* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download (A) and B chains - Michael P. Ready
Agarose gel electrophoresis wikipedia , lookup
Paracrine signalling wikipedia , lookup
Gene expression wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Signal transduction wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Peptide synthesis wikipedia , lookup
Size-exclusion chromatography wikipedia , lookup
Expression vector wikipedia , lookup
Magnesium transporter wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Interactome wikipedia , lookup
Biosynthesis wikipedia , lookup
Genetic code wikipedia , lookup
Metalloprotein wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Gel electrophoresis wikipedia , lookup
Point mutation wikipedia , lookup
Protein purification wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Protein–protein interaction wikipedia , lookup
List of incidents involving ricin wikipedia , lookup
Biochemistry wikipedia , lookup
CH339K
Proteins: Primary Structure, Purification, and
Sequencing
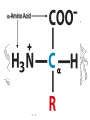
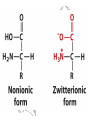
a-Amino Acid
a
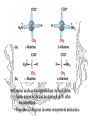
•All amino acids as incorporated are in the L-form
• Some amino acids can be changed to D- after
incorporation
• D-amino acids occur in some non-protein molecules
I prefer this layout, personally…
HOOC
R C H
NH2
L-amino acid
HOOC
H
C
R
NH2
D-amino acid
2 Amides
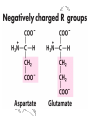
The Acidic and the Amide Amino Acids Exist
as Conjugate Pairs
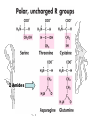
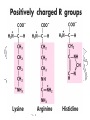
Ionizable Side Chains
Hydrogen Bond Donors / Acceptors
Disulfide formation
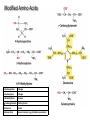
Modified Amino Acids
4-Hydroxyproline
Collagen
5-Hydroxylysine
Collagen
6-N-Methyllysine
Histones
g-Carboxygultamate
Clotting factors
Desmosine
Elastin
Selenocysteine
Several enzymes (e.g. glutathione peroxidase)
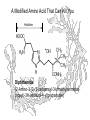
A Modified Amino Acid That Can Kill You
Histidine
Diphthamide
(2-Amino-3-[2-(3-carbamoyl-3-trimethylammoniopropyl)-3H-imidazol-4-yl]propanoate)
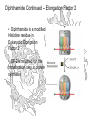
Diphthamide Continued – Elongation Factor 2
• Diphthamide is a modified
Histidine residue in
Eukaryotic Elongation
Factor 2
• EF-2 is required for the
translocation step in protein
synthesis
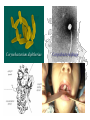
Corynebacterium diphtheriae
Corynebacteriophage
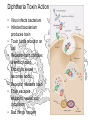
Diphtheria Toxin Action
• Virus infects bacterium
• Infected bacxterium
produces toxin
• Toxin binds receptor on
cell
• Receptor-toxin complex
is endocytosed
• Endocytic vessel
becomes acidic
• Receptor releases toxin
• Toxin escapes
endocytic vessel into
cytoplasm
• Bad things happen
Diphtheria Toxin Action
• Diphtheria toxin
adds a bulky group
to diphthamide
• eEF2 is inactivated
• Cell quits making
protein
• Cell(s) die
• Victim dies
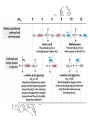
Other Amino Acids
Every a-amino acid has at
least 2 pKa’s
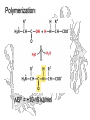
Polymerization
DG0’ = +10-15 kJ/mol
In vivo, amino acids are
activated by coupling to
tRNA
Polymerization of activated
a.a.:
DGo’ = -15-20 kJ/mol
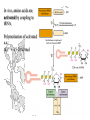
• In vitro, a starting amino acid
can be coupled to a solid matrix
• Another amino acid with
• A protected amino group
• An activating group at the
carboxy group
• Can be coupled
• This method runs backwards
from in vivo synthesis (C N)
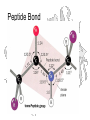
Peptide Bond
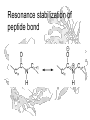
Resonance stabilization of
peptide bond
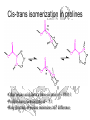
Cis-trans isomerization in prolines
•Other amino acids have a trans-cis ratio of ~ 1000:1
•Prolines have cis:trans ratio of ~ 3:1
•Ring structure of proline minimizes DG0 difference
Physical Methods
or
How to Purify and Sequence a
Weapons-Grade Protein
First Question
How do I measure the amount
of protein I have?
