* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download LECT35 trans1
Nicotinamide adenine dinucleotide wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Deoxyribozyme wikipedia , lookup
RNA silencing wikipedia , lookup
Catalytic triad wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Citric acid cycle wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Point mutation wikipedia , lookup
Metalloprotein wikipedia , lookup
Polyadenylation wikipedia , lookup
Protein structure prediction wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Gene expression wikipedia , lookup
Peptide synthesis wikipedia , lookup
Proteolysis wikipedia , lookup
Messenger RNA wikipedia , lookup
Biochemistry wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Genetic code wikipedia , lookup
Epitranscriptome wikipedia , lookup
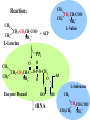
Protein Biosynthesis • Away from the Ribosome Genetic code Charging tRNA Ribosomes • On the Ribosome Initiation complex Elongation factors Peptide bond formation Termination Characteristics of the Code • • • • • • • • non-overlapping degenerate triplet basically universal over all living species polar no punctuation subject to miscues defines a reading frame Framing the Code OLD MAN AND THE SEA Ernest Hemmingway Garbled OLD MAA NAN DTH ESE A Insert A Garbled OLD MAA ANA NDT HES EA Insert A A In frame OLD MAA ANA AND THE SEA Insert AAA N In frame OLD MAA NAD THE SEA ^ Insert A delete N tRNA Charging tRNA Q: What is meant by Charging A: Charging means placing an amino acid on the 3’ (acceptor) end of the tRNA Q: So, what’s the big deal? A: There are 20 amino acids; the code is degenerate There could be 4 “isoaccepting tRNAs” competing for one Q: I still don’t see a problem A: One enzyme must recognize 4 different tRNA species and select the correct amino acid. Q: One enzyme does all that? A: No, each tRNA has its own enzyme Q: What is this enzyme called? A: Its call Aminoacyl-tRNA Synthetase Q: So, there are 20 of these enzymes A: Yes Q: That makes the job a recognition a little easier then? A: Yes, but the enzymes still have to distinguish between look-alikes such as leucine and valine, glutamine and glutamate, tyrosine and phenylalanine. Q: Are all aminoacyl-tRNA synthetases alike? A: Yes and no. Yes, they perform the same function, i.e., to recognize and transfer the correct amino acid to tRNA. Q: Why no? A: Because one class (Class I) looks for the anticodon on the tRNA, the other (Class II) looks for other features. Q: What else? A: Class I puts the amino acid on the 2’ position of the terminal ribose on tRNA, Class II only the 3’. Q: So, how does aminoacyl-tRNA synthetase discriminate amino acids and different tRNA species? A: The key lies in the tRNA itself. Besides the anticodon, tRNAs have other bases that set them apart. These bases called “identity elements” are found in the terminal ends (acceptor stem) and internal in the tRNA. Q: Do they also proofreading? A: Yes, but sparingly Q: How sparingly? A: Enough to keep errors down to isoleucine mistaken for a valine once every 50,000 times. Ile-tRNA synthetase actually hydrolyzes the valine-AMP precursor. CH3 Reaction: CH3 CH3 CH2 CH-COOCH3 NH3+ L-Valine CH2-CH2CH-COO- + ATP NH3+ L-Leucine PPi CH3 CH3 O O CH2-CH2CH-C ~O-P-O-CH2 O NH3+ O Ad L-Isoleucine Enzyme Bound HO tRNA OH CH3 CH2CH-COOCH3CH2 NH3+ Codon-Anticodon Interactions Polarity 3’ 5’ 5’ Anticodon on tRNA 3’ Codon on mRNA 3’ 5’ Anticodon loop mRNA 5’ 3rd position Alanine C G G C I A (C, U) Wobble base on anticodon 3’ mRNAs are always read 5’ to 3’. Crevice Ribosomes: The Staging Areas of Protein Synthesis Ribonucleoprotein Particles 70S (80S) Monosomes 30S (40S) 16S RNA (18S) 23 Peptides (33) 50S (60S) 23S RNA (28S) 31 Peptides (49) 5S RNA (5S + 5.8S) * Mammalian tRNA sites mRNA 30S 50S Polysomes Groups of ribosomes attached to a single mRNA