* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download PowerPoint Presentation - No Slide Title
Copy-number variation wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Behavioural genetics wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Metagenomics wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Gene expression programming wikipedia , lookup
Genome evolution wikipedia , lookup
Gene desert wikipedia , lookup
Genome (book) wikipedia , lookup
History of genetic engineering wikipedia , lookup
Human genome wikipedia , lookup
Ridge (biology) wikipedia , lookup
Designer baby wikipedia , lookup
Genomic imprinting wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Public health genomics wikipedia , lookup
Population genetics wikipedia , lookup
Minimal genome wikipedia , lookup
Pathogenomics wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene expression profiling wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Human genetic variation wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microevolution wikipedia , lookup
HLA A1-B8-DR3-DQ2 wikipedia , lookup
Molecular Inversion Probe wikipedia , lookup
A30-Cw5-B18-DR3-DQ2 (HLA Haplotype) wikipedia , lookup
Haplogroup G-M201 wikipedia , lookup
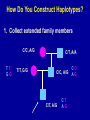
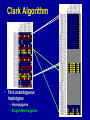
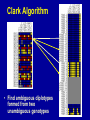
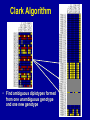
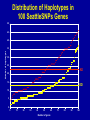
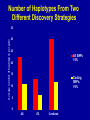
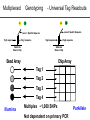
SNP Discovery and Genotyping Workshop • SNP discovery strategies Debbie Nickerson • Identifying SNPs by association for genotypephenotype analysis of candidate genes Chris Carlson • Identifying haplotypes for genotype-phenotype analysis of candidate genes Dana Crawford • SNP genotyping strategies Debbie Nickerson SNP Discovery and Genotyping Strategies Debbie Nickerson - [email protected] • Overview of Variation in the Human Genome • SNP Discovery Strategies and Status • SNP Data in the PGAs • Genotyping SNPs Total sequence variation in humans Population size: 6x109 (diploid) Mutation rate: 2x10–8 per bp per generation Expected “hits”: 240 for each bp Every variant compatible with life exists in the population BUT: Most are vanishingly rare Compare 2 haploid genomes: 1 SNP per 1331 bp* *The International SNP Map Working Group, Nature 409:928 - 933 (2001) Strategies to Find SNPs • Mine them from Existing Genome Resources • Targeted SNP Discovery in Candidate Genes Berkeley PGA - http://pga.lbl.gov CardioGenomics - http://www.cardiogenomics.org InnateImmunity - http://innateimmunity.net SeattleSNPs - http://pga.mbt.washington.edu Southwestern - http://pga.swmed.edu Sequence-based SNP Mining BAC library BAC Overlap Genomic DNA mRNA RRS Library or Sampling cDNA Library Shot gun Overlap EST Overlap Sequence Overlap SNP discovery GTTTAAATAATACTGATCA GTTTAAATAATACTGATCA GTTTAAATAGTACTGATCA GTTTAAATAGTACTGATCA ~ 4.1 Million SNPs Available http://www.ncbi.nlm.gov/SNP/ Mining Finds Only A Small Fraction of the SNPs Fraction of SNPs Discovered A 1.0 96 48 G 24 16 8 0.5 2 0.0 0.0 0.1 0.2 0.3 Minor Allele Frequency 0.4 0.5 Total Estimated SNPs and Fraction in dbSNP minimal allele expected SNPs expected SNP frequency (millions) frequency (bp) expected % in database 1% 11.0 290 11-12 5% 7.1 450 15-17 10% 5.3 600 18-20 20% 3.3 960 21-25 30% 2.0 1570 23-27 40% 0.97 3280 24-28 L. Kruglyak and D. Nickerson, Nat Genet 27:234-236 2001 Surfactant B - Locus Link dbSNP (http://www.ncbi.nlm.nih.gov/SNP/) Surfactant B - dbSNP Confirmation of SNP Resource in New Sample Potential Pitfalls Confirmed Multiple Method Report in dbSNP 100% 90% 80% 70% 60% 50% 40% 30% 20% 10% 0% Confirmed Unique Method Report in dbSNP Strategies to Find SNPs • Mine them from Existing Resources • Targeted SNP Discovery in Candidate Genes Berkeley PGA - http://pga.lbl.gov CardioGenomics - http://www.cardiogenomics.org