1 1T 1 1 1T 1 2 T† 1(2 ) 1 1T 1 1 1 1 1 1T† 1 1T 1T 1 1 1 1T 1T† 1 1 1 1 1 1 1T 1 1 1T 1 1 1 1 1 1 1 1M 1 1T 1 1 3 1 1 1 1 1 1 1(2T) 1 2 1T 1 1 P. pentosaceus ATCC25745 2.3.1.30 2.3.1.31 2.5.1.47 2.5.1.48 2.5.1.49 4.2.1.22 1 1 1 1 1 1 O. oeni PSU-1 1 1 1 1 1(2) L. mesenteroides ATCC8293 1 1 1(2T†) 1 1 1 1 1 Lb. sakei 23K Lb. salivarius UCC118 1 Lb. rhamnosus GG 1 1 1 1 1 1(2) Lb. reuteri DSM20016 1(2T†) 1 1 1 1 1 Lb. plantarum WCFS1 ## 1 1 1 1 1(2) 1 1 1 1(2) 1(2) 1(2) 1 Lb. johnsonii NCC533 Lb. kefiranofaciens ZW3 1 0? 1 1 1 ? 1(2) 0 (2) Lb. crispatus ST1 1 1 1C Lb. casei ATCC334 homoserine dehydrogenase (hom) homoserine kinase (thrB) aspartate kinase III (thrA, BS_yclM) aspartate kinase I and II (lysC/dapG) threonine synthase (thrC) Lb. helveticus DPC4571 Lb. gasseri ATCC33323 Lb. fermentum IFO3956 Lb. delbr. bulg. ATCC BAA-365 ## Lb. buchneri NRRL B-30929 Lb. acidophilus NCFM 1.1.1.3 2.7.1.39 2.7.2.4 2.7.2.4 4.2.3.1 ECnumbe r enzyme names* Lb. brevis ATCC367 E. faecalis V583 # Lb. amylovorus GRL1112 TABLE 1 Presence and regulation of the genes encoding central cysteine and methionine metabolism in Enterococcus, Lactobacillus and Leuconostoc genomes. 1 1 1 1 0? 1 1 1 1 1(2) 1 1T 1T 1 1T 1T 1 1T 1 1 1 1 1T 1T 2 1T 35 2.1.1.10/ 1.5.1.20 2.1.1.10 2.1.1.14? 2.1.1.14 4.4.1.21 homocysteine S-methyltransferase 5,10 methylenethf reductase (yitJ) homocysteine S-methyltransferase (mmuM) amino acid permease (mmuP) methionine synthase (yxjH/yxjG) methionine synthase (metE) S-ribosylhomocysteinase (luxS) 2.1.1.37 cytosine-5-methyltransferase 1T 1 1T 1T 1T 1 1T(2) 1T 1 1 3.2.2.16/ 5'-methylthioadenosine nucleosidase (mtn) 3.2.2.9 1 1 1 1 1 1 1 1T 1T 1 1 1 1 1 1 2.5.1.6 S-adenosylmethionine synthetase (metK) 1S 1S 1S 1S 1S 1S 1S 1S 1S 1S 1S 1S 1S 1S Lipoprotein_9 (metQ-like) 2T(3) 2T 1T 1T 1T(2) 2T 2T 1T 2T(3) 1T 1T 1T 1T 2T 1 2T 1T 1T 1T 1 1T 1T 1T 1 1 1 T T 1 1 1T (3) 1 1T T T T 1 1 (2 ) 1 1T 1T 2T 1T 1T T 1 1T 1 1T 1 0? 1 1M 1T(2T) 1T 1 1T 1 T T T 1 (2 ) 1(2 ) 1T T 1 1 1T 1 1 1 1T 1 1T 1T 1T 1T 1 1 1 1 1 1 1 1 1 1 1 1S 1S 1S 1S 1s? 1s? 1S 3T 3T 1T 1T(2) 3 - The genes present in the same operon are indicated by a similar coloring of the cells and or by †. In case more than one closely related sequence was present, the total number is given. The number is put between brackets in case not all are present in the same operon and/or preceded by a similar putative binding-site. In case the gene might be present but was not called it is indicated by “0?”. The genes have been grouped in five clusters on basis of the composition of the operons and in case of shared EC numbers. - Putative regulator binding sites upstream of the indicated gene are given in Capitals in superscript: C, CmbR motif; M, MetR motif; N, CodY motif; S, SMK-box; T, T-box. Low-scoring putative binding sites are indicated by small caption and a question mark. In several cases we found putative binding sites for two regulators. - Species name abbreviations: E. = Enterococcus; Lb. = Lactobacillus; L. = Leuconostoc; O. = Oenococcus; P. = Pediococcus. Abbreviations in enzyme names: thf, tetrahydrofolate - The related data and NCBI gi-codes can be found in supplementary file 1 and the analysis of upstream regions in supplementary