* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Mechanisms
Lipid signaling wikipedia , lookup
Biosynthesis wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
DNA repair protein XRCC4 wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Mitochondrion wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Transformation (genetics) wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Point mutation wikipedia , lookup
Proteolysis wikipedia , lookup
Biochemical cascade wikipedia , lookup
Western blot wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Free-radical theory of aging wikipedia , lookup
Reactive oxygen species wikipedia , lookup
Signal transduction wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
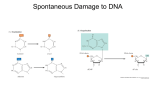
Reactions with target molecules Cellular deregulation Repair mechanisms “Essentials of Toxicology” by Klaassen Curtis D. and Watkins John B Chapter 3 Stages of toxicity …See Figure 3.1 1. Delivery 2a. Interaction with target molecule 2b. Alteration of biological environment 3. Cellular dysfunction 4. Repair or repair failure Successful repair No Toxicity No/inadequate repair Toxicity 1. Delivery Delivery to target site Concentration at target site Absorption Distribution toward target Re-absorption Activation Toxicity Elimination Distribution away from target Excretion De-activation No Toxicity Stages of toxicity …See Figure 3.1 1. Delivery 2a. Interaction with target molecule 2b. Alteration of biological environment 3. Cellular dysfunction 4. Repair or repair failure Mechanisms of toxicity Molecular targets are usually proteins, lipids, coenzymes, or nucleic acids, but rarely carbohydrates Three basic mechanisms – Formation of a stable non-covalent complex with receptor, enzyme, cofactor – Induction of a physicochemical change, e.g. pH, pO2, solvation, physical damage – Formation of reactive intermediate that binds covalently to macromolecules and/or triggers immune response Mechanism of action Effect on specific biochemical process that leads to disruption/alteration of cellular function that eventually results in impaired physiological function (health effect) (Transient or permanent…) Symptom is the observed manifestation of a health effect (outward, macroscopic) Mechanisms of action • Disruption or destruction of cell membrane (oxidative species, e.g. radical species) • Direct binding to cell molecule (CO+Hb; adducts, lead) • Enzyme inhibition – Cofactor • Inactivation (sequestration of cofactor) • Competition (replacement) – Binding to active site • Directly (classic enzyme inhibitors) • Indirectly: toxic metabolite binds See also Chapter 3 of Casarett and Doull’s “Toxicology” Mechanisms of action • Secondary action: release of endogenous substance that causes damage (histamine, neuropeptides, metals displacement) • Free-radical cascade reactions (damage to proteins, DNA, lipids, mitochondria) • Structural analogue properties – – – – Neuroendocrine context Receptor involvement Agonists (mimic action of endogenous substance) Antagonists (block action of endogenous substance) Cytochrome oxidase inhibition by cyanide stops mitochondrial respiration Mechanism of action - dioxin http://www.stanford.edu/group/whitlock/research.html Metabolism of bromobenzene to reactive epoxide intermediates which deplete glutathione and cause liver toxicity Metabolism of halothane leads to direct and indirect (immune) toxicity Carbon tetrachloride toxicity via free radical formation Redox cycling of herbicide Paraquat produces reactive oxygen species Coupling reactions: (H2O2) HOOH CAT 2H2O HOOH O2 2H+ O2- . SOD HOOH O2- . O2 GSSG 2GSH (H2O2) 2H2O HOOH GPX Effects of oxidative species on