* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Three-Point Binding Model
Western blot wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Polyadenylation wikipedia , lookup
Citric acid cycle wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Messenger RNA wikipedia , lookup
List of types of proteins wikipedia , lookup
Gene expression wikipedia , lookup
Biochemistry wikipedia , lookup
Catalytic triad wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Genetic code wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
P-type ATPase wikipedia , lookup
Metalloprotein wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Ligand binding assay wikipedia , lookup
Non-coding RNA wikipedia , lookup
Cooperative binding wikipedia , lookup
Bottromycin wikipedia , lookup
Expanded genetic code wikipedia , lookup
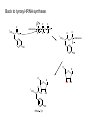
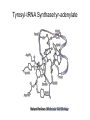
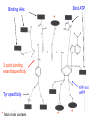
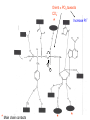
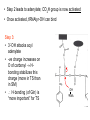
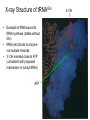
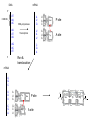
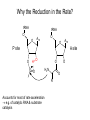
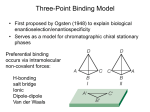
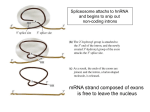
Three-Point Binding Model • First proposed by Ogsten (1948) to explain biological enantioselection/enantiospecificity • Serves as a model for chromatographic chiral stationary phases Preferential binding occurs via intramolecular non-covalent forces: H-bonding salt bridge Ionic Dipole-dipole Van der Waals Enantioselection by an Enzyme HO * glycerol kinase OH OH achiral CH2OH moieties are different because of non-equivalent binding sites in the enzyme 2- HO H OPO3 OH 3-glycerol phosphate Three-Point Binding Model - Enantiospecificity • Only one enantiomer binds to enzyme & is involved in reaction A A 3 interactions D B C b c good fit, high H d D B b C c d With the other enantiomer… A C 1 interaction D poor fit, low H B b d c B A b 2 interactions D poor fit, low H C c d • we get enantiospecificity (substrate & biomolecule are chiral) • To do this efficiently, we need a large biomolecule to align three binding sites to give high specificity • One problem with model: – Model is a static representation → “lock & key” Binding • The cost of binding: E + S K E.S (Michaelis Complex) Km (Michaelis constant): small value indicates high affinity for substrate Kbinding ( ~ 1/Km) Strong binding → K > 1 ΔG= -RT ln K ΔG must be –ve ΔGbinding = ΔHbinding- TΔSbinding For 2 molecules in, 1 out: ΔS is –ve (-TΔS) term is +ve Entropy disfavors binding of substrate to enzyme To get good binding, need –ve ΔH (i.e. bond formation) • Each non-covalent interaction is small (H-bond ~ 5 kcal/mol), but still gives a –ve ΔH • Enzymes use many FG’s to sum up many weak noncovalent interactions (i.e. 3 points) Back to tyrosyl-tRNA synthase: O O + O NH3 Adenosine P O P O P O O O O O O + O O NH3 O P Adenosine O OH OH R O R O + B O NH3 O OH OH tRNA Tyr OH B OH Tyrosyl-tRNA synthase • Use binding to orient CO2- nucleophile adjacent to P specifically as electrophile → specificity • Many non-covalent interactions overcome entropy of binding: H-bonds Enz.Tyr .ATP Tyrosine + ATP + Enz Enz.Tyr--AMP + H2P2O73(released) bind tRNA OH Enz.Tyr--OtRNA + AMP Enz.Tyr--AMP . tRNA (released) Enz + Tyr--OtRNA Can isolate this complex in the absence of tRNA Tyrosyl-tRNA Synthase.tyr-adenylate Bind ATP Binding AAs * 3 point binding enantiospecificity ATP, not dATP Tyr specificity * Main chain contacts * * Orient PO4 towards CO2Increase P+ * O P P * Main chain contacts * • We have examined the crystal structure of tyrosyl-tRNA synthase (Tyr & ATP bound) – Key contacts – 3 point binding model for (S)-tyrosine • We inferred geometry of bound ATP prior to reaction (i.e. ATP is no longer bound to enzyme) Step 1: • CO2- attacks