files 3 and 4. The original data is provided at www.cmbi.ru.nl/bamics/supplementary/Liuetal_2012_CysMetregulation. * For most enzymes the related gene names are provided. In most cases these represent names common in all Lactobacilli. In cases with little uniformity the names are derived from the orthologs found in the Bacillus subtilis genome (also see Materials and Methods). # The E. faecalis V583 genome has a copy of the cmbR gene (indicated by #), and the Lb. delbrueckii bulgaricus and Lb. plantarum genomes have a copy of the metR gene (indicated by ##). In Lb. plantarum the second copy of hom is associated with metA and cysD in an operon. 36 1 1 1T S. oralis Uo5 S. parasanguinis ATCC15912 S. parauberis KCTC11537 S. sanguinis SK36 S. suis 05ZYH33 1 1 1n? 1N 1 1 1n? 1 1 1 1N 1N 1N 1N 1 1N 1 1 1N 1N 1 1 1N 1N 1N 1N 1n? 1N 1N 1N 1n? 1 1 1 1M 1C 1H 1M 1 1M 1C 1M 1 1 1M 1C 1M 1 1 1M, 1 1M 1c? 1M 1 1 1M 1C 1H 1 1 1M 1c? 1C 1M 1 1M 1C 1M 1 1n? 1m? 1c? 1C 1M 1(2) 1M, H 1C 1H, N 1C 1 1 1 1 1H 1M 1 1M 1C 1M 1 H M M H M M M 1M 1C, N 1H, N 1 1M 1M 1M, C 1M 2.3.1.30 2.3.1.31 2.5.1.47 2.5.1.48 2.5.1.49 4.2.1.22 4.4.1.8/ 4.4.1.1 4.4.1.8 /4.4.1.1 4.4.1.8 /4.4.1.1 serine-acetyl transferase (cysE) homoserine O-acetyl transferase (metA) cysteine synthase (cysK/ytkP) cystathionine gamma synthase (yjcL) O-acetyl-homoserine sulfohydrolase (cysD) cystathionine beta synth. rev. pathway (yrhA) cystathionine beta/gamma-lyase (yrhB) cystathionine beta/gamma lyase (yjcJ) PLP-dependent C-S lyase (patB) 1 1 1 N 1 1C(2) 1C 1C 1N 1 1 5,10 methylenethf reductase (metF) homocysteine S-methyltransferase 5,10 2.1.1.10/ 1.5.1.20 methylenethf reductase (yitJ) 2.1.1.10 homocysteine S-methyltransferase (mmuM) 1C n? C 1C 1C S. uberis 0140J S. mutans UA159 1N 1N 1n? 1N 1 1 1n? 1 S. thermophilus CNRZ1066 S. mitis B6 1 1 1n? 1N 1N 1N 1N 1n? S. pyogenes M1-GAS S. gordonii Challis CH1 1N 1N 1N 1N Homoserine dehydrogenase (hom) Homoserine kinase (thrB) Aspartate kinase III (thrA) Threonine synthase (thrC) S. pneumoniae TIGR4 S. equi 4047 1 1 1 1N 1.1.1.3 2.7.1.39 2.7.2.4 4.2.3.1 S. pasteurianus ATCC43144 S. dysgalactiae GGS124 1N 1N 1N 1N S. agalactiae 2603V/R 1 1 1n? 1 Enzyme names Lc. lactis IL1403 1 1 1n? 1 ECnumber 1.5.1.20 S. gallolyticus ATCC_BAA2069 TABLE 2 Presence and regulation of the genes encoding central cysteine and methionine metabolism in Streptococcal genomes. 1C,n? 1 1 N 1 1 (2) 1 1M 1 1 1m? 1M 1 1M 1M 1M,H 1M,H 1m? 1M 1m? 1 H 1 (2) 1 1M 1M,H 1M 1M 37 2.1.1.14? 2.1.1.14 4.4.1.21 amino acid permease (mmuP) methionine synthase (yxjH/yxjG) methionine synthase (metE) S-ribosylhomocysteinase (luxS) 2.1.1.37 cytosine-5-methyltransferase 1 1M 1 1m? 1 1M 1 1 1M 1m? 1M 1 1 1 1 4 1 2 1 1 1 1 1 1 1S 1S 1S,C 1S,M 1S 3.2.2.16/ 3.2.2.9 5'-methylthioadenosine nucleosidase (mtn) 1 2.5.1.6 1S,n? 1S S-adenosylmethionine synthetase (metK) 1 1M 1M,N 1M,H 1 1m? 1M 1M,H 1M 1 1 1M 1M 1 1 N 1 1 1 1 1S 1S 1S,M 1S 1 1m? 1M 1 1M 1M 1M,C 1 1 1 1 1 1 1 1 1 1 1S, C 1S 1S 1M,N 1M,H 1 1 1 3 M 1S,M 1S 1M 2M(3) 1M 1 1 1 1 1S 1S - The genes present in the same operon are indicated by a similar coloring of the cells. In case more than one closely related sequence was present, the total number is given. The number is put between brackets in case not all are present in the same operon and/or preceded by a similar putative binding-site. In case the gene might be present but was not called it is indicated by “0?”