InnateImmunity - http://innateimmunity.net SeattleSNPs - http://pga.mbt.washington.edu Southwestern - http://pga.swmed.edu Sequence-based SNP Identification Amplify DNA 5’ 3’ Sequence Phrap Phred Sequence each end Base-calling Contig assembly of the fragment. Quality determination Final quality determination PolyPhred ATAGACG ATAGACG ATACACG ATACACG ATAGACG ATACACG Polymorphism detection Consed Sequence viewing Polymorphism tagging Analysis Homozygotes Heterozygote Polymorphism reporting Individual genotyping Phylogenetic analysis Sequence-Based Detection and Genotyping of SNPs Jim Sloan, Tushar Bhangle (PolyPhred) Matthew Stephens, Paul Scheet (Quality Scores for SNPs) Phil Green, Brent Ewing, David Gordon (Phred, Phrap, Consed) PGA SNPs • The PGAs provide a validated SNP resource (Allele Frequency Data) • Novel Views of the Variation Data Emerging Pathway Interfaces Color Fasta Formats Gene Structure Views Visual Genotypes Linkage Disequilibrium Views TagSNPs Haplotypes • Many New Formats Under Development Toward comprehensive association studies • 5-7 million common variants exist in genome • Testing all for association is impractical today • Can the list be reduced w/o loss of power? – SNPs in Coding (Amino Acid Changes) – Linkage disequilibrium (SNPs in other functional regions, i.e. regulatory elements) cSNPs - Both Deep and Average Coverage Available from the PGAs CD36 - Southwestern PGA - Deep cSNP Discovery Strategy Healthy, High Cholesterol, High Triglycerides, Congential Cardiac Abnormalities, Left Ventricular Hypertrophy ……. CD36 - SeattleSNPs PGA - Average cSNP Discovery Strategy Healthy only SIFT (Sorting Intolerant From Tolerant) Coding Changes CYP4F2 Trp (W) Gly (G) Predicted to be tolerated Val (V) Gly (G) Predicted not to be tolerated Ng and Henikoff, Gen. Res. 2002 SNP-Based Association Studies Indirect: Use dense map of SNPs and test for linkage disequilibrium (use association to find sites in entire sequence (non-coding) with function) 5’ Arg-Cys Val-Val Collins, Guyer, Chakravarti Science 278:1580-81, 1997 3’ SNP Discovery and Genotyping Workshop • SNP discovery strategies Debbie Nickerson • Identifying SNPs by association for genotypephenotype analysis of candidate genes Chris Carlson • Identifying haplotypes for genotype-phenotype analysis of candidate genes Dana Crawford • SNP genotyping strategies Debbie Nickerson Selecting SNPs for Genotype-Phenotype Analysis Using Allelic Association (Linkage Disequilibrium) Christopher Carlson [email protected] Candidate Gene Association Analysis • Describe existing genetic variation – Rare SNPs (deep exonic resequencing) – Common SNPs (complete resequencing) • Select a subset of SNPs for genotyping – cSNPs (amino acid changes) – htSNPs (resolve haplotypes) – tagSNPs (patterns of genotype) • Test for genotype/phenotype correlations SeattleSNPs Resequencing Strategy I • Resequence the complete genomic region of each gene – 2000 bp upstream of first exon – 1500 bp downstream of poly-A signal – All exons and introns for genes below 35 kbp Image courtesy of GeneSNPs VG2 • Visual Genotype 2 – – – – Web interface Visualize genotypes View SNPs by frequency Sort on similarity between sites – Sort on similarity between samples – Visualize LD SeattleSNPs Resequencing Strategy II • Resequence candidate genes from inflammation and coagulation pathways • Resequence 47 individuals – 24 African American – 23 European American Homozygote common Heterozygote Homozygote rare Missing Data VG2 • Visual Genotype 2 – – – – Web interface Visualize genotypes View SNPs by frequency Sort on similarity between sites – Sort on similarity between samples – Visualize LD VG2 • Visual Genotype 2 – – – – Web interface Visualize genotypes View SNPs by frequency Sort on similarity between sites – Sort on similarity between samples – Visualize LD VG2 • Visual Genotype 2 – – – – Web interface Visualize genotypes