proteins: Oxidation of: • sulphydryls • amines • alcohols • aldehydes Aminoacids targets: • cystein • methionine • tryptophan • tyrosine Inactivation/inhibition of enzymes in cellular compartments Effects of oxidative species on lipids: • Polyunsaturated fatty acids (PUFA): primary target of O3 peroxidation of membrane lipids • Most important mechanism of O3-induced injury O3 + PUFA carbonyl oxide aldehydes H2O Hydroxyhydroperoxy compound . HO H2O2 Lipid peroxidation cascade Lipid fragmentation Malondialdehyde (MDA) 8-isoprostane LTB4 (PMN chemotractant) Lipid peroxidation cascade Effects on nucleic acids Electrophiles react with strong nucleophilic atoms of nucleic acids . DNA + HO Imidazole ring-opened purines or ring-contracted pyrimidines Strand breaks Blocked DNA replication Formation of adducts depurination (apurinic sites: mutagenic) Reactions with target molecules • Non-covalent – – – – Receptors Ion channels Enzymes Co-factor depletion • Covalent binding – DNA – Proteins • H removal (neutral radicals) – Amino acid CH2 – Proteins • e- transfer – Hemoglobin Fe2+ hemoglobin Fe3+ (methemoglobin) • Enzymatic reactions – Protein toxins (diphtheria, cholera) Effects on target molecules • Dysfunction – – – – Mimics endogenous molecule Inhibition, blocking (receptors, ion channels) Conformational change DNA mis-pairing • Destruction – Cross linking – Fragmentation – Oxidation/degradation (lipids) Effects on target molecules • Antigenicity Immune response Unchanged – Dinitrobenzene – Nickel – Penicillin Following change – Quinones – Biotransformation products Hapten formation and immune reaction: penicillin G Stages of toxicity …See Figure 3.1 1. Delivery 2a. Interaction with target molecule 2b. Alteration of biological environment 3. Cellular dysfunction 4. Repair or repair failure Alteration of biological environment • Alter pH (methanol, ethylene glycol, 2,4dinitrophenol) • Solvents and detergents • Direct chemical effect (phosgene, sulfuric acid) • Physical space occupation (silica, asbestos, ethylene glycol, CO2) Ethylene glycol toxic metabolites Stages of toxicity …See Figure 3.1 1. Delivery 2a. Interaction with target molecule 2b. Alteration of biological environment 3. Cellular dysfunction 4. Repair or repair failure 3. Cellular impairment 1. Cell regulation (fig. 3.6) A. Gene expression a. Transcription b. Signal transduction (fig. 3.7) c. Extracellular signal (hormone) B. Cellular activity (table 3.1) a. Excitable cells - neurotransmission b. Other cells (Kupffer, exocrine, pancreatic) Cellular impairment 2. Internal maintenance a. ATP depletion (Fig. 3.8, table 3.2) Oxidative phosphorylation b. Intracellular Ca+ increase (Table 3.3) Influx to cytosol » from outside (channels, membrane) » from mitochondria/ER Efflux out of cytosol » Ca+ transporters » ATPase inhibition c. ROS, RNS, radicals ATP Effects of increased cytosolic Ca+ • Inhibition of ATPase – Mito loading with Ca2+ – Dissipation of membrane potential – Reduced ATP synthesis, oxidative phosphorylation and Ca2+ cycling • Microfilament dissociation – Membrane rupture • Hydrolysis - enzyme increase – Protein, phospholipids, DNA • ROS, RNS production – Ca2+ activates dehydrogenases in citric acid cycle --> e- transport increase --> ROS, RNS Inter-relationships Ca2+ channels that control cytosolic Ca2+ need ATP Ca2+ in cytosol ATP Mito potential Ca2+ ROS, RNS Inactivated pump Inter-relationships Enzyme inhibition ROS, RNS ATP ONOO- DNA damage PARP NAD+ Mito Permeability Transition Ca2+ uptake Membrane potential ROS, RNS ATP MPT Mitochondrial damage leads to cell death Pores open (1500 Da) Ca2+ from mito to cytosol Influx of protons, negative potential ATP synthesis Osmotic H20 influx Glycolysis Energy Mito swelling ATP hydrolysis Burst Two options for cell death Robertson JD & Orrenius S. Critical Rev. Toxicology 2000, Sep; 30(5):609-27 “Molecular mechanisms of apoptosis induced by cytotoxic chemicals” http://www.roche-applied-science.com/prodinfo_fst.htm?