PO42- () giving pentacoordinate P (trigonal bipyramidal) intermediate O O O O + -O P -O O Ad O O -O P O -O Ad Step 2: • Diphosphate must leave • Cannot “see” this step PPi has already left the enzyme site in the crystal structure • However, can use model building to include P & P of ATP: Thr40 & His45 form H-bonds to P **Stronger Hbonds are formed in TS than in trig. Bipyramidal intermediate Lower TS energy accelerate collapse of intermediate Gln195 Tests of Mechanism 1) Site-directed mutagenesis – Replace Gln195 with Gly (Gln195Gly) • Rate slows by > 1000 fold • ΔΔG ~ 4 kcal/mol • Developing -ve charge (on oxygen) in TS is no longer stabilized • Energy diagram? • Other mutants: – Tyr34Phe – His48Gly – These other mutations showed smaller decreases in ΔG – All contribute in some way to stabilize TS 2) Do Thr-40 & His-45 really bind / phosphates? Thr 40 Ala ( 7000 fold) His 45 Gly ( 300 fold) Both decelerate the reaction Double mutant 300,000 fold slower! A Chemical Model for Adenylate Reaction Mimic the proximity effect in an enzyme with small organic molecules: O O O O O O P O NO2 O P O O O + -O Rate is comparable to tyrosyl-adenylate formation unimolecular reaction NO2 Detect by UV • Step 2 leads to adenylate; CO2H group is now activated • Once activated, tRNAtyr-OH can bind Step 3: • 3’-OH attacks acyl adenylate • -ve charge increases on O of carbonyl Hbonding stabilizes this charge (more in TS than in SM) • H-bonding (of Gln) is “more important” for TS OH tRNA X-ray Structure of tRNAGln • Example of tRNA bound to tRNA synthase (stable without Gln) • tRNA (red) binds to enzyme via multiple H-bonds • 3’-OH oriented close to ATP (consistent with proposed mechanism in tyrosyl-tRNA) ATP 3-’OH Unique Role of Methionine • Recall, Methionine is the 1st amino acid in a peptide/protein (start codon) • As seen previously, Met is also formylated From N-formyltetrahydrofolate O fMet S tRNA S OH H O HO2C NH2 NH2 tRNA O Met Met-tRNA fMet S X O tRNA O N H O H fMet-tRNA fMet protected Reaction is catalysed by becoming pseudintramolecular (recall DNA template synthesis): Ribosome holds pieces together Ribosome is cellular “workbench” Protection with formyl group allows condensation one way around only (only one nucleophile) tRNAfMet falls off P site Dipeptide moves over to P site Control of Sequence • mRNA (messenger RNA) made by copying sequence of DNA in gene • Goes to ribosome, along with rRNA (ribosomal RNA-part of ribosome structure) & tRNA (with AAs attached) • In mRNA, 3 nucleotides of specific sequence encode 1 amino acid (CODON) • R-tRNAR has 3 nucleotides complementary through base pairing to the codon for R • Specific binding at A site • Codons for start & stop control the final protein length DNA mRNA 5' dT dA dC CODON dA dT dA RNA polymerase Transcription dG dC dT 3' A U G U A C Met P site U A U A U A Tyr A site C G A Rxn & translocation mRNA A G C U A U A U A Tyr C G A G C U Arg O Met P site A site H O N H Met Tyr Arg O Catalysis of Reaction? • Synthesis on ribosome is faster by 107 than rxn without ribosome • Peptide formation is not catalyzed by protein → no protein within 20 Ǻ of “active site” • rRNA (catalytic RNA) has been proposed : A2486 NH2 N N Adenosine from rRNA N N tRNA O O H R N pep O H O tRNA NH2 N N N N N N N H NH2 N + O O tRNAO tRNA R N H pep pep O O OH tRNA R N H O O tRNA • However, modification of bases has shown little effect on catalytic activity (2-fold decrease) • May be the 2’-OH (of tRNA) at last nucleotide on P site: i.e., the substrate! (see Nature Struct. Mol. Biol. (2004), 11, p 1101 Modified sugar at 3’OH: • OH → H • OH → F Both substitutions reduce rate by 106! adenosine Why the Reduction in the Rate? tRNA tRNA O O O A76 O A76 P site A site O H O O O H2N O R Accounts for most of rate acceleration e.g. of catalytic RNA & substrate catalysis O