View SNPs by frequency Sort on similarity between sites – Sort on similarity between samples – Visualize LD VG2 • Visual Genotype 2 – – – – Web interface Visualize genotypes View SNPs by frequency Sort on similarity between sites – Sort on similarity between samples – Visualize LD VG2 • Visual Genotype 2 – – – – Web interface Visualize genotypes View SNPs by frequency Sort on similarity between sites – Sort on similarity between samples – Visualize LD VG2 • Visual Genotype 2 – – – – Web interface Visualize genotypes View SNPs by frequency Sort on similarity between sites – Sort on similarity between samples – Visualize LD Preliminary Analyses • Hardy Weinberg Equilibrium • Population specificity • Nucleotide diversity • Pop genetics statistics (e.g. Tajima’s D) SNP Selection: cSNPs • Genotype SNPs which change amino acids • Genotype other “good story” SNPs – SNPs in known regulatory elements – SNPs in Conserved Noncoding Sequences Image courtesy of GeneSNPs SNP Selection: htSNPs • Genotype “haplotype tagging” SNPs which resolve existing common haplotypes SNP Selection: htSNPs • Genotype “haplotype tagging” SNPs which resolve existing common haplotypes SNP Selection: tagSNPs • Resequence a modest number of samples – Describe patterns of genotype at all common SNPs – Genotype tagSNPs which efficiently capture existing patterns of genotype Linkage Disequilibrium A B Haplotype is the pattern of alleles on a single chromosome – 4 possible haplotypes Linkage Disequilibrium (LD) describes the allelic association between two SNPs Two popular LD statistics: D´ r2 Complete LD A B Unequal allele frequency Allelic association is as strong as possible – 3 haplotypes observed – No detected recombination between SNPs – Genotype is not perfectly correlated D´ = 1 r2 < 1 Perfect LD A B Equal allele frequency Allelic association is as strong as possible – 2 haplotypes observed – No detected recombination between SNPs – Genotype is perfectly correlated D´ = 1 r2 = 1 Rational SNP Selection Select SNPs to genotype on the basis of LD • Some SNPs are in LD with many other SNPs • Some SNPs are in LD with no other SNPs • SNPs between a pair of associated SNPs are not necessarily associated with the flanking SNPs LD SNP Selection Example CSF3 in European Americans •5200 bp •17 SNPs LD SNP Selection Example CSF3 in European Americans •5200 bp •17 SNPs •10 common SNPs (above 10% minor allele frequency) LD Site Selection Algorithm • Find minimal set of SNPs for assay, such that each SNP is either assayed directly or above r2 threshold with an assayed SNP •Calculate all pairwise r2 values •Set r2 threshold based on power estimates for study LD Site Selection Algorithm • Find minimal set of SNPs for assay, such that each SNP is either assayed directly or above r2 threshold with an assayed SNP •Calculate all pairwise r2 values •Set r2 threshold based on power estimates for study CSF3 Site Selection • Threshold LD: r2 > 0.64 – Bin 1: 4 sites – Bin 2: 4 sites – Bin 3: 2 sites • Genotype 1 SNP from each bin, chosen for biological intuition or ease of assay design Power and LD • Given – All common SNPs described – Patterns of LD between common SNPs are known • Select SNPs such that every SNP is either – Directly assayed – Associated with an assayed SNP • Test for disease associations with assayed SNPs • Power to detect disease associations at unassayed SNPs depends on r2 between assayed and unassayed SNPs LD Selection and Haplotype • LD selected SNPs provide the highest possible haplotype diversity for a given number of SNPs assayed • LD selection is robust to recombination and hotspot structure • LD selection is sensitive to population stratification SNP Selection Summary • It is possible to test all common variants in a candidate gene directly for risk association (main effects) with meaningful null negative results • Caveat: Higher order risks unaddressed – Haplotype (G X G effects