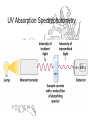
UV Absorption Spectrophotometry
Beer-Lambert Law
I
Transmitta nce : T 10cl
Io
or alternativ ely
Absorbance : A log T cl
c = concentration
l = path length
= extinction coefficient
An Absorbance = 2 means that only 1% of the incident beam is
getting through.
Transmittance and Absorbance
Absorbance vs. Concentration
Transmittance vs. Concentration
Second Question
How can I spot my protein in the
great mass of different proteins?
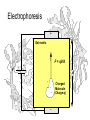
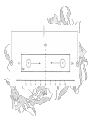
Electrophoresis
+
Gel matrix
V
F = qV/d
+
-
d
d
-
-
-
Charged
Molecule
(Charge q)
V
F f q qE
d
q charge
E field strength
Fb fv
f frictional coefficien t
v velocity
At equilibriu m :
fv qE
or
v q
E f
v q
E f
The frictional coefficient f depends on the size of the
molecule, which in turn depends upon the molecular mass,
so:
q
va
M
i.e. the velocity depends on the charge/mass ratio, which
varies from protein to protein
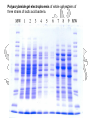
Polyacrylamide Gels
Polyacrylamide gel electrophoresis of whole cell proteins of
three strains of lactic acid bacteria.
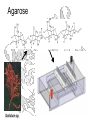
Agarose
Gelidium sp.
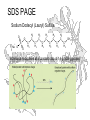
SDS PAGE
Sodium Dodecyl (Lauryl) Sulfate
O
+
Na O
H2
C
O
S
O
C
H2
H2
C
C
H2
H2
C
C
H2
H2
C
C
H2
H2
C
C
H2
C
H2
CH3
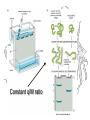
SDS binds to proteins at a constant ratio of 1.4 g SDS/g protein
Constant q/M ratio
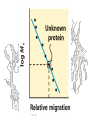
1
Rf
logM r
Disulfide cleavage
Disulfide cleavage and chain separation
+ bME
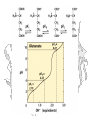
Isoelectric Point
Abrin A - Predicted Charge
30
20
Predicted pI
5.088
Charge on Protein
10
0
-10
-20
-30
-40
0.0
2.0
4.0
6.0
8.0
pH
10.0
12.0
14.0
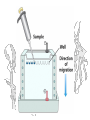
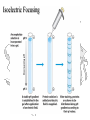
Isoelectric Focusing
pH
Carrier Ampholytes
• Amphoteric Electrolytes
• Mixture of molecules containing multiple
amino- and carboxyl- groups with
closely spaced pIs
• Partition into a smooth, buffered pH
gradient
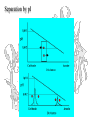
Separation by pI
Isoelectric Focusing
Below the pI, a protein has a positive charge and migrates
toward the cathode
Above the pI, a protein has a negative charge and migrates
toward the anode
Isoelectric Focusing
Foot Flesh Extracts from Pomacea flagellata and Pomacea patula
catemacensis
STOP
HERE
Protein Purification Steps
1 unit = amount of enzyme that catalyzes
conversion of 1 mmol of substrate to product in 1
minute
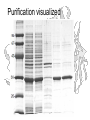
Purification visualized
Example:
Purification of Ricin
Georgi Markov
1929-1978
Ricinus communis – castor oil
plant
Ricin
Ricin B chain
(the attachment bit)
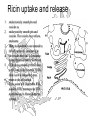
Ricin uptake and release
1.
2.
3.
4.
5.
6.
7.
endocytosis by coated pits and
vesicles or,
endocytosis by smooth pits and
vesicles. The vesicles fuse with an
endosome.