. The genes have been grouped in five clusters on basis of the composition of the operons and in case of shared EC numbers. - Putative regulator binding sites upstream of the indicated gene are given in Capitals in superscript: C, CmbR motif; M, MetR motif; N, CodY motif; S, SMK-box; T, T-box. Low-scoring putative binding sites are indicated by small caption and a question mark. In several cases we found putative binding sites for two regulators. - Species name abbreviations: Lc. = Lactococcus; S. = Streptococcus. Abbreviations in enzyme names: thf, tetrahydrofolate - The related data and NCBI gi-codes can be found in supplementary file 1 and the analysis of upstream regions in supplementary file 3. The original data is provided at www.cmbi.ru.nl/bamics/supplementary/Liuetal_2012_CysMetregulation. * For most enzymes the related gene names are provided. In most cases these represent names common in all Lactobacilli. In cases with little uniformity the names are derived from the orthologs found in the Bacillus subtilis genome (also see Materials and Methods). 38 1 1 1 4 3.5.1.16 /3.5.1.18 (dapE-like) ABC_tran-NIL, BPD_transp_1 1 1C 1C 1 1M(2M) 1c? 1c? 1c? 2M 1c? 1c? 1c? 1M 1c? 1c? 2C(3) 1c? 1M 1m? 1C 1C C C 2 (3 ) 1C 1M 1M 1M 1M 1M 1M 1m? 1M 1M 1M 1M 1M 1M 1m? 1M 1M C 1 1M 1M 1 1m? 1C 1C 1C 1M 1C 1C 1C 1 1 (2 ) 1 1M 1M 1m? 1M 1 1M 1m? 1M 1 1M C C C c? 1M 1C 1C 1C 1C S. uberis 0140J S. thermophilus CNRZ1066 S. suis 05ZYH33 S. sanguinis SK36 S. pyogenes M1-GAS S. pneumoniae TIGR4 S. pasteurianus ATCC43144 S. parauberis KCTC11537 S. parasanguinis ATCC15912 S. oralis Uo5 S. mutans UA159 S. mitis B6 S. gordonii Challis CH1 S. gallolyticus ATCC_BAA2069 S. equi 4047 S. dysgalactiae GGS124 Lc. lactis IL1403 controlled transport systems SBP_bac_3 (tcyA/yxeM) BPD_transp_1, ABC_tran (tcyBC/yxeNO) SBP_bac_3 (tcyA/yxeM) Lipoprotein_9 (metQ-like) S. agalactiae 2603V/R TABLE 3 Presence and regulation of the cysteine and methionine ABC transport systems in Streptococcal genomes. 1 1C 1c? 1c? 1C 1c? C C 1 (2 ) 1C 1M(2) 1M 1C 0? 1M 1C 1M 1M - The genes present in the same operon are indicated by a similar coloring of the cells. In case more than one closely related sequence was present the total number is given. In case the gene might be present but was not called it is indicated by “0?”. The number is put between brackets in case not all are present in the same operon and/or preceded by a similar putative binding-site. - Putative regulator binding sites upstream of the indicated gene are given in Capitals in superscript: C, CmbR motif; M, MetR motif. Low-scoring putative binding sites are indicated by small caption and a question mark. In several cases we found putative binding sites for two regulators. - Species name abbreviations: Lc. = Lactococcus; S. = Streptococcus. - The related data and NCBI gi-codes can be found in supplementary file 1 and the analysis of upstream regions in supplementary file 3. The original data is provided at www.cmbi.ru.nl/bamics/supplementary/Liuetal_2012_CysMetregulation. 39