within a locus) – Epistasis (G X G effects between loci) – Environment (G X E effects) SNP Discovery and Genotyping Workshop • SNP discovery strategies Debbie Nickerson • Identifying SNPs by association for genotypephenotype analysis of candidate genes Chris Carlson • Identifying haplotypes for genotype-phenotype analysis of candidate genes Dana Crawford • SNP genotyping strategies Debbie Nickerson Identifying Haplotypes for Genotype-Phenotype Analysis Dana C. Crawford [email protected] Outline of discussion • Constructing or inferring haplotypes • Haplotype tools available in PGA • Description of haplotypes in SeattleSNPs genes • Use of VH1 tool to visually inspect – Haplotype blocks – Haplotype diversity – Hotspots of recombination • Summary of SeattleSNPs haplotype data What is a Diplotype ? • Humans are diploid • At each SNP there are two alleles, which are observed as a genotype • At each gene there are two haplotypes, which are observed as a multi-site genotype, or diplotype What is a Haplotype? A: “…a unique combination of genetic markers present in a chromosome.” pg 57 in Hartl & Clark, 1997 VH1 – haplotype visualization tool How Do You Construct Haplotypes? 1. Collect extended family members C/C, A/G TT GG T/T, G/G C/T, A/A C/C, A/G C/T, A/G CT AG CC AG How Do You Construct Haplotypes? 2. Go from diploid to haploid via somatic cell hybrids e.g. Patil et al 2001 How Do You Construct Haplotypes? 3. Allele-specific PCR SNP 1 C/T SNP 2 A/G How Do You Construct Haplotypes? 4. Statistical inference • Clark Algorithm • EM (Arlequin) • Phase Ligation (HAPLOTYPER) • PHASE Clark Algorithm • Find unambiguous haplotypes – Homozygotes – Single Heterozygotes Clark Algorithm • Find ambiguous diplotypes formed from two unambiguous genotypes Clark Algorithm • Find ambiguous diplotypes formed from one unambiguous genotype and one new genotype Clark Algorithm • Iterate until either all haplotypes resolve, or ambiguous haplotypes are inconsistent with any inferred haplotype Haplotype Algorithm Comparison • Clark – Intuitive – Fast • PHASE – Complete solution – Slightly more accurate than EM – Slow version 2 faster • EM – Complete solution – Slightly more accurate than Clark – Robust to ambiguity • Haplotyper (Ligation) – Fast – Better than Clark – Less accurate than EM or PHASE Haplotype Tools in the PGA InnateImmunity • 25 genes re-sequenced in innate immunity pathway • 4 populations: European and African-Americans, Hispanics, Asthmatics • PHASE and Haplotyper results posted on website http://innateimmunity.net Haplotype Tools in the PGA SeattleSNPs • • • • 120 genes re-sequenced in inflammation response 2 populations: European- and African-Americans PHASE results posted on website Interactive tool (VH1) to visualize and sort haplotypes http://pga.gs.washington.edu Distribution of Haplotypes in 100 SeattleSNPs Genes 50 45 40 Number of haplotypes 35 30 25 AD 20 15 ED 10 5 0 0 10 20 30 40 50 60 Number of genes 70 80 90 100 Common Haplotypes in 100 SeattleSNPs Genes (Frequency >5%) Population >5% MAF Average Range ED 4.54 1-8 AD 4.99 0 - 11 Haplotype Sharing Between Populations in 100 SeattleSNPs Genes 1 0.9 0.8 0.7 0.6 0.5 0.4 0.3 0.2 0.1 0 Non-shared Shared ED AD Number of Haplotypes From Two Different Discovery Strategies Average number of haplotypes per gene 35 30 25 All SNPs >5% 20 15 Coding SNPs, >5% 10 5 0 AD ED Combined Haplotype Structures Are Similar Across Discovery Strategies… FGB – African-Americans Coding SNPs 29 SNPs >5% 13 SNPs >5% …But, Not For All Genes F10 – African-Americans Coding SNPs 48 SNPs >5% 13 SNPs >5% Are Blocks Preserved Using Different Discovery Strategies? Four-gamete test: A B a b A B a A b b a B Yes*, for some: 10% of genes in AD 25% of genes in ED Fewer “blocks” with fewer SNPs/kb HaploBlockFinder; Zhang and Jin 2003 *>75% of the blocks are preserved Using Visualization Tools (VH1) To Identify Haplotype Blocks IL10: • Rare sites