/apoptosis MTP - cell death • • • • • • Necrosis Extensive damage All mito Multiple metabolic defects Random sequence ATP severely depleted Cell swelling and lysis • • • • • • Apoptosis Less extensive Some/many mito Some metabolic defects Ordered sequence Some ATP available Cell shrinkage, membrane bound fragments Stages of toxicity …See Figure 3.1 1. Delivery 2a. Interaction with target molecule 2b. Alteration of biological environment 3. Cellular dysfunction 4. Repair or repair failure Levels of repair Molecular repair • • • • • Proteins reduction (re-activation) NADPH Protein refolding (heat-shock proteins) Protein degradation and re-synthesis Lipid reduction (GPO, GR, NADPH) DNA repair DNA damage repair • Direct: photolyase (UV-dimers, O6-methyl-G removal) • Excision DNA glycosylase (removes AP site) AP endonuclease (PO3 bond) DNA polymerase (replicates sequence) Ligase (ties the ends) PARP (multiple ADP ribose - unfolds/facilitates repair) • Recombination Sister chromatid exchange Cellular/Tissue repair • Single cell - regeneration (neurons) • Tissue – Apoptosis – Proliferation • Chemokine priming (G0-G1) :TNFa, IL-6 • Chemokine progression (G1-GM) :HGF, TGFa – Migration – ECM (Stellate cells, PDGF, TGFb) Inflammation Macro’s IL-1, TNFa endothelia, fibroblasts Vascular dilation Leukocyte infiltration Release of PAF, LTB4, cytokines Leuko-endo adhesion Side reactions - Inflammatory oxidative burst . Three pathways of HO generation: • NAD(P)H oxidase • Nitric oxide synthase (NOS) • Myeloperoxidase (MPO) L-arginine + O2 L-citruline NOS H+ . . O 2 NAD(P)+ H+ HOOH + H+ +Cl- NO2 . NO Oxidase NAD(P)H + O2 (macro’s and granulo’s) (macro’s) (granulo’s) MPO H20 HOCl Fenton HO O2 Cl- . More side reactions • Gene expression – Cytokines IL-6, IL-1, TNFa – Acute phase proteins • • • • Minimize injury Facilitate repair (inhibit lysosomal proteases) Plasma proteins CYP450, GSTs (detox) • Generalized reactions – Fever (IL-1, IL-6, TNFa) hypothalamus – Pituitary (ACTH --> cortisol) (negative feedback) Repair failure • Tissue necrosis – No apoptosis, no proliferation, dose matters • Fibrosis – ECM deposition – TGFb matrix synthesis, autocrine control Repair failure cont. • Carcinogenesis – Failure to repair DNA in: • Proto-oncogenes - activating mutation (GF, recept., TF, signal transduction proteins) • Tumor suppressor genes - inactivating mutation (p53, protein kinase inhibitors, TF) – Failure of apoptosis – Failure to stop proliferation • • • • • Mutation accumulation Repair is less likely Neoplastic transformation (reduced methylation) Reduced cell-cell contact Inhibition of cell-matrix contact Factors determining specificity • Sensitivity – Neurons and heart require high levels of O2 to make ATP via mitochondrial respiration; CO and CN are therefore very toxic to these organs. – Bone marrow and gut epithelium contain rapidly dividing cells; mitotic substances damage these tissues. • Distribution – Inorganic mercury (Hg) cannot cross the blood-brain barrier; methylmercury can. • Selective uptake – Strontium 90 into bone instead of calcium – Paraquat mistaken for polyamines in lung cells Factors determining specificity • Metabolism – Localized activation in Clara cells of lung – Absence of detoxification: eye lacks formaldehyde dehydrogenase (methanol blindness) • Lack of repair mechanism – Liver has high capacity to remove O6alkylguanine but brain capacity is low