Many ricin molecules are returned to
the cell surface by exocytosis, or
the vesicles may fuse to lysosomes
where the ricin would be destroyed.
If the ricin-containing vesicles fuse
to the Trans Golgi Network, (TGN),
there ís still a chance they may
return to the cell surface.
Toxic action will occur when RTA,
aided by RTB, penetrates the TGN
membrane and is liberated into the
cytosol.
Ricin Action
• Ricin and
related
enzymes
remove an
adenine base
from the large
ribosomal RNA
• Shut down
protein
synthesis
The possibility that
ricin might be used
as an asymmetric
warfare weapon has
not escaped the
attention of the
armed services.
The last time I was
qualified to know for
sure, there were no
effective antidotes.
Significant Terrorist Incidents Involving Chemical and Biological Agents
Year
1946
1970
1972
1974
1980
1984
1991
1990-1995
1995
1995
1998
2001
2003-2004
Organization
Agents
DIN
Arsenic Compounds
("Revenge" in Hebrew; also
Dahm Y'Israel Nokeam, "Avenging
Israel's Blood")
(Germany)
Weather Underground
Tried to obtain agents from Ft. Detrick by
(United States)
blackmailing a homosexual serviceman.
R.I.S.E
Typhoid, diphtheria, dysentery, meningitis and several
(United States)
others to be delivered by aerosol.
Aliens of America
Nerve Agents
(Alphabet Bomber)
(United States)
R.A.F.
Botulinum toxin
(Rote Armee Faktion)
(Germany)
Rajneshee Cult (United States)
Salmonella enterica serovar typhimurium
Minnesota Patriots Council
(United States)
Aum Shinrikyo
(Japan)
Aryan Nation
(United States)
The Covenant and the Sword
(United States)
Republic of Texas
(United States)
Unknown (United States)
Fallen Angel (United States)
Ricin
Bacteria and viral agents, toxins, organophosphorus
nerve agents.
Yersinia pestis
Ricin
Bacterial and viral agents
Bacillus anthracis
Ricin
Raw
Extract
(NH4)2SO4
Cut
Affinity
Gel
Filtration
Salting In – Salting out
n
1 2
Ionic Strength : I ci zi
i 1 2
ci concentrat ion of ion i
zi charge on ion i
• salting in: Increasing ionic strength increases protein
solubility
• salting out: Increasing further leads to a loss of solubility
Salting in – salting out
The solubility of haemoglobin in different electrolytes as a
function of ionic strength.
Derived from original data by Green, A.A. J. Biol. Chem. 1932, 95, 47
Salting in: Counterions
help prevent formation of
interchain salt links
Solubility reaches
minimum at pI
Salting out: there’s simply less water available to solubilize
the protein.
Different proteins have different
solubilities in (NH4)2SO4
Lyotropic ChaotropicSeries
Cations: N(CH3)3H+> NH4+> K+> Na+> Li+> Mg2+>Ca2+> Al3+>
guanidinium / urea
Anions: SO42−> HPO42−> CH3COO−> citrate > tartrate > F−> Cl−>
Br−> I−> NO3−> ClO4−> SCN−
1) Bring to 37% Saturation – ricin still soluble, many other
proteins ppt
2) Collect supernatant
3) Bring to 67% Saturation – ricin ppt, many remaining
proteins still soluble
4) Collect pellet
5) Redissolve in buffer
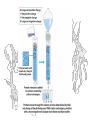
Dialysis and Ultrafiltration
(How do you get the %@$&#! salt out?)
Raw
Extract
(NH4)2SO4
Cut
Affinity
Gel
Filtration
Separation by chromatography
Basic Idea:
You have a stationary phase
You have a mobile phase
Your material partitions out
between the phases.
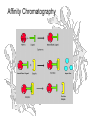
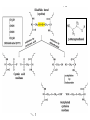
Affinity Chromatography
Structure of Agarose
Agarose is a polymer of agarobiose, which in turn consists of one
unit each of galactose and 3,6-anhydro-a-L-galactose.
Ricin sticks to galactose, so store-bought agarose acts as an
affinity column right out of the bottle, with ricin binding the beads
while other proteins wash through.