removed • Sorted by related sites • “Block” structure evident Using VH1 to Identify Highly Divergent Haplotypes • Some haplotypes are highly divergent • More likely to have functional consequences? • Mixed Blessing: – Easier to detect – Harder to dissect Using Haplotypes To Identify Hotspots Of Recombination CD36 haplotypes, sorted by sample Linkage Disequilibrium and Hotspots Associated Sites CD36 Hotspot in between sites need to be typed from both ends Detection of Recombination Hotspots In Candidate Genes HOTSPOTTER • Developed by Na Li and Matthew Stephens • Multilocus model for LD: Does not rely on “block-like” patterns Relates LD to underlying recombination process Incorporated into new version of PHASE (v2.0) students.washington.edu/lina/software/ CD36 – combined population CD36 – AD and ED populations HOTSPOTTER Preliminary Results 15 out of 100 genes have evidence of a hotspot: AGTR1 APOB CD36 IL1B IL21R IL4 NOS3 PLAUR PON1 SERPIN45 SELP SFPA2 SFTPB VCAM1 VEGF SeattleSNPs Haplotype Summary • More haplotypes per gene than previously described • <50% of African-American chromosomes are represented by common shared haplotypes • Block structure is preserved across discovery strategies for only a fraction of the genes • Evidence for hotspots of recombination in human genes SNP Discovery and Genotyping Workshop • SNP discovery strategies Debbie Nickerson • Identifying SNPs by association for genotypephenotype analysis of candidate genes Chris Carlson • Identifying haplotypes for genotype-phenotype analysis of candidate genes Dana Crawford • SNP genotyping strategies Debbie Nickerson Ideals for SNP Genotyping • High Sensitivity - PCR but moving towards direct genomic DNA detection • High Specificity - Accurate • Simple process - Easy to automate - High Throughput • Multiplexing - Perform many assays at once - decrease costs • Cheap SNP Genotyping Matched Probe and Target C Allele C Target C G Mis-Matched T Allele C Allele-Specific Hybridization Hybridize +ddCTP Polymerase Extension Target Oligonucleotide Ligation C Target Invader Taqman C Target C Target A Fail to hybridize C G A C incorporat ed C Fails t o incorporate C G A Ligate Fail to ligat e C G A Cleave Fail to cleave C G Degrade C A Fail t o degrade C Allele-Specific PCR C Target G A Amplify Fail to amplify SNPTyping Formats Microtiter Plates - Fluorescence eg. Taqman - Good for a few markers - lots of samples - PCR Size Analysis by Mass or Electrophoresis eg. Sequenom or SnapShot - Moderate Multiplexing reducing costs Arrays - Custom or Universal eg. Affymetrics, Illumina or ParAllele - Highly multiplexed HighThroughput - Genotype directly on genomic DNA Taqman Genotyping with fluorescence-based homogenous assays (single-tube assay) A G Genotype Calling - Cluster Analysis Genotyping by Mass Spectrometry Multiplex ~ 5 SNPs Comparative Genotyping in Populations Pooled DNA Population 1 PCR Pooled DNA Quantitative Assay Estimate Allele Frequency Polymorphism 60/40 Pooled DNA Population 2 PCR Pooled DNA Quantitative Assay Estimate Allele Frequency Polymorphism 85/15 Pooled Genotyping Advantages: Speed, Cost Major Disadvantages: Loss of haplotype information Loss of stratification by phenotype or environmental factors SNP Genotyping Custom SNP Genotyping Chips: Multiplexed C Genotyping - Universal Tag Readouts G A T Locus 2 Specific Sequence Locus 1 Specific Sequence Tag1 sequence Tag2 sequence cTag1 sequence cTag2 sequence Subst rate Bead or Chip Substrate Bead or Chip Bead Array Chip Array Tag 1 Tag 2 Tag 3 Tag 4 Illumina Multiplex ~1,000 SNPs Not dependent on primary PCR ParAllele Illumina Genotyping - Gap Ligation 1,000 SNPs Assayed on 96 Samples SNP Genotyping Lots of systems - Still costly but dropping Offering Moderate to High throughputs Systems vary in price $$ -$$$$ Laboratory Information Management Systems (Key: Track - Samples, - Assays - Completion rate - Reproducibility/Error Analysis)