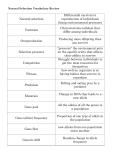
* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Variation
Saethre–Chotzen syndrome wikipedia , lookup
Gene expression profiling wikipedia , lookup
X-inactivation wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
SNP genotyping wikipedia , lookup
Public health genomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Gene desert wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Gene therapy wikipedia , lookup
Behavioural genetics wikipedia , lookup
Point mutation wikipedia , lookup
Genome (book) wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Genomic imprinting wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genetic engineering wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Gene nomenclature wikipedia , lookup
Helitron (biology) wikipedia , lookup
Gene expression programming wikipedia , lookup
Human leukocyte antigen wikipedia , lookup
Human genetic variation wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
Heritability of IQ wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Population genetics wikipedia , lookup
Genetic drift wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
phenotype - morphological, physiological, biochemical or behavioral characteristics, resulting from a variety of sources genotype - genetic constitution (of an organism or a population) e.g., A1A1 or A1A2 or A1B1 / A1B2, etc. i.e., homozygote, heterozygote, multilocus gene - classically phenotypic, but technically a DNA sequence that is transcribed and translated into protein, as well as its related promoters and modifiers allele - a version of a gene haplotype - a haploid genotype or allele (it the alleles are distinguished by nucleic acid sequence), or a profile of alleles (e.g., fingerprint) mutation - verb: alteration of genetic material; noun: an altered form of genetic material locus - a position or site on a chromosome, as small as a base or as large as a gene polymorphism - more than one allele at a locus in a population polygenic trait - more than one locus controlling a phenotype additivity phenotype of offspring is due to the combined or "added" effects of 2 or more loci A allele adds one unit of effect to phenotype B allele adds two units genotypes effect B 1 B1 2 B1B2 4 B2B2 6 A1A1 3 A1A2 4 A2A2 5 5 7 9 6 8 10 7 9 11 dominance - the property of one allele affecting phenotype to the complete or incomplete exclusion of another allele may be rare or common or neither, advantageous or deleterious or neither recessive - opposite of a dominant allele epistasis - nonadditive interaction of two or more loci on a phenotype or fitness; modification of phenotypic expression of one locus by another; analogous to dominance of one locus over another pleiotropy - multiple effects of a single gene e.g.1, Waardenberg syndrome in humans, defective allele for pigmentation responsible for both (some cases of) blue eyes and deafness; 40% of white cats with blue eyes are deaf, or ipsilaterally blue-eyed and deaf e.g.2, phenylketonuria defective allele of phenylalanine hydroxylase gene responsible for converting phenylalanine to tyrosine mental retardation, eczema, light skin e.g.3, sickle cell anemia "tower skull", impaired mental function, heart failure, pneumonia, rheumatism, paralysis, abdominal pain, kidney failure, fibrosis of spleen incomplete dominance = semidominance = - intermediate phenotype of heterozygote but not equal effects from both alleles codominance - protein products of both alleles in heterozygote detectable phenotypically overdominance (heterosis, heterozygote advantage, or hybrid vigor) - heterozygote has phenotype or fitness characteristics outside the range of either homozygote (more fit) underdominance - heterozygote disadvantage linkage disequilibrium - nonindependent assortment of 2 or more loci or of particular alleles at two or more loci due to physical proximity or meiotic drive/segregation distortion or selection genotype frequency - the proportion of a population with a certain genotype, e.g., 25% A1A1 50% A1A2 25% A2A2 allele frequency - the proportion of gene copies in a population with a certain allele (remember that there are 2 gene copies per diploid organism) allele freq = sum (2 per homozygote + 1 per heterozygote) total number of gene copies of all alleles example genotypes A1A1 A1A2 A2A2 genotype frequencies (0.25) (0.50) (0.25) alleles A1 A2 allele frequencies (0.50) (0.50) i.e., 1(0.25) + ½(0.50) = 0.50 Sources of Phenotypic Variation Genotype Environment Epigenesis – mitotically and meiotically heritable changes in gene expression that do not involve a change in DNA sequence, or differentiation and morphogenesis from “above” (other than) nucleotide sequence gene regulation by non-coding RNA (ncRNA), including small (sRNA), micro (miRNA), and inhibitory (RNAi) cytosine modifications, e.g., methylation histone modifications, e.g., lysine acetylation repressor proteins endosymbionts maternal effects, e.g., nutrition, toxins indirect effects, e.g., enlargement of the vertebrate brain due to hydrostatic pressure within the neural tube Heritability heritability is the proportion of phenotypic variance that is attributable to genetic variance Conceptually: 2 h = VG VG + VE = VG VP VG = genotypic variance VP = phenotypic variance VE = environmental variance Heritability Phenotype of offspring h2 is actually the coefficient of determination (r2) in a regression analysis, i.e., how much of the variation in y is explained by variation in x (values range from 0.0-1.0) r² = 0.8749 phenotype of parents