Begin adding 0.2 M
Lactose
Raw
Extract
(NH4)2SO4
Cut
Affinity
Gel
Filtration
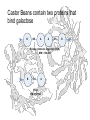
Castor Beans contain two proteins that
bind galactose
B
SS
A
A
SS
Ricinus communis Agglutinin (RCA)
MW = 120,000
B
SS
A
Ricin
MW = 60,000
B
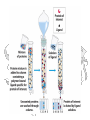
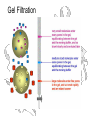
Gel Filtration
Gel Filtration
Gel Filtration (aka Size
Exclusion)
You knew I couldn’t leave it at that…
Vm = matrix volume
Vo = void volume
Vp = pore volume
Vt = total volume
Ve = elution volume
(1a) Vt = Vo + Vp or
(1b) Vp = Vt - Vo
(2) Ve = Vo + Kav*Vp
Combining 1b with 2
Ve Vo
K av
Vt V0
• a and b represent the effective separation range
• c corresponds to the exclusion limit
Kav
Note: smaller = slower,
whereas in SDS-PAGE,
smaller = faster.
Note
Fig. 3. Measurement of molecular weight of native NAGase enzyme of green crab by gel
filtration on Sephadex G-200: standard proteins (empty circles); green crab NAGase (filled
circle).
From Zhang, J.P., Chen, Q.X., Wang, Q., and Xie, J.J. (2006) Biochemistry (Moscow) 71(Supp. 1)
855-859.
Gel Filtration Separation of Ricin
Ricin
RCA
Raw
Extract
(NH4)2SO4
Cut
Affinity
Gel
Filtration
Okay, Now Let’s Sequence
the A-Chain
Bovine Insulin
21 residue A chain
31 residue B chain
Connected by disulfides
In order to sequence the protein, the
chains have to be separated
Chain Separation
• Interchain disulfide broken by high
concentrations of bME
• Chains are about the same size – but
can take advantage of different pIs
– B-Chain
– A-Chain
pI ~ 5.3
pI ~ 7.2
Ion Exchangers
•Apply bME – treated ricin to DEAE-cellulose at pH 7
•At pH 7:
•A chain (pKa 7.2) is essentially uncharged,
•B chain (pKa 4.8) is highly negative
•A chain washes through the column
•B chain sticks, eluted with gradient of NaCl
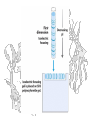
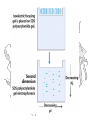
2-D Electrophoresis (an aside)
• Can use two different properties of a
protein to separate electrophoretically
• For analysis of cellular protein content,
often use 2-dimensional
electrophoresis:
• 1st dimension is isoelectric focusing
• 2nd dimension is SDS PAGE
2-D Electrophoresis (cont.)
• Can use other protein properties to
separate
– Simple PAGE at 2 different pHs
– PAGE and SDS PAGE
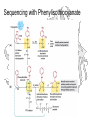
Sequencing with Phenylisothiocyanate
• Applied Biosystems 492 Procise Protein
Sequencer
Chain Cleavage: Cyanogen Bromide
C-Terminal Sequencing
• Carboxypeptidases are enzymes that
chew proteins from the carboxy
terminus
• Can incubate a protein (preferably
denatured – more later) with a
carboxypeptidase
• Remove aliquot at intervals (time
course)
• Run amino acid analysis of aliquots
C-Terminal Sequencing of Rat Plasma
Selenoprotein
From Himeno et al (1996) J. Biol. Chem. 271: 15769-15775.
Tandem Mass Spectrometry can also be used to
determine peptide sequences
MOLECULAR EVOLUTION
Sequence differences among vertebrate
Time of Divergence
hemoglobins
|-------------|-------------|------------|------------|------------|------------|
┌───────────────────────────────Shark
│
│
┌─────────────────────Perch
└─────────┤
│
┌─────────────Alligator
└───────┤
│
┌──────Horse
└──────┤
│
┌───Chimp
└──┤
│
└───Human
|-------------|-------------|------------|------------|-----------|------------|------------|------------|
Sequence Difference
Neutral Theory of Molecular Evolution
• Kimura (1968)
• Mutations can be:
– Advantageous
– Detrimental
– Neutral (no good or bad phenotypic effect)
• Advantageous mutations are rapidly
fixed, but really rare
• Diadvantageous mutations are rapidly
eliminated
• Neutral mutations accumulate
What Happens to a Neutral
Mutation?
• Frequency subject to random chance
• Will carrier of gene reproduce?
• Many born but few survive
– Partly selection
– Mostly dumb luck
• Gene can have two fates
– Elimination (frequent
– Fixation (rare)
Genetic Drift in Action
Our green
genes are
evolutionarily
superior!
Ow!
Never
mind…
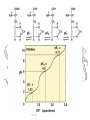
Simulation of Genetic Drift
• 100 Mutations x 100 generations:
• 1 gets fixed
• 2 still exist
• 97 eliminated (most almost immediately)
1
Frequency
0.8
0.6
0.4
0.2
0
0
25
50
Generation
75
100
Rates of Change
Overall Rate RT RM RF
where :
RM mutation rate
RF fixation rate
and RM and RF are both related to population size N
RM a N
1
N
Therefore population size cancels out.
R T depends only on the probabilit y of mutation t imes the probabilit y of fixation.
RF a
Therefore change accumulati on is CONSTANT.
Therefore change accumulati on can be a MOLECULAR CLOCK
Protein Evolution Rates
Different proteins have different rates
Protein Evolution Rates
Different proteins have different rates
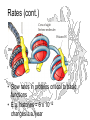
Rates (cont.)
• Slow rates in proteins critical to basic
functions
• E.g. histones ≈ 6 x 10-12
changes/a.a./year
Rates (cont.)
Fibrinopeptides
• Theoretical max
mutation rate
• Last step in blood
clotting pathway
• Thrombin converts
fibrinogen to fibrin
Fibrinopeptides keep fibrinogens from sticking together.
Rates (cont.)
• Only constraint on sequence is that it has to
physically be there
• Fibrinopeptide limit ≈ 9 x 10-9 changes/a.a./year
Relationships
among plant
hemoglobins
Arredondo-Peter, Raul,
et al (1998) Plant
Physiol. 118: 1121-1125
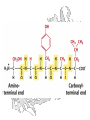
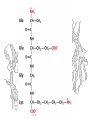
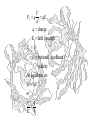
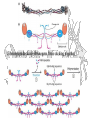
Amino acid sequences of
several ribosome-inhibiting
proteins
Phylogenetic trees built from
the amino acid sequences of
type 1 RIP or A chains (A)
and B chains (B) of type 2
RIP (ricin-A, ricin-B, and lectin RCAA and RCA-B from castor bean;
abrin-A, abrina/b-B, and agglutinin
APA-A and APA-B from A.
precatorius; SNAI-A and SNAI-B,
SNAV-A and SNAV-B, SNAI'-A and
SNAI'-B, LRPSN1-A and LRPSN1-B,
LRPSN2-A and LRPSN2-B, and SNAIV from S. nigra; sieboldinb-A,
sieboldinb-B, SSAI-A, and SSAI-B
from S. sieboldiana; momordin and
momorcharin from Momordica
charantia; MIRJA from Mirabilis
jalapa; PMRIPm-A and PMRIPm-B,
PMRIPt-A and PMRIPt-B from
Polygonatum multiflorum;
RIPIriHol.A1, RIPIriHol.A2, and
RIPIriHol.A3 from iris hybrid; IRAr-A
and IRAr-B, IRAb-A and IRAb-B from
iris hybrid; SAPOF from S.
officinalis; luffin-A and luffin-B from
Luffa cylindrica; and karasurin and
trichosanthin from Trichosanthes
kirilowii)
Hao Q. et.al. Plant Physiol. 2010:125:866-876
Phylogenetic tree of Opisthokonts, based on nuclear
protein sequences
Iñaki Ruiz-Trillo, Andrew J. Roger, Gertraud Burger, Michael W. Gray & B. Franz
Lang (2008) Molecular Biology and Evolution, Jan 9